Clear Sky Science · en
Ultrasensitive non-enzymatic protein detection using proximity immunoassay with photonic resonator absorption microscopy
Why finding tiny traces of proteins matters
Doctors and researchers increasingly rely on proteins in blood to warn of cancer, heart disease, infections, or harmful inflammation long before symptoms become severe. But many of these molecular warning signs appear at vanishingly low levels that today’s standard lab tests struggle to detect quickly or cheaply. This study introduces a new testing method, called PINATA, that can spot extremely small amounts of a protein linked to inflammation while using simple equipment and room‑temperature steps, opening the door to more accessible and sensitive diagnostics.
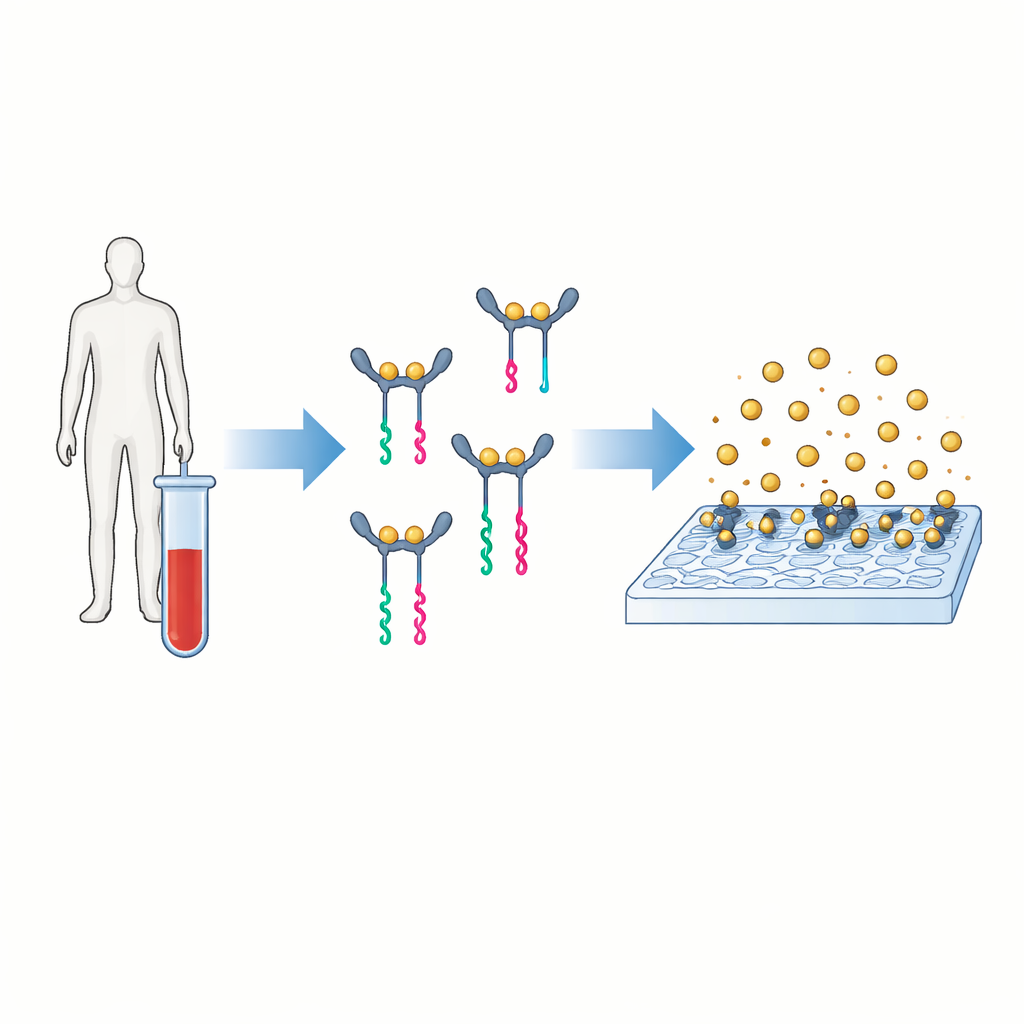
A new way to turn proteins into readable signals
The core idea behind PINATA is to translate the presence of a protein into a short piece of DNA that is easy to amplify and count. The authors focus on interleukin‑6, a signaling molecule in the blood that rises in conditions ranging from infection to cancer. In conventional tests, two antibodies latch onto the protein and carry an enzyme that generates a colored or fluorescent signal. PINATA keeps the basic notion of two antibodies recognizing the same protein, but instead of enzymes it attaches short DNA strands to each antibody. When both antibodies bind the same protein and move close together, their DNA partners cooperate to release a separate DNA fragment called a reporter. Each protein molecule can thus trigger the release of many identical DNA reporters.
Using DNA “traffic rules” instead of enzymes
At the heart of the method are carefully designed DNA circuits that behave like molecular traffic systems, directing when strands can bind, unbind, or swap partners. These circuits are built so that, unless a protein pulls the two antibody‑linked DNA pieces together, the reporter DNA stays locked in place and no signal is generated. When the protein is present, its bridging action frees the reporter. This released reporter then participates in a secondary amplification process on a prepared surface. There, it repeatedly engages in strand‑exchange reactions that allow a single reporter molecule to recruit many gold nanoparticles to the surface, creating a strong digital signal without using any enzymes or temperature cycling.
