Clear Sky Science · en
Inferring signaling pathway abnormalities from histopathological images via logic-constrained gene-pathway heterogeneous knowledge graph
Seeing Hidden Cell Signals in Cancer Slides
When doctors examine cancer under a microscope, they mostly see shapes and colors, not the complex molecular conversations happening inside the cells. Yet these invisible signaling "conversations" between genes and pathways often determine how a tumor grows and how it might respond to treatment. This study introduces a new artificial intelligence system that reads routine pathology images and infers which cellular communication routes, or pathways, are going wrong—offering a way to glimpse molecular behavior without extra lab tests.
From Single Gene Clues to the Bigger Picture
Traditional approaches try to predict the status of individual genes from tissue images: is a particular gene mutated or not? That is useful, but cancer rarely depends on a single faulty gene. Instead, groups of genes work together to disrupt entire signaling pathways that control growth, survival, and immunity. The authors argue that focusing on single genes misses this cooperative behavior and can fail to explain why tumors look and act the way they do under the microscope.
Building a Map Between Images, Genes, and Pathways
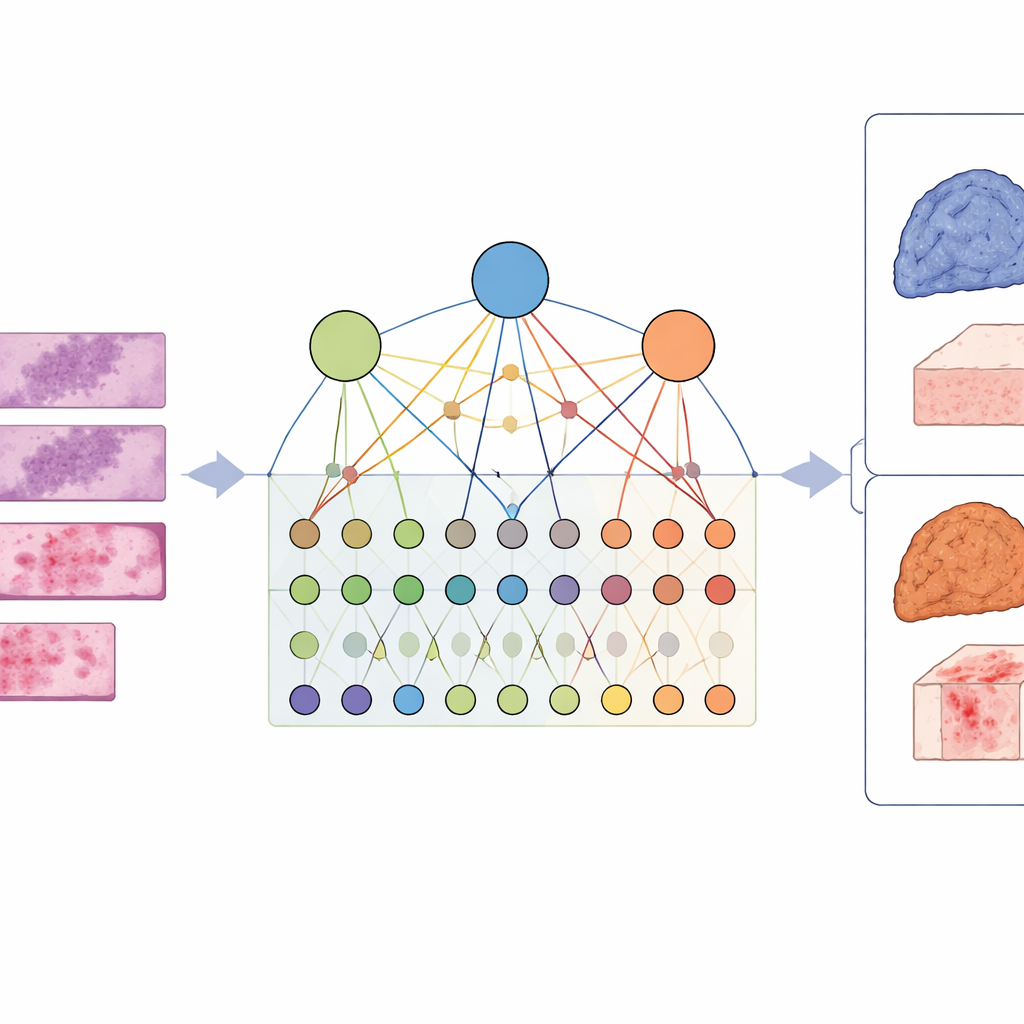
To capture this complexity, the researchers designed a framework called LCG-HGNN that links three layers of information. First, they break each whole-slide image of lung adenocarcinoma into many small patches and represent these patches as a graph, where nearby or similar-looking regions are connected. Second, they connect gene markers to signaling pathways using established biological databases, building a second graph that encodes which genes belong to which pathways. Finally, they let an AI model pass information through this combined graph, so that image patterns, genes, and pathways influence each other rather than being treated in isolation.
