Clear Sky Science · en
Correlative multimodal imaging for microscale spatial mapping of collagen-gene activity interactions in human tissues
Why looking inside muscles matters
Diseases that slowly scar our organs, like Duchenne muscular dystrophy (DMD), unfold at scales too small for the naked eye. Gene activity happens inside individual cells, while the supporting tissue around them is made of tiny protein fibers. Until now, scientists have rarely been able to see these two worlds together in the very same slice of human tissue. This study introduces a way to do exactly that, revealing how messages from genes and the body’s structural scaffolding interact at the microscale.
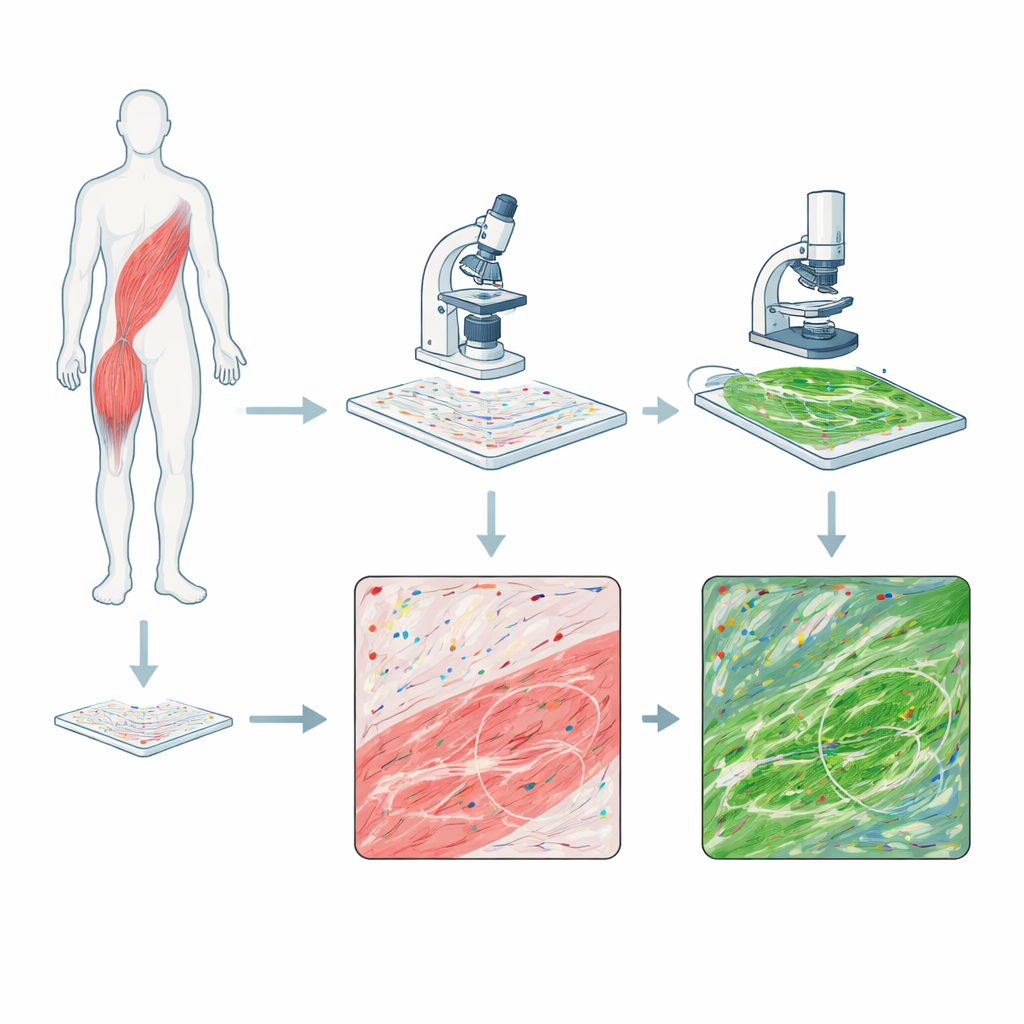
Two cameras on the same piece of tissue
The researchers combined two powerful imaging methods on single sections of human skeletal muscle biopsies from people with DMD and from a healthy volunteer. One method, called RNAscope, stains individual RNA molecules—the working copies of genes—as colored dots, showing where specific genetic messages are present. The second method, multiphoton microscopy, uses laser light to make collagen fibers in the tissue glow without any added dye, capturing the architecture of the connective tissue that thickens during scarring and fibrosis. By imaging first the RNA signals and then the collagen signals on the exact same slide, they could align the two pictures so that each dot of gene activity could be compared with the surrounding fiber network, pixel by pixel.
What the separate views already tell us
Looking at each imaging method on its own already highlighted clear differences between diseased and healthy muscle. RNAscope showed that muscles from the DMD patients contained fewer dystrophin transcripts—the RNA messages needed to make the missing dystrophin protein—than the healthy sample. The remaining signals also differed in size depending on which part of the gene was targeted, hinting at differences in how the long dystrophin message is processed. Meanwhile, multiphoton images showed that DMD muscles were packed with long, dense collagen fibers, a visual sign of fibrotic scarring, whereas the healthy muscle had fewer, more evenly distributed fibers. These findings match clinical experience: in DMD, muscle tissue is gradually replaced by fat and scar.
