Clear Sky Science · en
Plasmon-enhanced bioassay for amplification-free detection and quantification of SARS-CoV-2 RNA
Why faster, simpler virus tests matter
The COVID-19 pandemic exposed how much we rely on lab-based tests that are slow, expensive, and hard to scale when millions of people need answers quickly. The work described here presents a new kind of lab test that can spot and count tiny amounts of coronavirus genetic material without the usual amplification step used in PCR. This approach aims to bring hospital-grade accuracy closer to simple, low-cost testing that could be deployed more widely during future outbreaks.
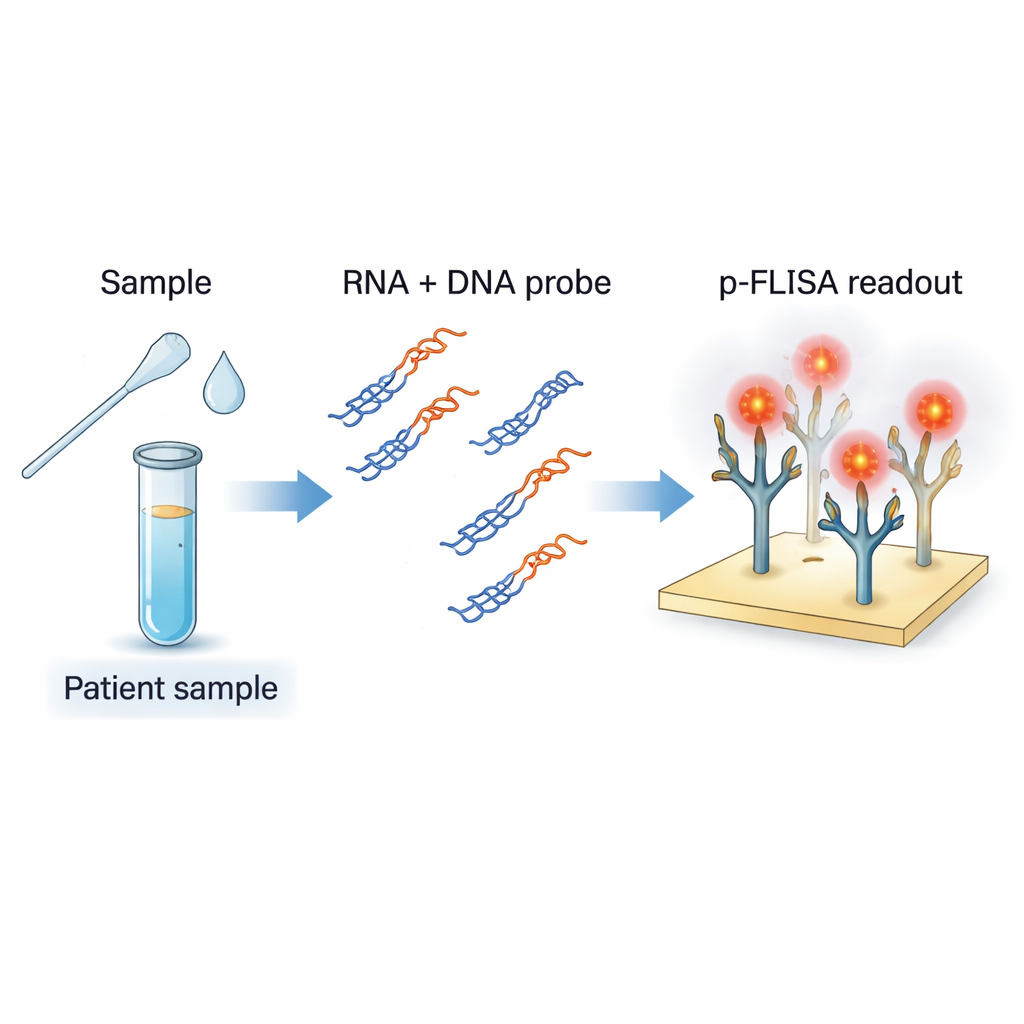
A new way to see viral genetic traces
Standard COVID-19 diagnosis uses RT-PCR, which makes many copies of fragments of viral RNA so they can be detected. While very sensitive, PCR requires complex equipment, trained staff, and time, and usually reports a "yes or no" answer rather than a precise viral load. The authors set out to build an assay that works more like a highly upgraded version of an ELISA blood test: a simple plate-based format, but tuned to detect viral RNA directly and quantify how much is present. Their method targets SARS-CoV-2 RNA in samples such as nasal swabs and saliva, but is designed to be easily adapted to other RNA viruses.
