Clear Sky Science · en
Nonequilibrium chemomechanical transduction of ATP-driven protein unfolding in the 26S proteasome
How a Tiny Machine Keeps Our Proteins in Line
Inside every cell, a microscopic shredder called the 26S proteasome constantly destroys damaged or no-longer-needed proteins, keeping the cell healthy. At the heart of this machine is a ring-shaped motor that burns chemical fuel (ATP) to grab, unfold, and pull proteins into a central chamber where they are chopped up. This paper uses advanced computer simulations to reveal how that motor converts chemical energy into mechanical motion, offering a detailed, quantitative picture of a process that underpins aging, brain disease, immunity, and cancer.
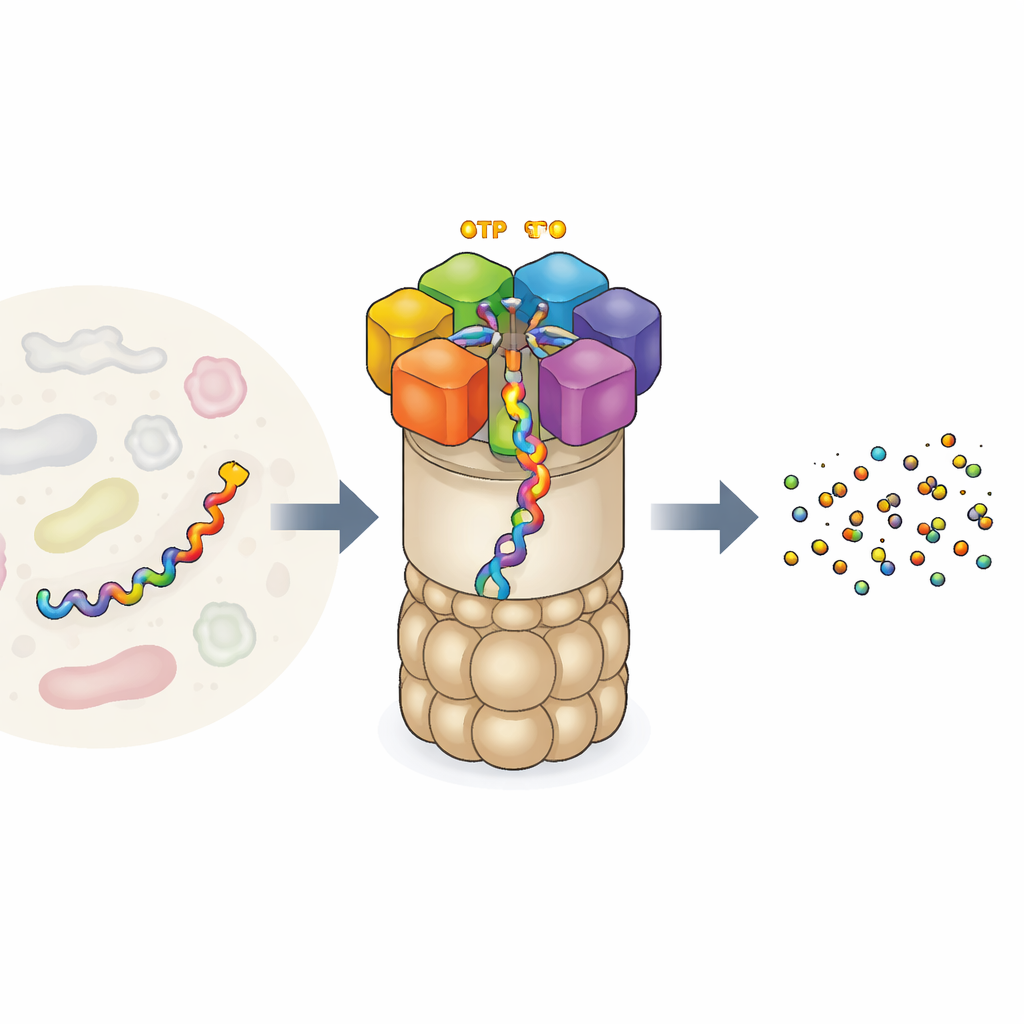
The Cell’s Protein Recycling Factory
The 26S proteasome is one of the largest and most intricate protein machines in our cells. It consists of a barrel-like core that cuts proteins and a regulatory cap that recognizes which proteins to destroy. Sitting at the gateway is a ring of six different motor units. Each unit can bind ATP, the universal energy currency of the cell, and uses that energy to grip a protein chain and pull it into the core for breakdown. This motor does not act randomly: earlier experiments suggested that the six units work in a coordinated “hand-over-hand” fashion, passing the protein along like climbers on a rope. But experiments could only see a few snapshots of this action, leaving open questions about the full sequence of movements and how exactly fuel burning is linked to mechanical work.
