Clear Sky Science · en
High-resolution dynamic full-field optical coherence microscopy: illuminating intracellular activity in deep tissue
Seeing Living Cells in Action, Without Any Dyes
Much of what we know about cells inside the body comes from stains and fluorescent dyes that can alter or even harm the tissue being studied. This paper introduces an advanced microscope that can watch the natural activity of cells deep inside organs like the liver and intestine without adding any labels at all. It turns tiny internal motions into vivid, almost fluorescent-looking pictures, opening a window onto living tissue that could one day help doctors diagnose disease in real time.
A New Way to Watch Cells Move
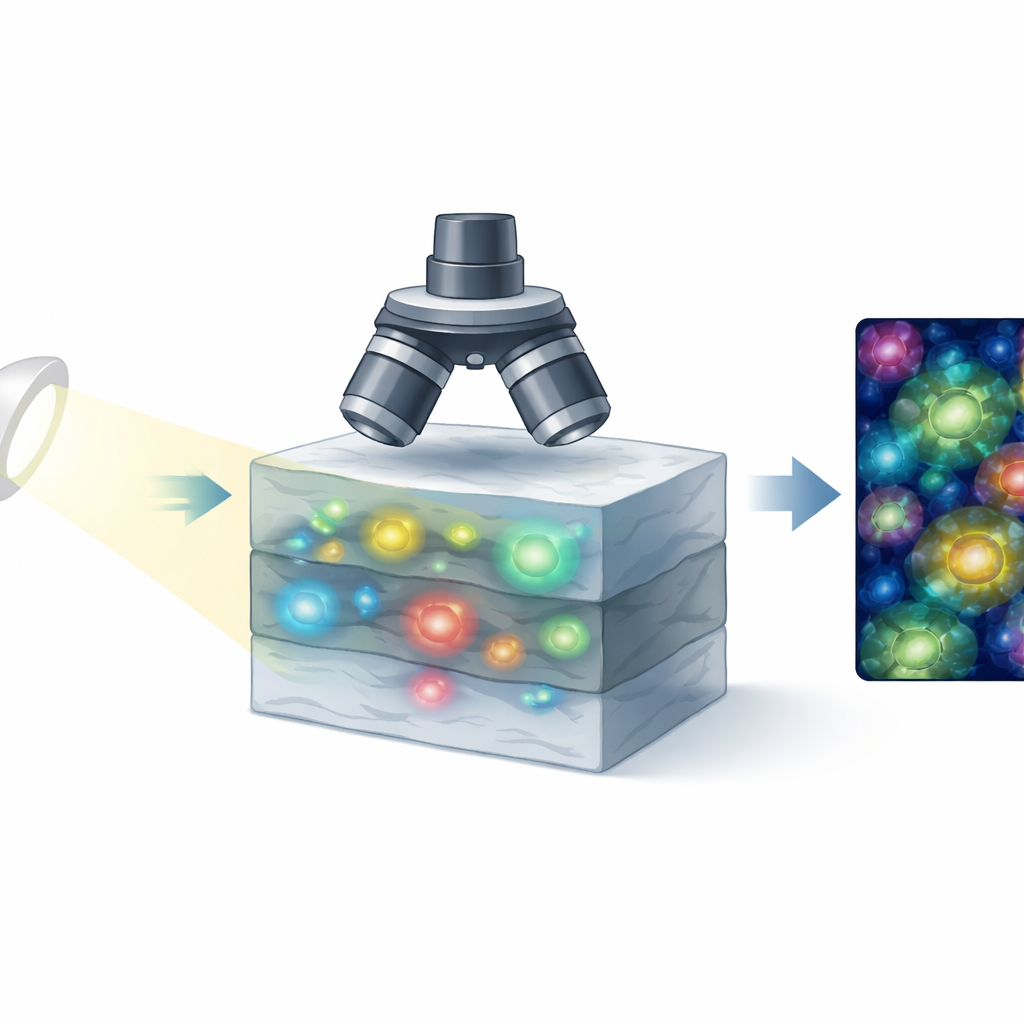
The technique at the heart of this work is called dynamic full-field optical coherence microscopy, a form of light-based imaging that detects how light echoes back from inside tissue. Instead of scanning point by point, it records an entire plane of tissue at once with a camera, then repeats this rapidly. The key idea is that living cells are never truly still: their internal parts shift, vibrate, and reorganize as they use energy and carry out their tasks. These microscopic motions subtly change the light signal over time. By carefully analyzing how the signal fluctuates at each point, the system builds images in which active structures stand out, much like they would in a fluorescent microscope, but without any added dyes.
Pushing Deep Into Cloudy, Real-World Tissues
Imaging deep inside real organs is difficult because tissue scatters and scrambles light, and microscopes usually face a trade-off between sharpness and depth. The authors redesigned the dynamic microscope to overcome these limits. They used powerful 100× oil-immersion lenses, which collect and focus light very tightly, and combined them with a special white light source driven by a laser. This source is both extremely bright and spatially incoherent, meaning it avoids the grainy speckle patterns that plague many laser-based systems. With this combination, the microscope achieves detail down to a few hundred nanometers—small enough to resolve fine cell structures—while still seeing up to about 120 micrometers deep into strongly scattering tissues such as liver. A smart, motorized reference arm continuously adjusts the optical path as the focus moves deeper, keeping the image contrast high throughout the volume.
