Clear Sky Science · en
A Metabolism-Informed Neural Network Identifies Pathways Influencing the Potency and Toxicity of Antimicrobial Combinations
Why this matters for everyday health
As more infections stop responding to our go-to antibiotics, doctors increasingly turn to combinations of drugs. But mixing medicines can be a double-edged sword: the right pair can clear stubborn bacteria, while the wrong mix can harm organs like the kidneys and liver. This study introduces a new computer-based approach, called CALMA, designed to help scientists and clinicians find drug combinations that are both powerful against microbes and gentler on patients.
Fighting germs without trial-and-error
Traditionally, discovering good antibiotic combinations has meant testing huge numbers of drug pairs or trios in the lab and in patients. The number of possible mixtures grows explosively with every added drug, making full experimental testing unrealistic. On top of that, most existing tools focus only on how strongly combinations kill bacteria, ignoring how they might damage human tissues. The authors argue that to design better treatments, we must consider both sides of the equation: potency against pathogens and safety for people.
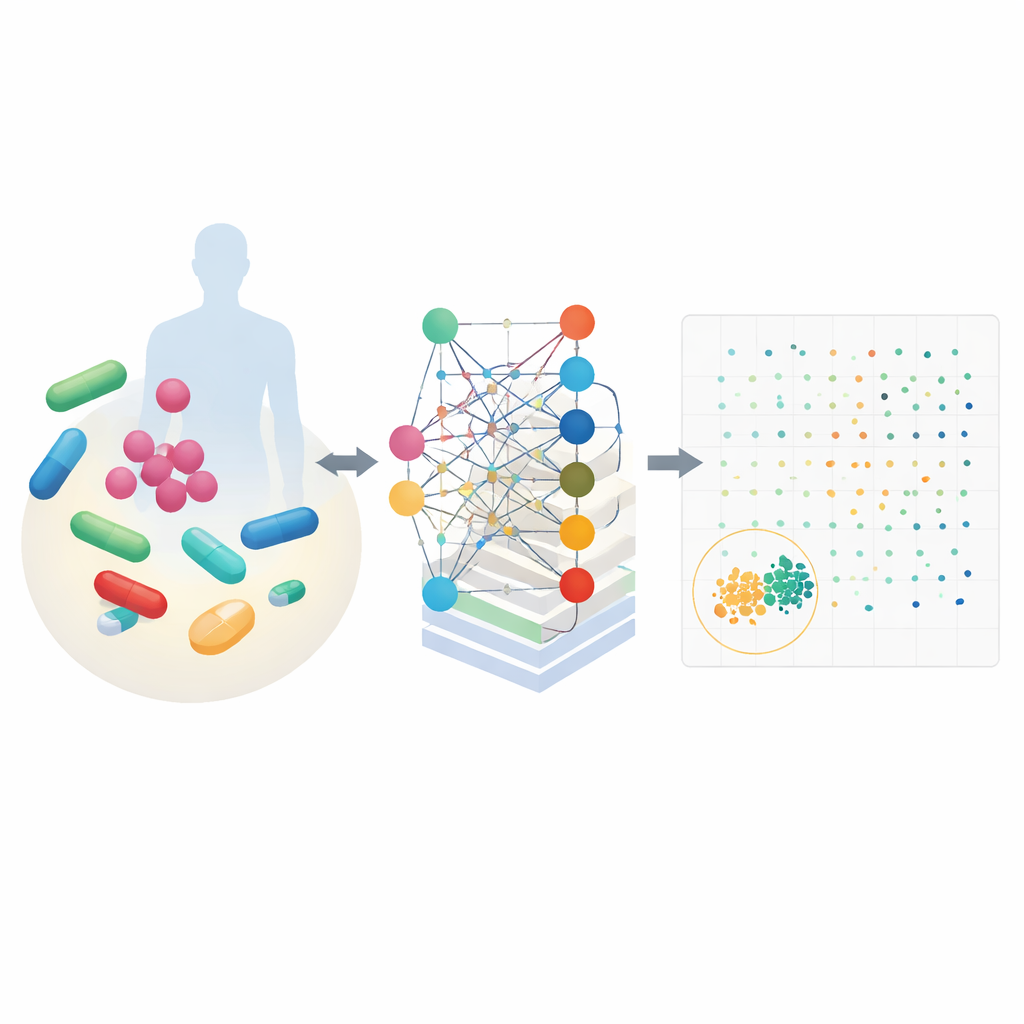
A smart model that understands cell chemistry
CALMA combines two powerful ideas. First, it uses detailed maps of cellular metabolism that describe how organisms turn nutrients into energy and building blocks. These maps, built for bacteria like Escherichia coli and the tuberculosis germ, simulate how thousands of chemical reactions change when a drug is present. Second, CALMA feeds these simulations into an artificial neural network whose structure mirrors metabolic pathways. By grouping information according to known biological routes—such as energy production or nucleotide recycling—the model learns patterns that link specific pathways to either effective bacterial killing or harmful side effects in human cells.
Scanning the landscape of drug mixtures
Using data from previous experiments and large safety databases, the researchers trained CALMA to assign each drug combination two scores: one for how strongly it attacks bacteria and another for how likely it is to be toxic in people. Plotting these scores creates a landscape where the best candidates sit in the lower left—strong against microbes and mild on human cells. When they applied CALMA to 35 clinically relevant drugs, the tool narrowed nearly 600 possible pairs down to a small set of promising options, shrinking the experimental search space by about 97 percent. Among the top suggestions were combinations involving common antibiotics such as azithromycin, vancomycin, isoniazid, and trimethoprim.
