Clear Sky Science · en
Prediction of outcome from spatial Protein profiling of triple-negative breast cancers
Why the layout of cancer cells matters
When doctors look at an aggressive breast cancer under the microscope, they see a crowded landscape of tumor cells, immune cells, and supporting tissue. But until recently, most tests have focused on how much of a given molecule is present, not where it sits. This study shows that the physical arrangement of proteins and cells inside triple-negative breast cancers can forecast how patients fare, and introduces a new way to read those spatial patterns directly from high-detail images.
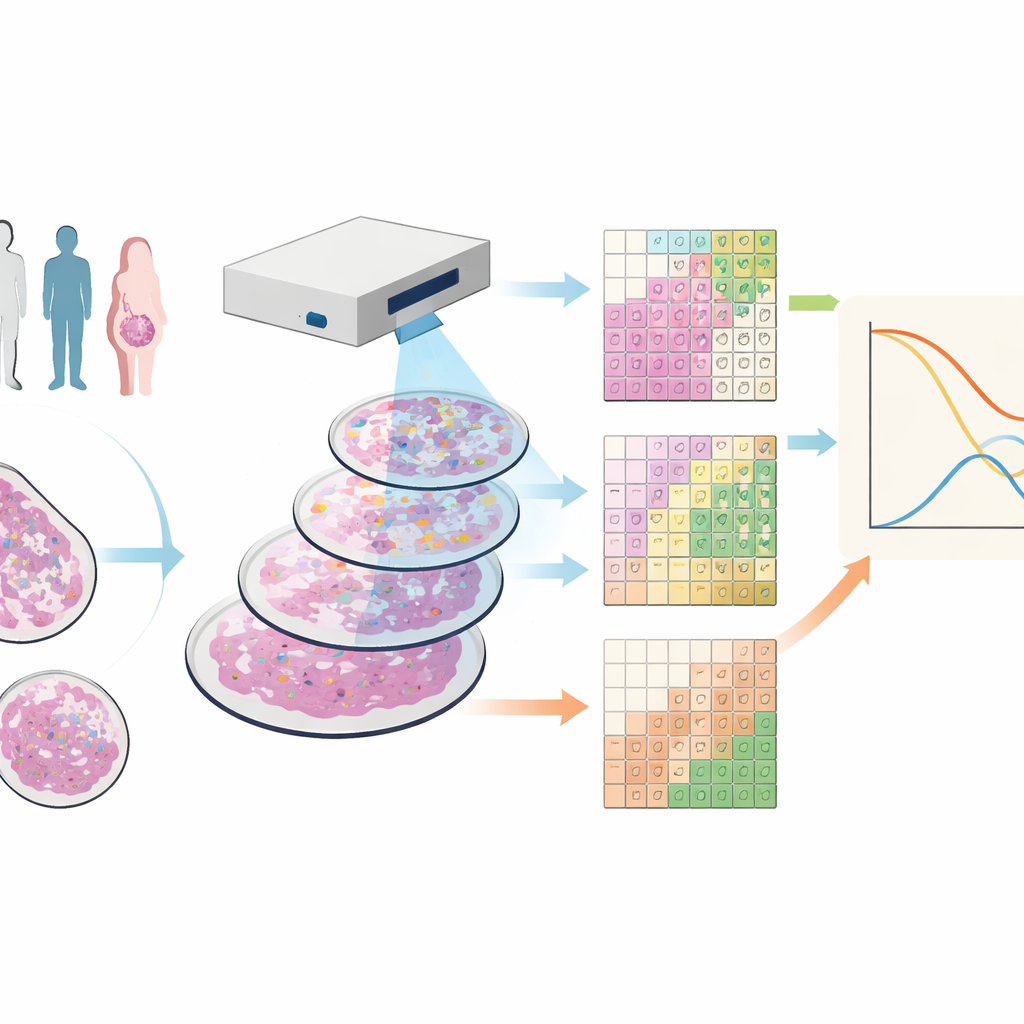
Seeing cancer tissue in many colors
The researchers worked with tissue samples from 88 people with triple-negative breast cancer, a form of the disease that lacks common drug targets and often has a poor outlook. They used a technique called imaging mass cytometry, which stains each sample with dozens of metal-tagged antibodies. Under a specialized scanner, this produces images where each point in the tissue has a detailed fingerprint of many different proteins, revealing tumor cells, various immune cells, blood vessels, and structural fibers all at once.
Breaking the image into tiles instead of cells
Most current tools try to draw boundaries around every cell and then classify them by type, a process that can be error-prone and slow, especially in thin tissue slices where cells are only partially captured. The team instead created SparTile, a method that skips drawing individual cell outlines. SparTile chops each image into many small overlapping squares, or tiles, and uses mathematical techniques to learn recurring patterns of protein combinations within those tiles. It first groups tiles into broad regions—tumor, supporting tissue (stroma), or immune-rich zones—and then further divides each region into more specific “microenvironments,” each with its own mix of proteins and neighboring cell types.
Hidden neighborhoods linked to patient outcome
With these microenvironments mapped, the scientists measured how much of each pattern appeared in every patient’s sample and compared this with long-term survival. Several striking links emerged. One tumor-centered microenvironment rich in a protein called MX1 and markers of myeloid immune cells was associated with a much higher risk of death. Another tumor pattern marked by vimentin—a sign that tumor cells have shifted toward a more mobile, invasive state—also tracked strongly with poor survival. In contrast, areas dominated by certain T cells were linked to better outcomes, while B cell–rich regions showed weaker effects, possibly because they were rare in the small tissue cores analyzed.
