Clear Sky Science · en
Label-free whole slide virtual multi-staining using dual-excitation photon absorption remote sensing microscopy
Seeing Tissues Without Destroying Them
When doctors diagnose diseases like cancer or kidney damage, they often rely on thin slices of tissue that have been soaked in chemical dyes. These stains reveal hidden structures but permanently alter or use up the sample, which can be a problem when only a tiny biopsy is available. This study introduces a way to “digitally stain” tissue using light and artificial intelligence, creating familiar-looking pathology images without adding any dye at all.
Why Traditional Stains Are a Mixed Blessing
Chemical stains such as hematoxylin and eosin, or special dyes for collagen, carbohydrates, and kidney structures, are the workhorses of modern pathology. They make transparent tissue visible and are indispensable for diagnosing cancer, infections, and organ damage. But these stains are destructive: the same section usually cannot be re-stained or used for advanced tests, and multiple stains quickly consume precious biopsy material. Each stain also requires carefully controlled lab work, trained staff, and can add hours or days to the time before a diagnosis is ready.
Light-Based Imaging That Reads the Tissue Itself
The researchers used a specialized microscope called Photon Absorption Remote Sensing (PARS), which reads how tissue molecules absorb and release energy from ultraviolet light. In this work they combined two ultraviolet colors, one at a shorter wavelength and one slightly longer, and fired them in an interlaced pattern at the same spot on the tissue. Each pulse produces both heat-related signals and faint glow-like emissions, giving four distinct channels of information from the same location. One wavelength is especially sensitive to DNA in cell nuclei, while the other highlights collagen, elastin, red blood cells, and dark pigments like melanin. Together they map out nuclei, supporting tissue, blood, and pigment in ways that resemble and even extend what pathologists see with traditional stains. 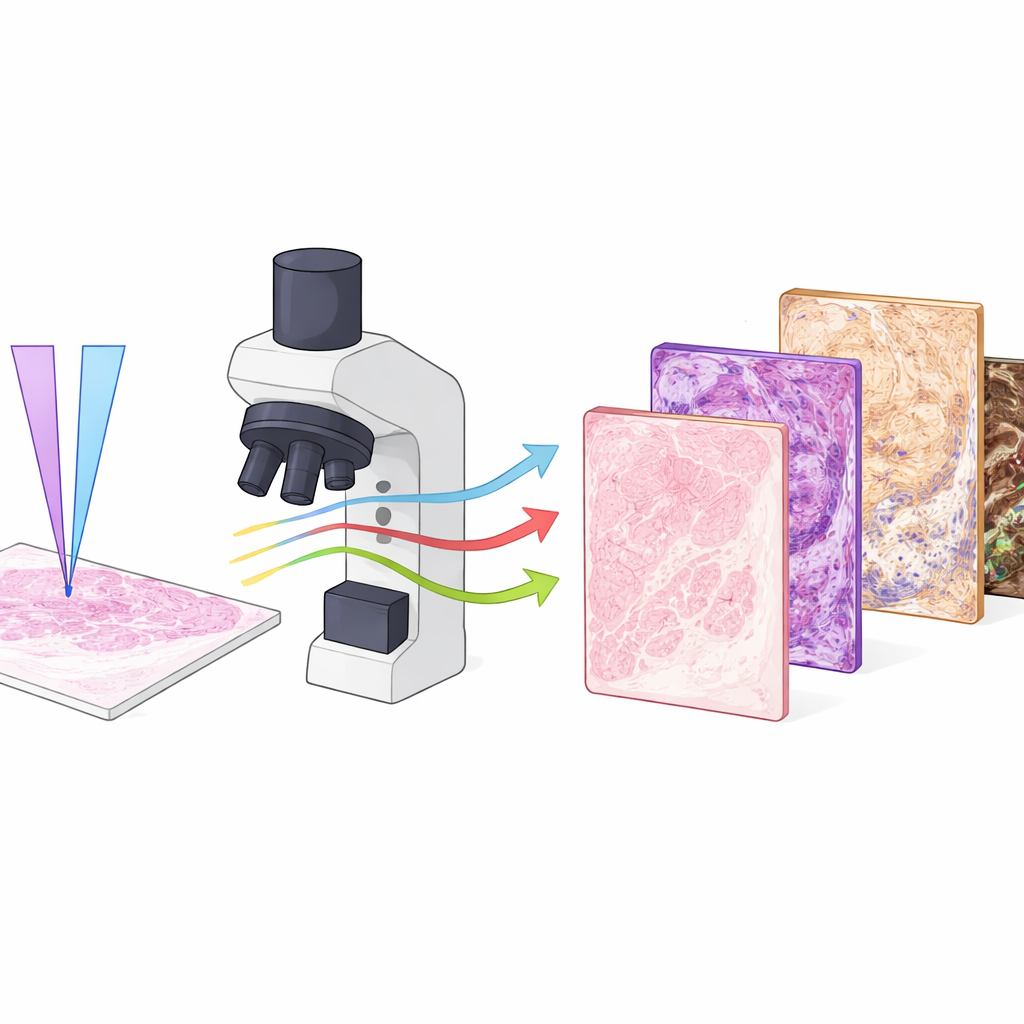
Teaching Computers to Paint Virtual Stains
Collecting rich optical signals is only half the story; the other half is turning them into images that look like standard stained slides. To do this, the team used a deep-learning framework called RegGAN. They first imaged unstained tissue with PARS, then chemically stained the very same slide and scanned it with a regular brightfield scanner. After carefully aligning these paired images, they trained neural networks to transform the multi-channel PARS images into versions that mimic specific stains, including routine hematoxylin and eosin as well as Masson’s trichrome, PAS, and Jones methenamine silver. Separate models were trained for each stain, so that a single label‑free input slide could later be “virtually re-stained” several different ways on demand.
