Clear Sky Science · en
Genomic characterization and sub-clustering of Escherichia coli clonal complex 38 reveal host associated genetic markers
Why this matters for everyday life
Antibiotic-resistant infections are no longer a distant hospital problem—they increasingly touch food, animals, travel, and community life. This study focuses on a particular group of Escherichia coli bacteria, called clonal complex 38 (CC38), that can cause serious bloodstream and urinary infections and often resist our most important antibiotics. By tracing where these bacteria come from and how they move between humans, animals, and the environment, the researchers reveal clues that could help health authorities stop dangerous strains before they spread widely.
A closer look at a troublesome bacterial family
Not all E. coli are harmful, but some lineages are repeat offenders in severe disease. CC38 has recently emerged as one of these problematic groups, second only to a well-known high-risk lineage (ST131) in Danish bloodstream infections. The team examined 242 resistant E. coli CC38 from Danish patients, 83 from food and farm animals, and more than 2,300 related genomes collected worldwide. They used whole-genome sequencing—essentially reading each bacterium’s DNA—to map how different branches of this family are related, what resistance genes they carry, and which hosts they tend to live in.
Following the trail from hospitals to farms and food
When the researchers compared E. coli CC38 from Danish patients with those from poultry, livestock, and food, they found two main Danish groups. One group was made up mostly of human infections, while the other contained bacteria from poultry, other farm animals, and food products. Importantly, there were no near-identical DNA matches between human and animal isolates, suggesting there were no clear-cut foodborne outbreaks in Denmark during the study period. However, statistical models that look at accessory DNA segments indicated that some human-associated subgroups probably have poultry origins, hinting at past or indirect spillover from chickens into people.
A global family tree with two big branches
Expanding to a worldwide dataset of 2,638 genomes, the team built a global “family tree” for CC38. Two major branches stood out. One was strongly tied to poultry and carried moderate levels of drug resistance along with many traits linked to serious disease. The other was dominated by human-associated subgroups that differed in resistance, disease potential, and preferred hosts. Some subgroups were clearly human-focused and heavily resistant, while others showed mixed patterns, turning up in humans, poultry, wildlife, water, and companion animals. This patchwork reflects CC38’s ability to adapt to different environments and host species, complicating efforts to control it.
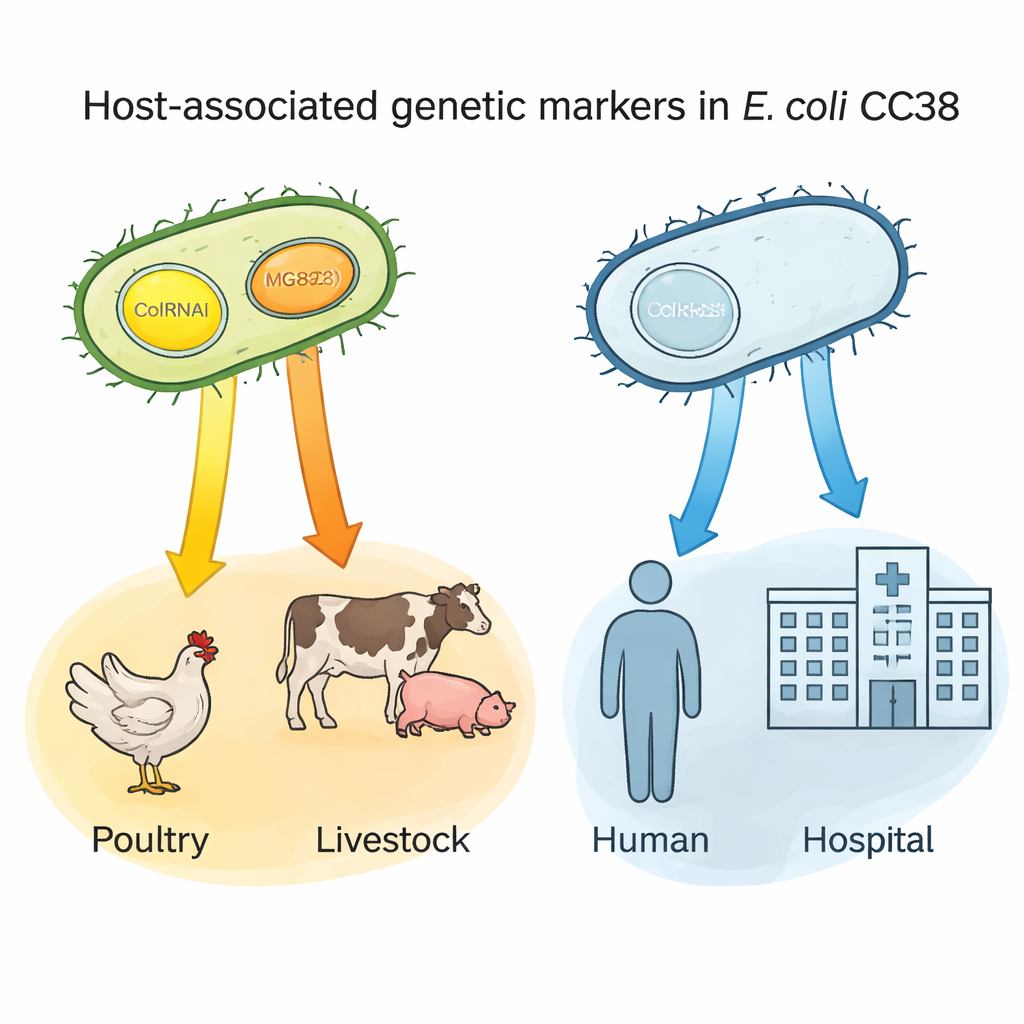
Small DNA circles as host “name tags”
A key discovery involved two tiny pieces of DNA—plasmids called ColRNAI and Col(MG828)—that can move between bacteria. These plasmids were common in CC38 from poultry and other farm animals but rare in the human-centered subgroups. Using statistical analysis, the authors showed that carrying either plasmid strongly predicted a poultry origin; carrying both together was especially characteristic of lineages linked to livestock and certain food sources. Because these plasmids also often carry resistance traits, they may act as genetic “name tags” that flag bacteria likely to have come from animal reservoirs, and may help track how resistant strains enter the food chain and reach people.
What this means for protecting health
For a non-specialist, the main message is that harmful, drug-resistant E. coli are not confined to hospitals or patients: they form interconnected populations spanning humans, poultry, livestock, wildlife, food, and the environment. This study shows that within one important lineage, CC38, some branches are strongly tied to animals and others to humans, and that specific mobile DNA elements can help reveal where a strain likely came from. Using these genetic markers in routine surveillance could give early warning when animal-associated resistant bacteria begin to appear in people. In turn, this supports a “One Health” approach—treating human, animal, and environmental health as parts of the same system—to design smarter, targeted strategies for preventing infections and slowing the spread of antibiotic resistance.
Citation: Roer, L., Rasmussen, A., Hansen, F. et al. Genomic characterization and sub-clustering of Escherichia coli clonal complex 38 reveal host associated genetic markers. Commun Med 6, 126 (2026). https://doi.org/10.1038/s43856-026-01402-2
Keywords: antimicrobial resistance, Escherichia coli, zoonotic transmission, poultry and livestock, genomic surveillance