Simulating a Molecular Tug-of-War
The authors built a probabilistic computer model that treats the motor as a system that hops between many possible shapes while molecules of ATP and its products bind and unbind. They defined 30 main conformations of the ring in which one, two, or three motor subunits lose their grip on the protein, plus a special tightly closed state in which all six hold on. Using a standard algorithm for simulating random chemical events, they tracked hundreds of thousands of steps in which ATP binding, ATP breakdown, and motor shape changes occur. From these runs they could predict how fast a protein is pulled through under different conditions, such as varying amounts of ATP, its spent form ADP, and a non‑breakable ATP mimic often used in experiments.
When Too Much Fuel Slows the Motor
The simulations reproduced several puzzling experimental findings. As ATP levels rise from low values, the motor pulls proteins faster, because fuel binding is the slowest step. But beyond about 1 millimolar ATP, the speed peaks and then falls: the ring spends more time in a jammed, non‑translocating conformation where all six subunits are occupied by ATP yet the protein does not move. Adding ADP or the non‑breakable ATP analog slows the motor steadily, because these molecules compete with ATP for binding pockets but cannot complete the full power stroke. The model also predicts how the motor behaves when it meets very tightly folded regions of a protein, which act as roadblocks. In those cases, the motor spends more time struggling against resistance, and the overall degradation rate drops, matching measurements with artificially stabilized protein domains.
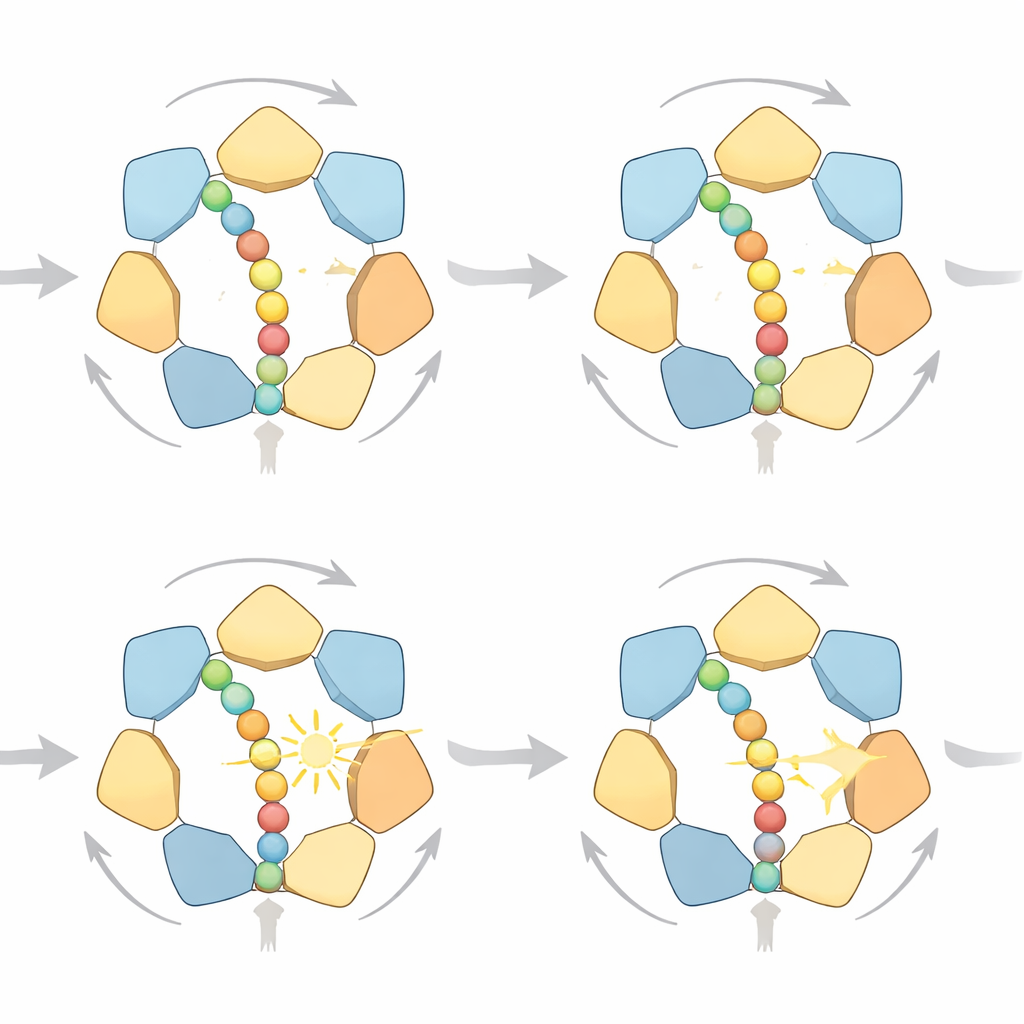
Many Ways to Take a Step
By examining the simulated trajectories in detail, the researchers found that the ring does not follow a single rigid cycle. Instead, there are multiple high‑probability pathways that all respect the same directional “hand‑over‑hand” rule: subunits near the exit of the ring release the protein after ATP breakdown, move to the top of a spiral staircase of subunits, and re‑grip the chain further up. Sometimes the protein advances by one small step, sometimes by two, depending on how