What the Virtual Slides Reveal
Across human and mouse tissues—including kidney cancers, melanoma, fungal skin infections, and normal organs—the virtual stains closely tracked their chemical counterparts. Tumor borders, nuclear shapes, collagen-rich scar tissue, red blood cells, fungal filaments, and fine kidney structures all appeared with high fidelity when both ultraviolet wavelengths were used together. Quantitative image-quality measures confirmed that combining the two excitations outperformed using either one alone, particularly for structures like collagen, blood cells, and fungal elements that rely on the added contrast from the longer wavelength. In a small masked study, three experienced pathologists rated both real and virtual images mainly as good or excellent for visual diagnostic quality, and they were unable to reliably tell which images were chemically stained and which were virtual.
Strengths, Limits, and Future Potential
While promising, the method is not yet ready to replace routine slide scanners. The current PARS system is slow, taking hours to cover the area that a clinical scanner can capture in minutes, and all data came from one imaging setup and one staining laboratory. The evaluation focused on visual similarity and select measurable features, rather than full clinical decision-making across many patients and centers. Nonetheless, the approach offers a unique advantage: because the label‑free imaging does not damage the tissue, the same slide can later be stained with traditional dyes or used for molecular tests, and multiple virtual stains can be generated from a single scan. 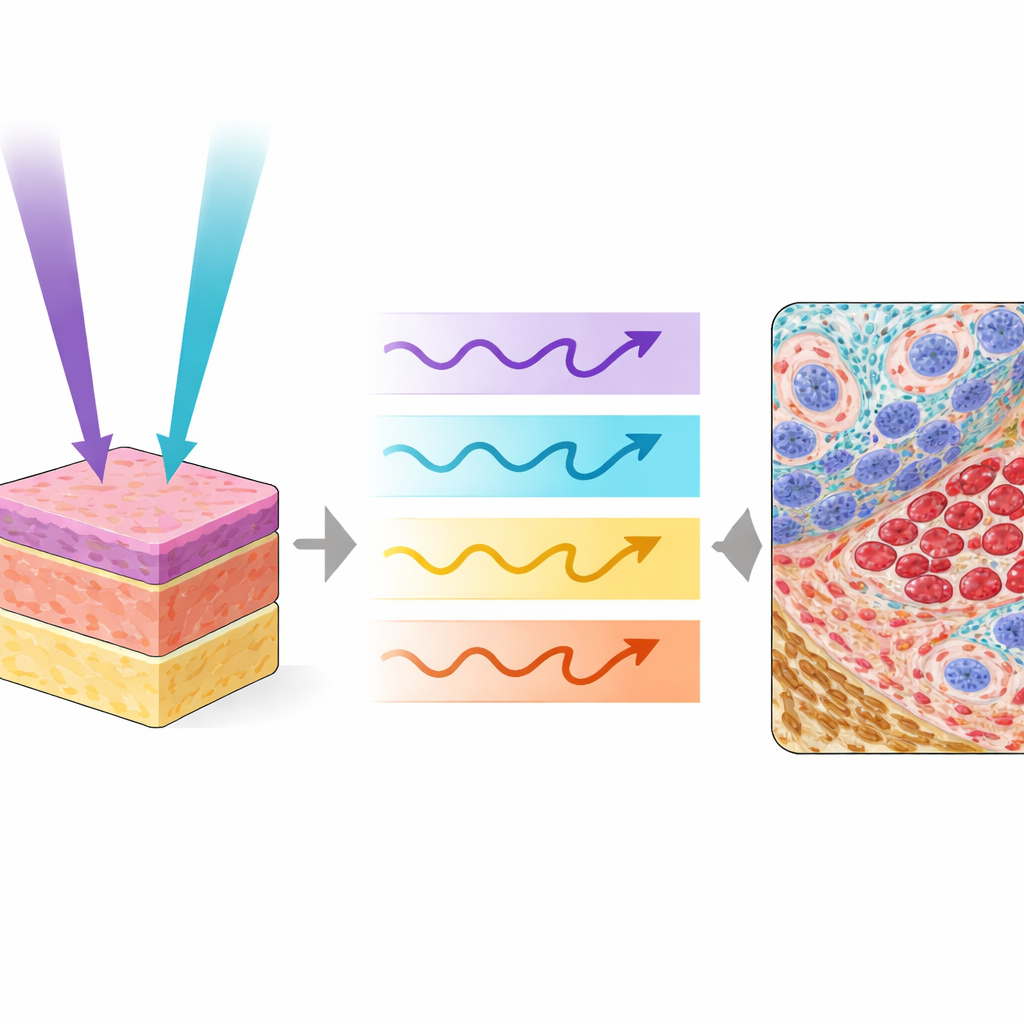
What This Means for Patients and Doctors
In simple terms, this study shows that it is possible to “read” tissue using light alone and then use artificial intelligence to recreate the familiar colors and patterns that pathologists trust, including several different stains from one section. The dual‑color PARS system provides enough information to virtually highlight nuclei, supporting tissue, blood, pigment, and specialized kidney structures without touching a drop of dye. With faster hardware and larger, multi‑center studies, this technology could become a powerful companion to standard pathology, conserving precious biopsies and offering pathologists a richer, non-destructive view of disease.
Citation: Tweel, J.E.D., Ecclestone, B.R., Tummon Simmons, J.A. et al. Label-free whole slide virtual multi-staining using dual-excitation photon absorption remote sensing microscopy. npj Imaging 4, 22 (2026). https://doi.org/10.1038/s44303-026-00154-x
Keywords: virtual staining, label-free microscopy, digital pathology, ultraviolet imaging, deep learning in histology