Zooming out: broad patterns across the tissue
After digitally registering the two kinds of images, the team divided each tissue section into a grid, treating each square as a tiny neighborhood. In every square they measured both collagen features—fiber length, orientation, and how winding or straight the fibers were—and the local density of dystrophin RNA dots. This “heat map” view revealed patchy, region‑to‑region variation in both gene activity and collagen structure within DMD muscle. However, when these measures were compared using statistical tests, transcript abundance and collagen properties showed almost no consistent relationship at this coarse scale. In other words, when averaging over relatively large regions, areas rich in dystrophin RNA were not reliably more or less fibrotic than areas with fewer RNA signals.
Zooming in: relationships at the cellular scale
The most revealing results emerged when the scientists focused on neighborhoods measured in only a few tens of micrometers—roughly the size of individual muscle fibers and their immediate surroundings. For every RNA dot, they drew a series of growing circles and asked how many collagen fibers lay inside, and how long those fibers were. Across both DMD and healthy tissues, collagen fibers were longest right next to the dystrophin RNA signals and became progressively shorter farther away. In the most severely scarred samples, transcript dots almost always had collagen nearby, even at very small distances, while in the healthy sample many RNA dots sat in regions with little or no nearby collagen. Control analyses using randomly placed points and slightly altered image‑processing settings showed that this pattern could not be explained simply by chance or by quirks of the software.
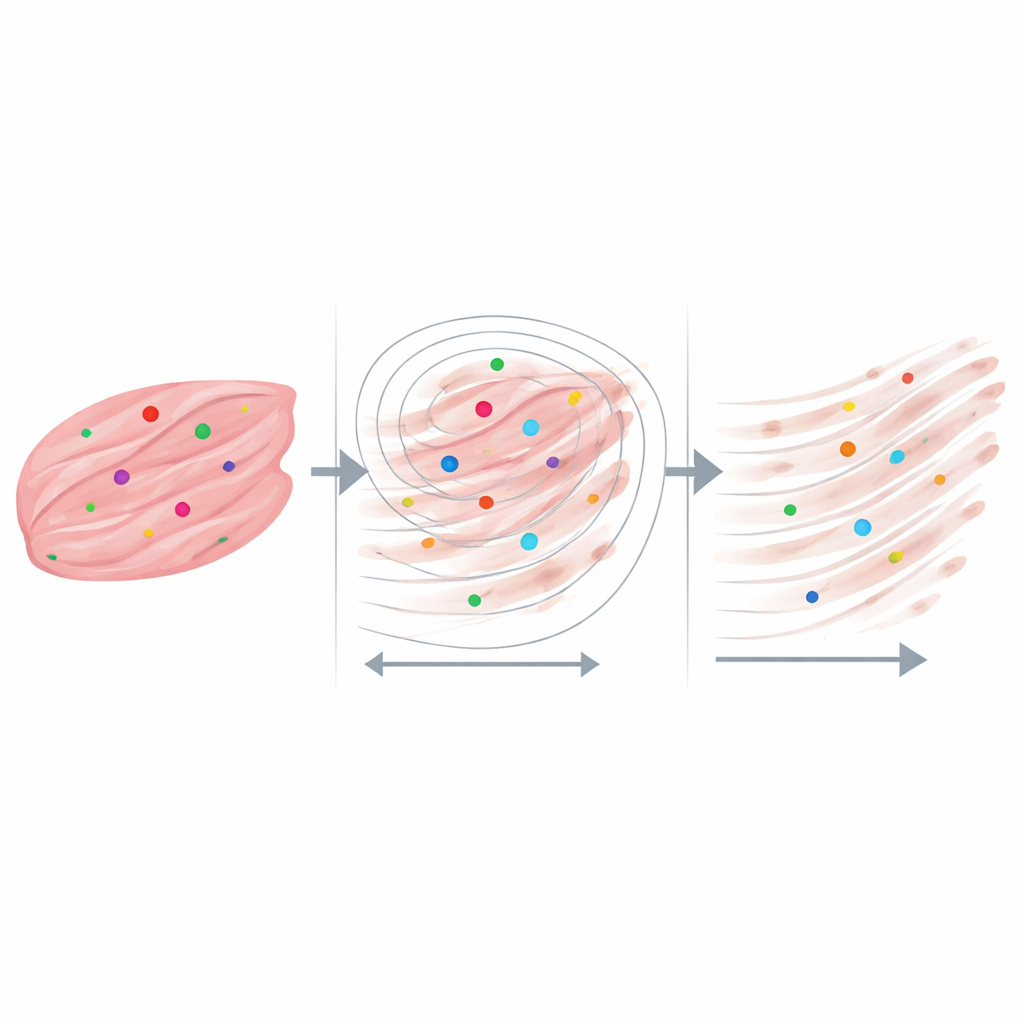
What this could mean for muscle disease and beyond
These findings suggest that important links between gene messages and tissue structure can be hidden when looking only at large averages, but become visible when examined at the cellular level. In this proof‑of‑concept study, regions enriched in specific dystrophin transcripts tended to sit near longer, denser collagen fibers, hinting that local gene activity and fibrotic remodeling may influence each other within tiny microenvironments of muscle. The authors emphasize that their patient group is small and the work is not yet a clinical test. Instead, they present a flexible platform that can be extended to other genes, tissue types, and imaging methods. By tying together where genes are active and how the tissue scaffold is built, this approach opens the door to discovering new spatial biomarkers of fibrosis, regeneration, and responses to emerging RNA‑based therapies.
Citation: Scodellaro, R., Mietto, M., Ferlini, A. et al. Correlative multimodal imaging for microscale spatial mapping of collagen-gene activity interactions in human tissues. npj Imaging 4, 17 (2026). https://doi.org/10.1038/s44303-026-00149-8
Keywords: multimodal imaging, Duchenne muscular dystrophy, collagen fibrosis, spatial gene expression, skeletal muscle biopsy