Turning RNA into a catchable target
The team uses short pieces of DNA that are designed to match specific regions of the SARS-CoV-2 genome. When mixed with RNA extracted from a patient sample and gently heated and cooled, these DNA probes pair up with any matching viral RNA to form DNA–RNA hybrids, like a small zipper with one RNA strand and one DNA strand. A special antibody called S9.6 acts as the catcher: it recognizes and binds strongly to these hybrids, but not to ordinary single- or double-stranded DNA or to unrelated RNA. By coating the bottom of a plate with S9.6, the assay selectively grabs only those hybrids that contain the virus’s genetic sequences of interest, filtering out the rest of the sample’s genetic clutter.
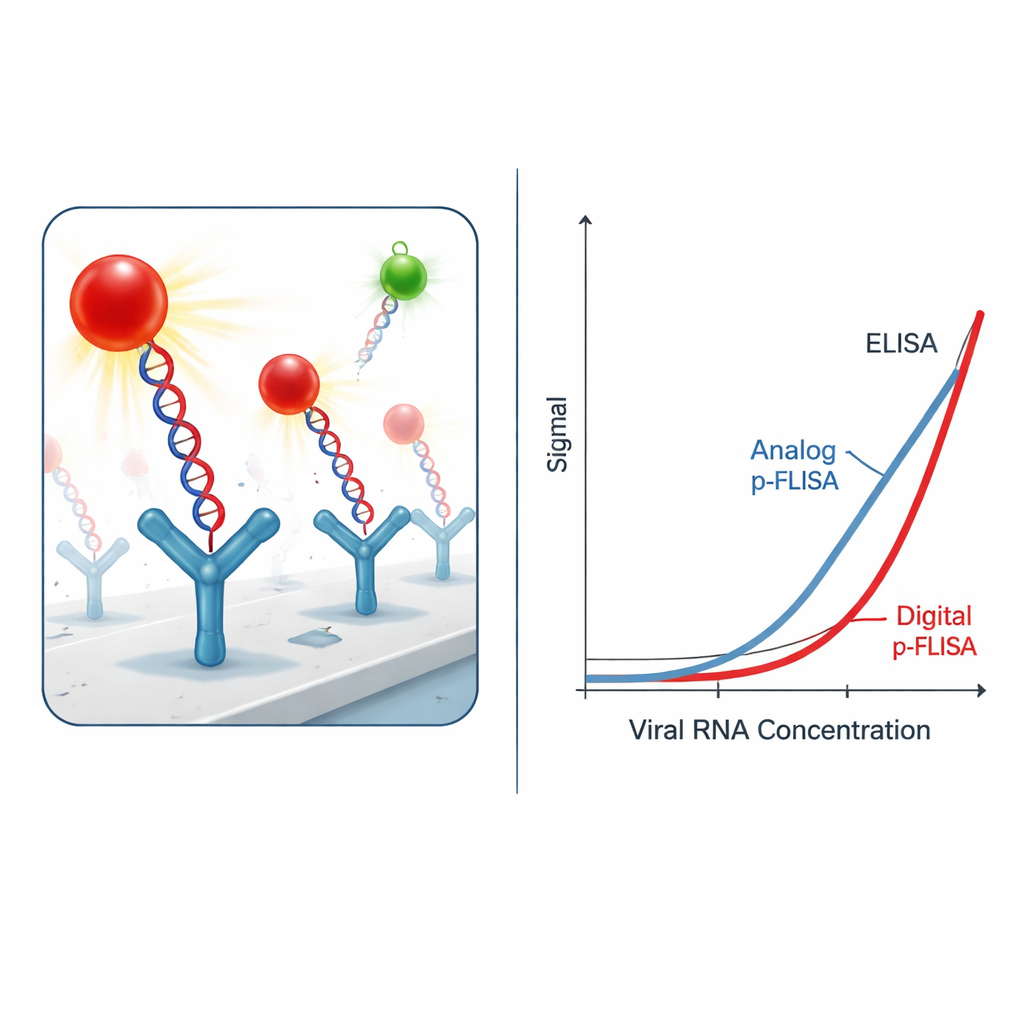
Making the signal ultrabright
Simply catching the viral hybrids is not enough; the challenge is seeing them above background noise. Instead of conventional fluorescent dyes, the researchers use "plasmonic-fluor" nanolabels—engineered nanoparticles that act like tiny antennas for light. Each label combines a metal nanorod with multiple fluorescent molecules and a coating that allows it to latch onto biotin tags on the DNA or antibody. These plasmonic labels emit light more than a thousand times brighter than standard dyes under the same conditions. In practice, that means far fewer viral hybrids are needed to produce a detectable glow, dramatically improving the test’s sensitivity and lowering the minimum viral RNA concentration it can measure.
From analog glow to digital counting
In its simplest “analog” form, the assay measures the overall brightness from each well of the plate, much like a classic fluorescent test. Even in this mode, the plasmon-enhanced system improves the detection limit and the lowest reliably quantifiable level of SARS-CoV-2 RNA by one to three orders of magnitude compared with a traditional ELISA using enzymes or standard fluorophores. The authors then push the idea further by switching to a “digital” format: rather than averaging the light over the whole well, they image the surface with a fluorescence microscope and count individual bright nanolabels using custom image analysis software. This single-particle counting approach yields another ten- to thirty-fold gain in sensitivity, leading to roughly 2,300-fold better detection limits and 460-fold better quantification limits than ELISA overall.
