Clear Sky Science · en
Whole-field, high-resolution Fourier ptychography with neural pupil engineering
Sharper Views Across the Whole Slide
Modern microscopes can reveal astonishing cellular detail—but usually only in a small sweet spot near the center of the image. At the edges of a large tissue slide, fine structures often blur and fade, limiting how much doctors and researchers can trust what they see. This paper introduces a new way to push a powerful imaging method, Fourier ptychographic microscopy, closer to its theoretical limits, delivering crisp detail across an entire large field of view without rebuilding the microscope from scratch.
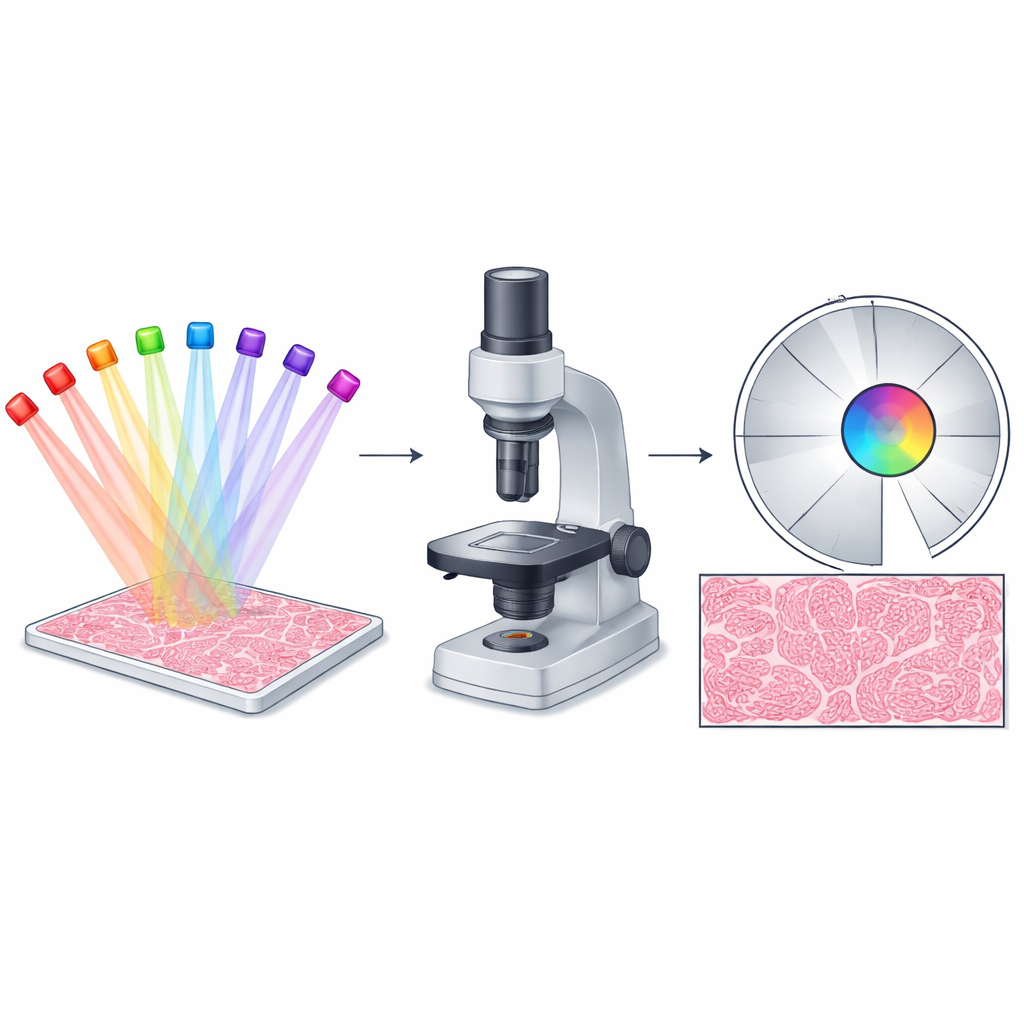
Why Microscopes Struggle at the Edges
Fourier ptychographic microscopy (FPM) works by shining light onto a sample from many different angles and then using a computer to combine the resulting low-resolution snapshots into a single, high-resolution picture. In principle, this strategy should give images that are both very sharp and very wide—ideal for whole-slide pathology, live cell studies, and industrial inspection. In practice, though, FPM performs best only near the optical center. Farther out, imperfections in lenses and the curved wavefronts from LED illumination break a simplifying assumption that the imaging system behaves the same everywhere. As a result, the edges of the field show artifacts, lost contrast, and missing fine details, even though the center looks excellent.
A Smart, Shape-Shifting Aperture
The heart of the problem lies in how FPM typically handles the microscope’s pupil function, an optical "window" that defines which parts of the light’s spatial frequencies get through. Standard FPM treats this window as a fixed, centered circle in a mathematical space related to spatial frequencies. The authors observed that in real experiments, especially for regions away from the center, the effective window is subtly shifted. Instead of trying to handcraft a more complicated physical model, they let a neural network learn how this window should move. Their approach, called neural pupil engineering FPM (NePE-FPM), represents the pupil as a continuous function encoded by a tiny neural network and a multi-resolution hash table. This setup allows the pupil to slide smoothly in frequency space during reconstruction so that the algorithm can adapt to off-axis behavior without adding extra hard-to-measure system parameters.
Clearer Cells and Crisper Patterns
To test their method, the researchers imaged plant root tissue and standard resolution targets. Compared with conventional FPM that uses a fixed pupil, NePE-FPM produced significantly sharper cell boundaries and higher image contrast at the edges of the field of view. Quantitative tests showed up to about 55% improvement in contrast in some regions, with individual stained cells becoming clearly distinguishable where they were previously blurred. On a publicly available resolution target designed to stress FPM, competing algorithms struggled to recover both amplitude and phase accurately when the illumination’s curvature was important. NePE-FPM, by contrast, preserved fine stripe patterns and yielded more accurate phase maps, a key requirement for quantitative, label-free imaging.