many subunits let go together. Under plentiful fuel, one‑step motions dominate because they waste less work against opposing forces; when fuel is scarce, the model predicts more frequent two‑step jumps. The simulations also connect mechanical load to the chemical state of the motor: as resistance increases and the protein stalls, ADP tends to accumulate in more of the six binding pockets—exactly what high‑resolution structural studies observe.
Energy Use and Shared Design Rules
The model allows the authors to map how the motor’s energy rises when ATP is broken down and falls when that energy is turned into motion. They calculate an efficiency curve showing that the motor works best at an intermediate opposing force: too little resistance and it burns ATP wastefully; too much and it grinds nearly to a halt. When the team compared their predictions to data from related protein‑destroying machines in bacteria and yeast, they found very similar trends in how a non‑breakable ATP mimic slows these motors. This suggests that many members of the same family of ring‑shaped enzymes likely share a common, conserved mechanism for pulling on proteins.
Why This Matters for Health and Disease
By turning scattered structural snapshots and biochemical measurements into a single, testable framework, this work shows in quantitative detail how a tiny molecular motor translates chemical fuel into force to recycle the cell’s proteins. The model not only explains a wide range of existing experiments but also makes predictions about how changes in fuel levels, mechanical load, or motor mutations could alter protein destruction. Because similar machines operate across all forms of life and are central to conditions from neurodegeneration to cancer, understanding their inner workings at this level may ultimately guide the design of drugs that tune, boost, or selectively block these microscopic shredders.
Citation: Wu, D., Ouyang, Q., Wang, H. et al. Nonequilibrium chemomechanical transduction of ATP-driven protein unfolding in the 26S proteasome. npj Biol. Phys. Mech. 3, 4 (2026). https://doi.org/10.1038/s44341-026-00034-w
Keywords: proteasome, AAA+ ATPase motor, protein degradation, molecular machines, chemomechanical coupling