Putting the test to work on real samples
To see whether the method holds up outside controlled lab mixtures, the researchers tested RNA extracted from nasal swabs and saliva of patients with COVID-19, including infections from different viral variants such as alpha/beta and delta, as well as samples from people with other respiratory viruses. Their plasmon-enhanced assay detected SARS-CoV-2 RNA in all PCR-positive samples, with no signal above background in PCR-negative samples or those harboring other viruses, indicating excellent clinical sensitivity and specificity comparable to RT-PCR. Moreover, the measured RNA concentrations showed an inverse relationship with PCR cycle threshold values: samples that needed fewer PCR cycles (indicating higher viral load) had higher RNA levels by the new assay, consistent with biological expectations and suggesting that it can provide meaningful quantitative viral load information.
What this could mean for future outbreaks
For non-specialists, the key message is that this assay offers a way to read out how much virus is present without the extra copying step that makes PCR slow and equipment-heavy. By combining a selective antibody for RNA–DNA hybrids with ultrabright nanoscale light sources and digital counting, the method approaches PCR-like performance while retaining a simple, plate-based workflow. With further validation and engineering, such plasmon-enhanced assays could be adapted to many RNA targets and possibly reshaped into rapid, point-of-care formats, helping clinicians not only diagnose infections but also gauge disease stage and contagiousness from an absolute measure of viral RNA.
Citation: Liu, L., Seth, A., Liu, Y. et al. Plasmon-enhanced bioassay for amplification-free detection and quantification of SARS-CoV-2 RNA. npj Biosensing 3, 13 (2026). https://doi.org/10.1038/s44328-026-00078-x
Keywords: SARS-CoV-2 RNA detection, plasmonic nanolabels, digital immunoassay, amplification-free diagnostics, viral load quantification