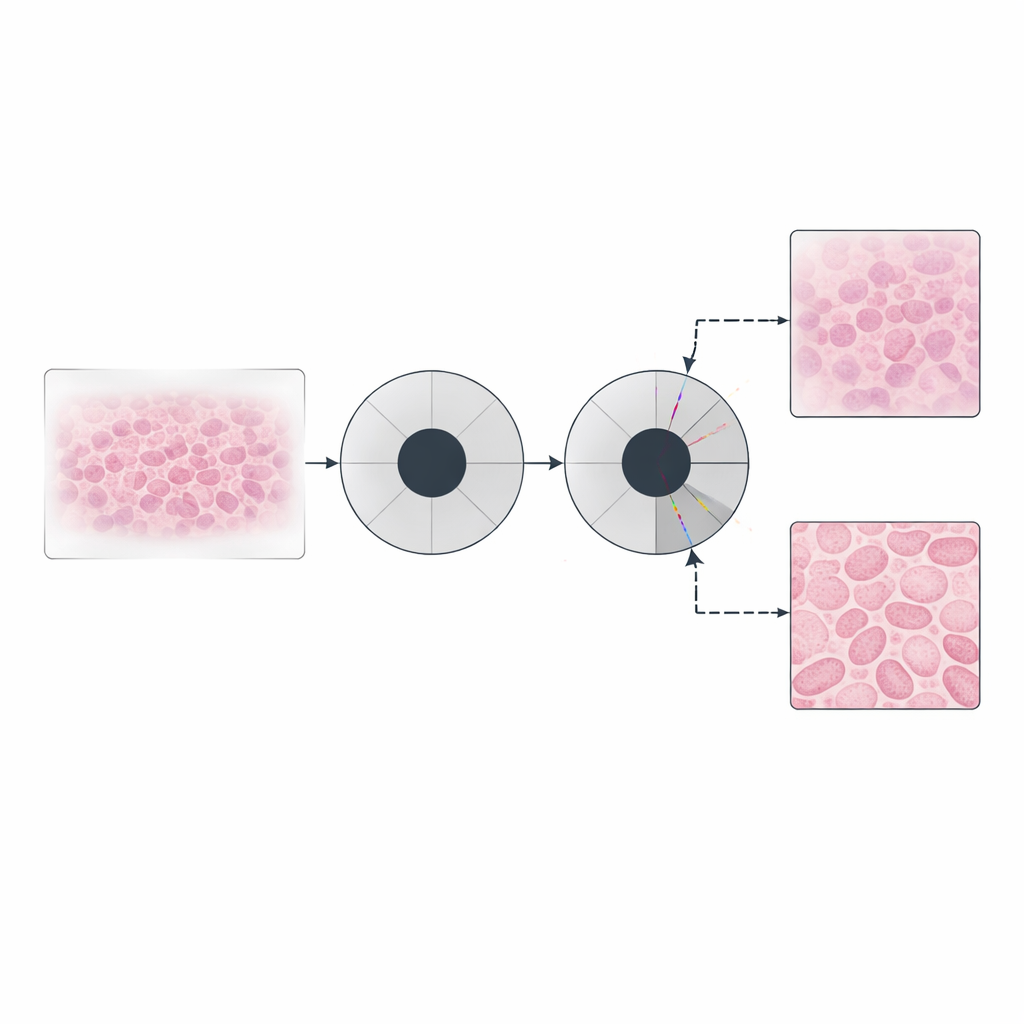
Learning Both the Sample and the Optics
The authors went further by allowing neural networks to represent not only the shifting pupil but also the sample itself. In this "double implicit" scheme, one network encodes how the sample modifies light, while another encodes how the optical window behaves across frequencies. Carefully chosen activation functions ensure that amplitudes and phases remain physically realistic. This continuous, coordinate-based description acts like a smart filter: it naturally smooths noise while preserving genuine transitions, avoiding the blocky artifacts that can appear when traditional methods rely heavily on certain kinds of regularization. Tests on tissue slices showed smoother, cleaner phase images with enhanced contrast, while still matching the underlying quantitative values.
Speeding Up for Real-World Use
Because whole-slide imaging involves huge datasets, speed matters. NePE-FPM is designed with efficiency in mind. The multi-resolution hash encoding lets the neural representation be queried in constant time, and the authors implemented custom CUDA code to handle the heavy lifting on a graphics processing unit. For typical datasets with millions of pixels and dozens of illumination angles, reconstruction times dropped to tens of seconds—about fifteen times faster than comparable CPU-based implementations—while still achieving large resolution gains across the entire field.
Bringing Theory Closer to Practice
In accessible terms, this work teaches the microscope’s "window" to move where it needs to be, instead of forcing it to stay fixed in an oversimplified model. By letting a compact neural network continuously adjust how light is filtered in frequency space, NePE-FPM recovers fine cellular details uniformly across large areas, narrows the gap between what FPM promises on paper and what it delivers in the lab, and does so at practical speeds. For applications like digital pathology or high-throughput inspection, it offers a path to gigapixel images where the edges are finally as trustworthy as the center.
Citation: Shuhe Zhang and Liangcai Cao, "Whole-field, high-resolution Fourier ptychography with neural pupil engineering," Optica 12, 1615-1624 (2025). https://doi.org/10.1364/OPTICA.575065
Keywords: Fourier ptychographic microscopy, computational imaging, neural pupil engineering, quantitative phase imaging, whole-slide microscopy