Teaching the System Known Biology
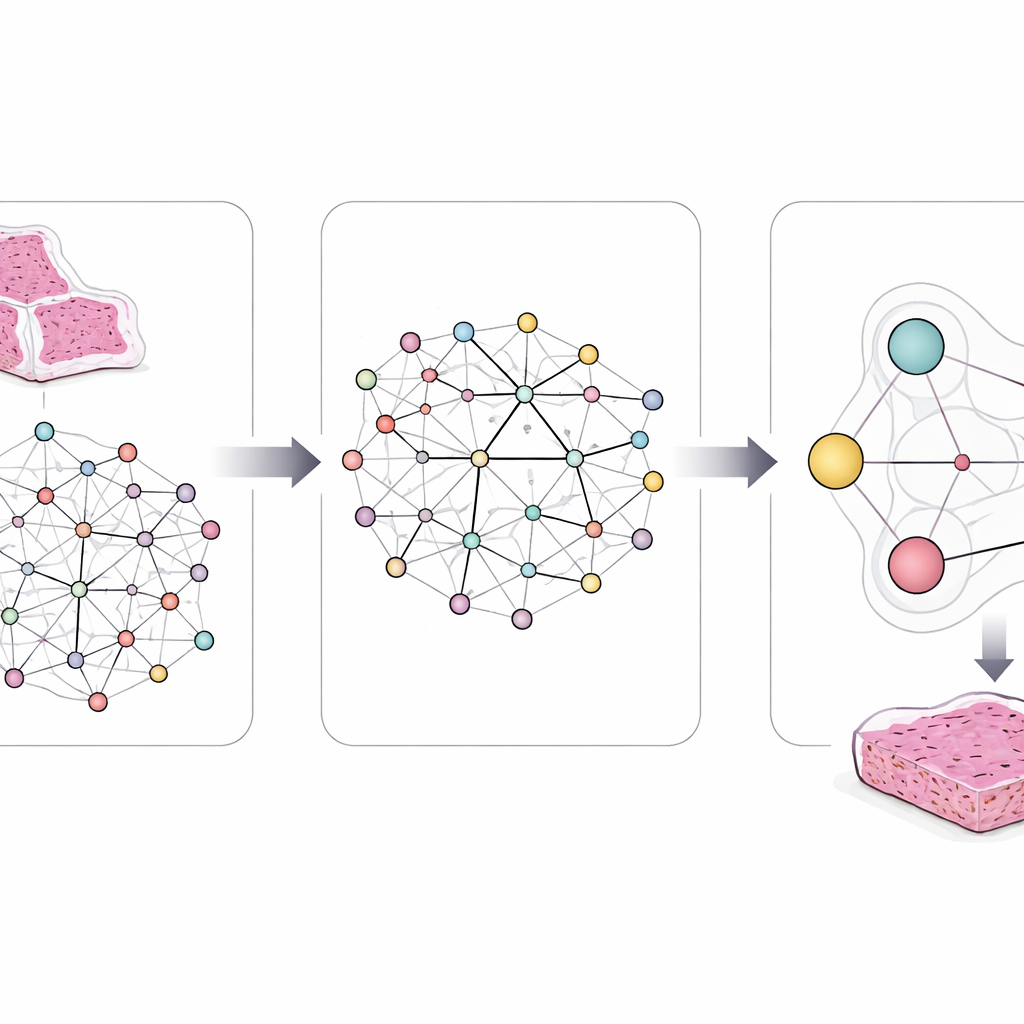
A key innovation is that the model is not allowed to learn freely without guidance. The authors embed clear, rule-like statements derived from decades of cancer research. For example, some mutations almost never appear together in the same tumor, and some pathways sit upstream or downstream of others. They encode such rules as logical constraints that gently steer the AI’s internal connections and predictions. If the model proposes a combination of gene and pathway changes that clashes with known biology or with similarities between patients’ slides, these rules push it toward more realistic alternatives.
Testing Accuracy Against Existing Methods
The team evaluated their system on over 1,600 lung cancer slides linked with genetic data. They compared it with standard deep learning models that focus on single genes and with more advanced methods that handle many genes at once but ignore pathway structure. Across several important pathways, including those involving cell growth signals and the tissue’s supporting scaffold, the new model predicted gene group changes and pathway abnormalities more accurately. It also produced image patches whose appearance closely matched real clinical examples known to be associated with specific mutations, suggesting that it was focusing on meaningful microscopic patterns rather than spurious image quirks.
Why This Matters for Patients
For a non-specialist, the main outcome is that this approach turns a routine pathology slide into a rich source of molecular insight. Instead of ordering separate, often costly genetic tests for every possible target, doctors could eventually use such models to screen for disrupted pathways directly from the images they already collect. While the method still needs larger, real-world validation and does not yet replace genomic testing, it points toward a future where the microscope view doubles as a window onto the tumor’s inner wiring—helping guide more precise and biologically informed cancer care.
Citation: Yu, Y., Shi, W., Chen, X. et al. Inferring signaling pathway abnormalities from histopathological images via logic-constrained gene-pathway heterogeneous knowledge graph. npj Biomed. Innov. 3, 23 (2026). https://doi.org/10.1038/s44385-026-00078-6
Keywords: cancer pathways, histopathology AI, graph neural networks, lung adenocarcinoma, digital pathology