Distance between tumor and immune cells as a warning sign
Beyond the presence of particular neighborhoods, their spatial relationships also mattered. The team asked how closely tumor regions and myeloid immune regions sat next to each other by computing a statistical measure of how much their protein signals overlapped in space. Patients whose tumors had myeloid-rich areas nestled closely among tumor cells tended to do worse than those where the two populations were more separated. This “distance” measure remained a strong predictor of risk even after accounting for standard clinical factors, and its importance held up when tested on two independent breast cancer image datasets from other research groups.
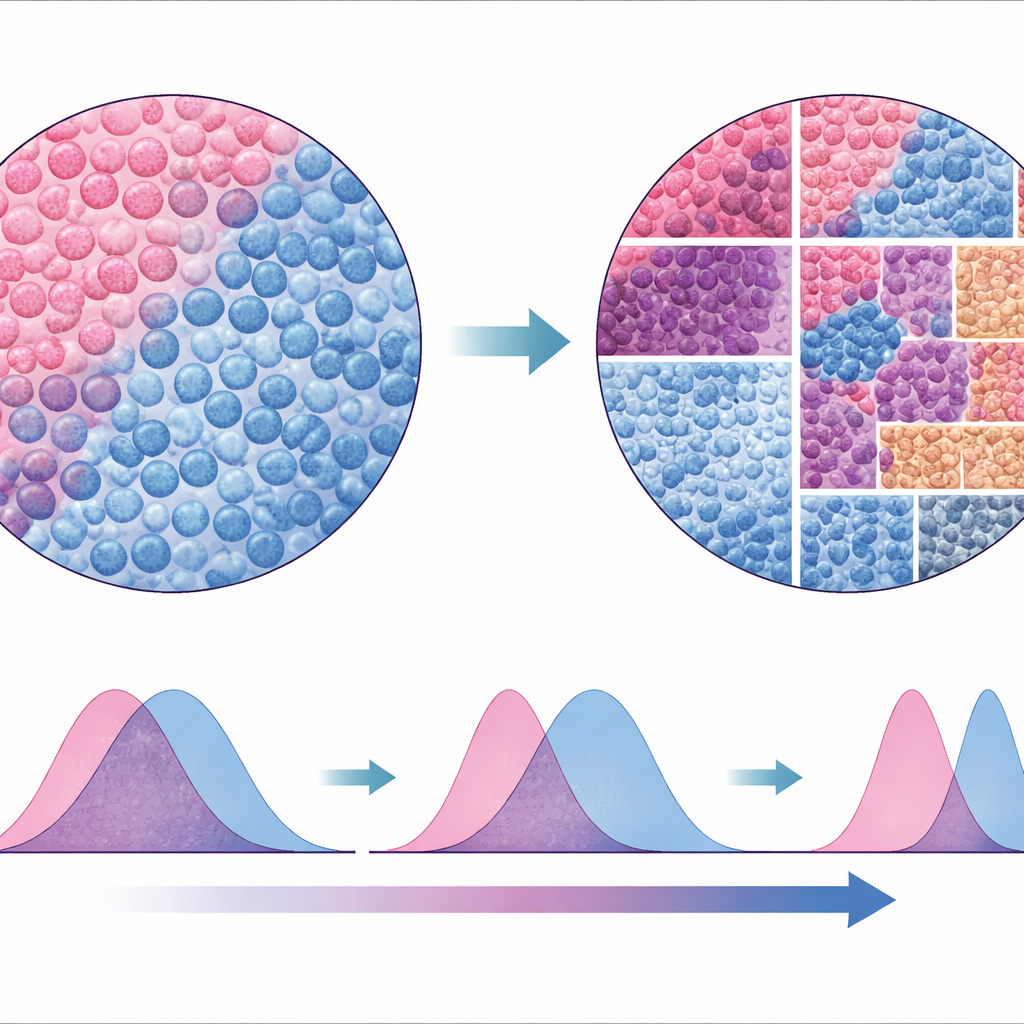
A new way to read the tumor landscape
To test how robust their approach was, the authors compared SparTile to traditional methods that rely on segmenting and classifying individual cells and then building neighborhoods around them. While the cell-based analyses did recover some of the same patterns, they were less consistent across datasets and less reliable at flagging certain risky microenvironments, such as those involving MX1 and myeloid cells. Because SparTile works directly on the raw protein images, it avoids many of the assumptions and technical pitfalls of cell-by-cell approaches, yet it can still be combined with them later when detailed cell types are needed for planning experiments or therapies.
What this means for patients and future care
This work shows that not only the ingredients, but the layout of a tumor matters for predicting how a patient with triple-negative breast cancer will do. By converting complex, multi-protein images into interpretable spatial patterns and simple distance measures, SparTile offers a way to spot high-risk patients from small tissue samples and to uncover biological arrangements that might be targeted by future treatments. While more validation is needed before it can guide routine care, the approach points toward a future where the “map” of a tumor becomes as important as its molecular parts list.
Citation: Foroughi pour, A., Wu, TC., Noorbakhsh, J. et al. Prediction of outcome from spatial Protein profiling of triple-negative breast cancers. Commun Med 6, 133 (2026). https://doi.org/10.1038/s43856-026-01400-4
Keywords: triple-negative breast cancer, tumor microenvironment, spatial proteomics, imaging mass cytometry, prognostic biomarkers