Revealing Hidden Architecture in the Liver
To test the system, the researchers imaged fresh mouse liver. Standard versions of the technique produced rather plain views: densely packed liver cells with vague borders and dark spots where nuclei sit. When they switched to dynamic imaging and analyzed the temporal fluctuations, the pictures transformed. Cell boundaries became crisp; filament-like networks consistent with mitochondrial activity appeared within many liver cells; and sinusoids—the tiny blood channels that weave between plates of cells—lit up across a wide range of fluctuation speeds. In zoomed-in views, individual red blood cells and small, mobile elements likely corresponding to platelets or immune cells could be discerned inside these channels, even many layers deep. The method also captured differences in how quickly various parts of the tissue fluctuated, by mapping slow, intermediate, and fast motions into different colors.
Peering Into the Gut’s Microscopic Landscape
The team then turned to the small intestine, imaging it from both the inner (mucosal) side and the outer (serosal) side. From the mucosal surface, they could see the finger-like villi that line the gut, with enterocytes forming a tight mosaic at the tips. Nuclei and structures consistent with microvilli were visible on the cell surfaces, along with likely goblet cells, which secrete mucus, and a variety of highly active cells in the supporting tissue beneath. From the serosal side, the microscope captured intricate nerve networks known as the myenteric and submucosal plexuses, as well as blood vessels threaded among them. Remarkably, it produced the first optical coherence images of Paneth cells at the base of intestinal crypts—specialized defenders of the gut—along with the surrounding crypt cells and possible supporting stromal cells, all distinguished by their dynamic signatures.
Why This Matters for Future Medicine
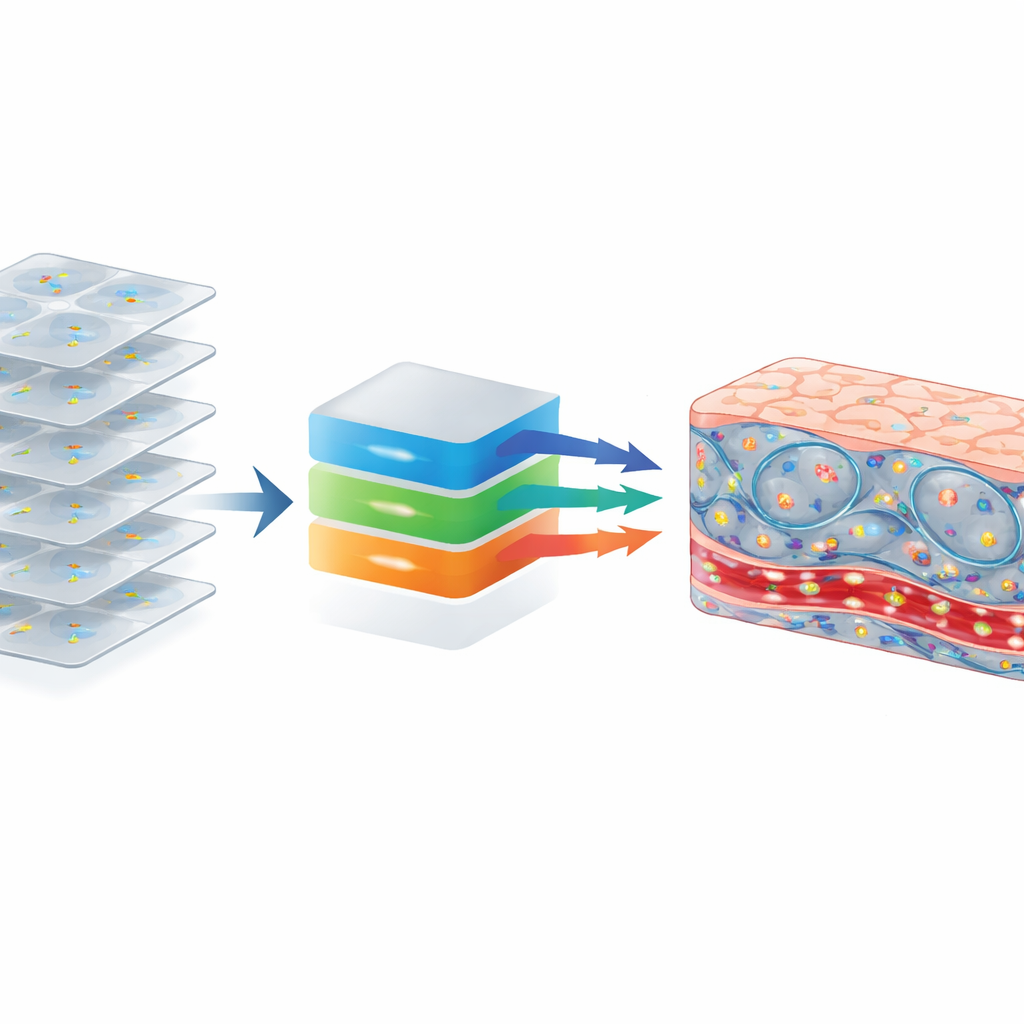
By combining high resolution, extended depth, and motion-based contrast, this new system shows that it is possible to obtain rich, fluorescence-like images of living tissue without any dyes or genetic modifications. It exposes the fine structure and activity of cells in complex organs such as liver and intestine, revealing blood flow, immune cell candidates, nerve networks, and compartmentalized activity within individual cells. With further engineering to handle motion and access in live animals or patients, the same approach could be adapted for in vivo use. That would offer clinicians a rapid, label-free way to see how cells are behaving in real time during surgery or diagnosis, potentially enabling earlier detection of disease and more precise, personalized treatments.
Citation: Tarvydas, E., Trečiokaitė, A. & Auksorius, E. High-resolution dynamic full-field optical coherence microscopy: illuminating intracellular activity in deep tissue. npj Imaging 4, 21 (2026). https://doi.org/10.1038/s44303-026-00153-y
Keywords: label-free microscopy, optical coherence imaging, liver tissue imaging, intestinal microstructure, cellular dynamics