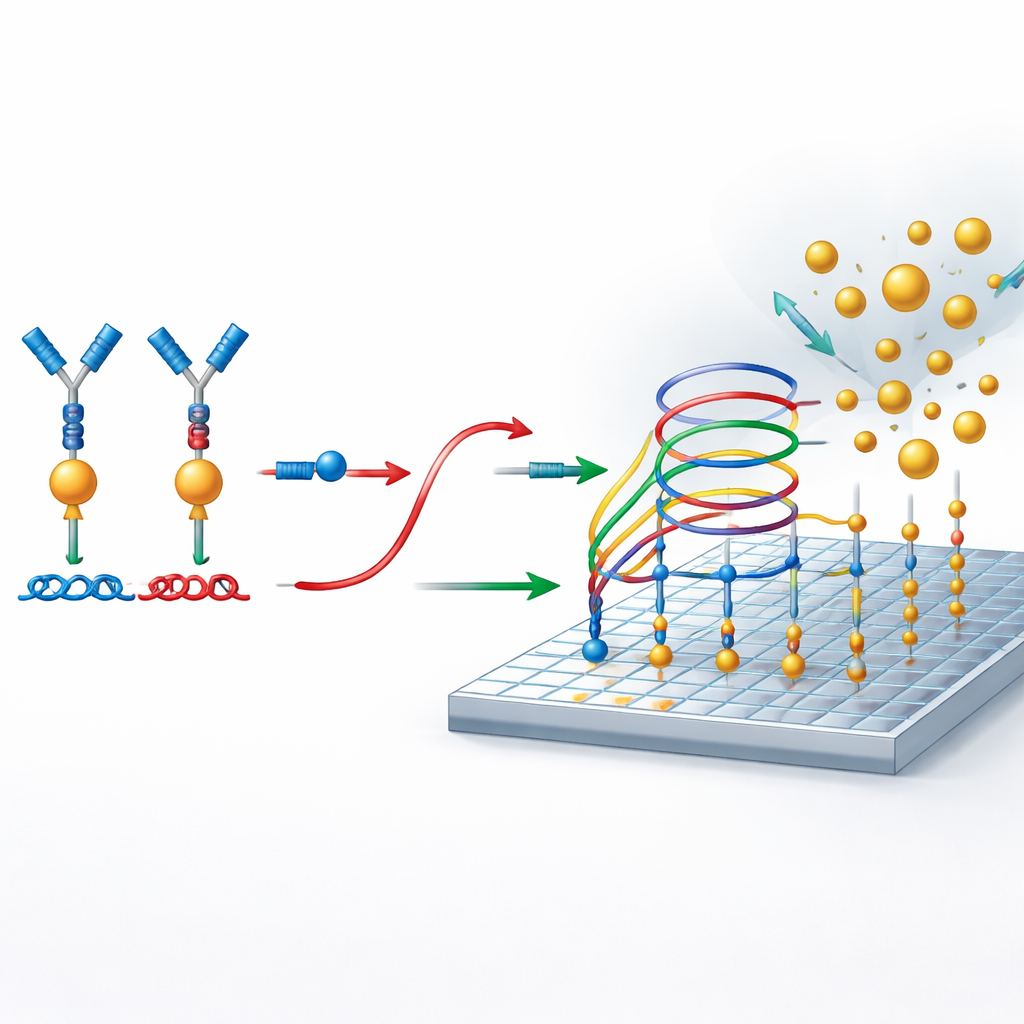
Counting single nanoparticles as yes-or-no events
To read out the result, the researchers use photonic resonator absorption microscopy, or PRAM. The sensing surface is a specially patterned material that strongly reflects light at a particular color. When gold nanoparticles land on this surface, they absorb that light and appear as dark dots in the microscope image. Because the system is designed so that nanoparticles only bind when reporter DNA is present, each dark dot represents a successful detection event linked back to a protein molecule. A simple, low‑cost optical setup and image‑processing software are then used to count these dots across the surface, turning the number of nanoparticles into a precise measurement of protein concentration.
How sensitive and selective is the test?
Using this approach, the team shows they can detect interleukin‑6 at levels as low as 37 femtograms per milliliter—roughly a few tens of molecules in a droplet—across a dynamic range spanning six orders of magnitude. The assay works in a straightforward two‑step, 90‑minute protocol performed entirely at room temperature. The authors also demonstrate that the test remains accurate even when interleukin‑6 is mixed into complex samples like human serum and plasma, which typically interfere with sensitive measurements. They further confirm that antibodies targeting interleukin‑6 do not respond to other, related proteins, underscoring the assay’s selectivity.
What this could mean for future diagnostics
For a non‑specialist, the key takeaway is that PINATA offers a way to detect disease‑related proteins at extremely low levels using a compact optical instrument instead of bulky, expensive lab hardware. By combining smart DNA circuitry with digital counting of nanoparticles, the method avoids fragile enzymes and heating steps, yet still reaches or surpasses the sensitivity of many advanced protein tests. With further development and adaptation to other targets, this strategy could enable earlier diagnosis, more frequent monitoring, and point‑of‑care testing for a wide range of conditions where tiny changes in protein levels carry big clinical meaning.
Citation: Shepherd, S., Bhaskar, S., Xu, H. et al. Ultrasensitive non-enzymatic protein detection using proximity immunoassay with photonic resonator absorption microscopy. npj Biosensing 3, 21 (2026). https://doi.org/10.1038/s44328-026-00090-1
Keywords: protein biomarker detection, ultrasensitive diagnostics, DNA strand displacement, interleukin-6 assay, digital biosensing