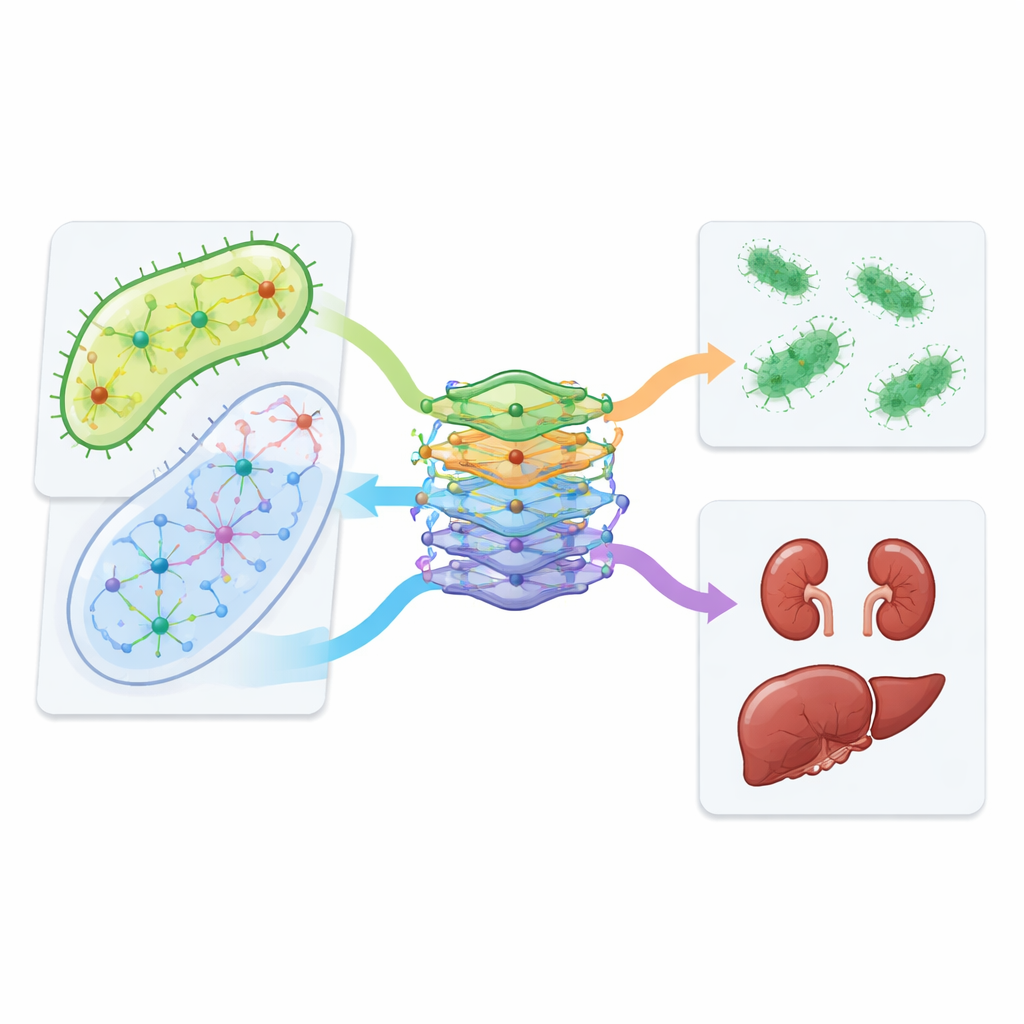
From computer predictions to lab tests and real patients
The team moved beyond simulations to test several predicted combinations in kidney and liver cell lines. They found that certain pairs, including azithromycin with vancomycin and isoniazid with trimethoprim, were less toxic to human cells than the single drugs alone, while still retaining activity against E. coli. CALMA’s design also allowed the researchers to probe which metabolic routes drive toxicity. A pathway that recycles DNA building blocks, known as nucleotide salvage, emerged as a key player: tweaking it in cell experiments changed how harmful some combinations were. Finally, the authors examined health records from hundreds of thousands of patients and observed that people receiving azithromycin together with vancomycin had fewer documented kidney problems than similar patients on vancomycin alone, echoing the lab findings.
What this means for future treatments
To a non-specialist, CALMA can be thought of as an intelligent filter that understands enough biology to suggest which drug mixtures are worth testing first. Instead of blindly mixing antibiotics and hoping for the best, researchers can focus on combinations that models predict will hit bacterial weak spots while sparing vulnerable human pathways. Although the method depends on available data and is not perfect, it offers a more rational route to designing combination therapies. Over time, tools like CALMA could help doctors treat resistant infections more safely, reduce trial-and-error in drug development, and even be adapted to other diseases where treatment combinations and side effects are major concerns.
Citation: Arora, H.S., Lev, K., Robida, A. et al. A Metabolism-Informed Neural Network Identifies Pathways Influencing the Potency and Toxicity of Antimicrobial Combinations. npj Drug Discov. 3, 11 (2026). https://doi.org/10.1038/s44386-026-00042-9
Keywords: antibiotic combinations, drug toxicity, artificial neural networks, metabolism, antimicrobial resistance