Clear Sky Science · en
IdentifiHR predicts homologous recombination deficiency in high-grade serous ovarian carcinoma using gene expression
Why this research matters for ovarian cancer patients
For people with high-grade serous ovarian cancer, one of the deadliest forms of ovarian cancer, treatment choices can be a matter of life or death. About half of these tumors have a weakness in how they repair damaged DNA, which makes them especially sensitive to certain drugs called PARP inhibitors. The challenge is figuring out, for each patient, whether their tumor has this weakness. This study introduces IdentifiHR, a new tool that reads patterns of gene activity, rather than DNA mutations alone, to predict which tumors have faulty DNA repair and could benefit most from these targeted treatments.
From DNA scars to gene activity patterns
When a cell loses a major repair pathway called homologous recombination, it begins to patch its DNA using more error-prone methods. Over time this leaves a characteristic pattern of “scars” across the genome—missing regions, extra copies, and broken chromosome segments. Existing clinical tests look for these scars directly in the DNA or for specific mutations in key genes such as BRCA1 and BRCA2. While powerful, these tests require extensive DNA sequencing and do not always capture the tumor’s current repair status. The authors asked whether a different layer of biology—the pattern of genes turned on or off in the tumor—could act as a live readout of this damage and be used to classify tumors as repair-deficient or repair-proficient.
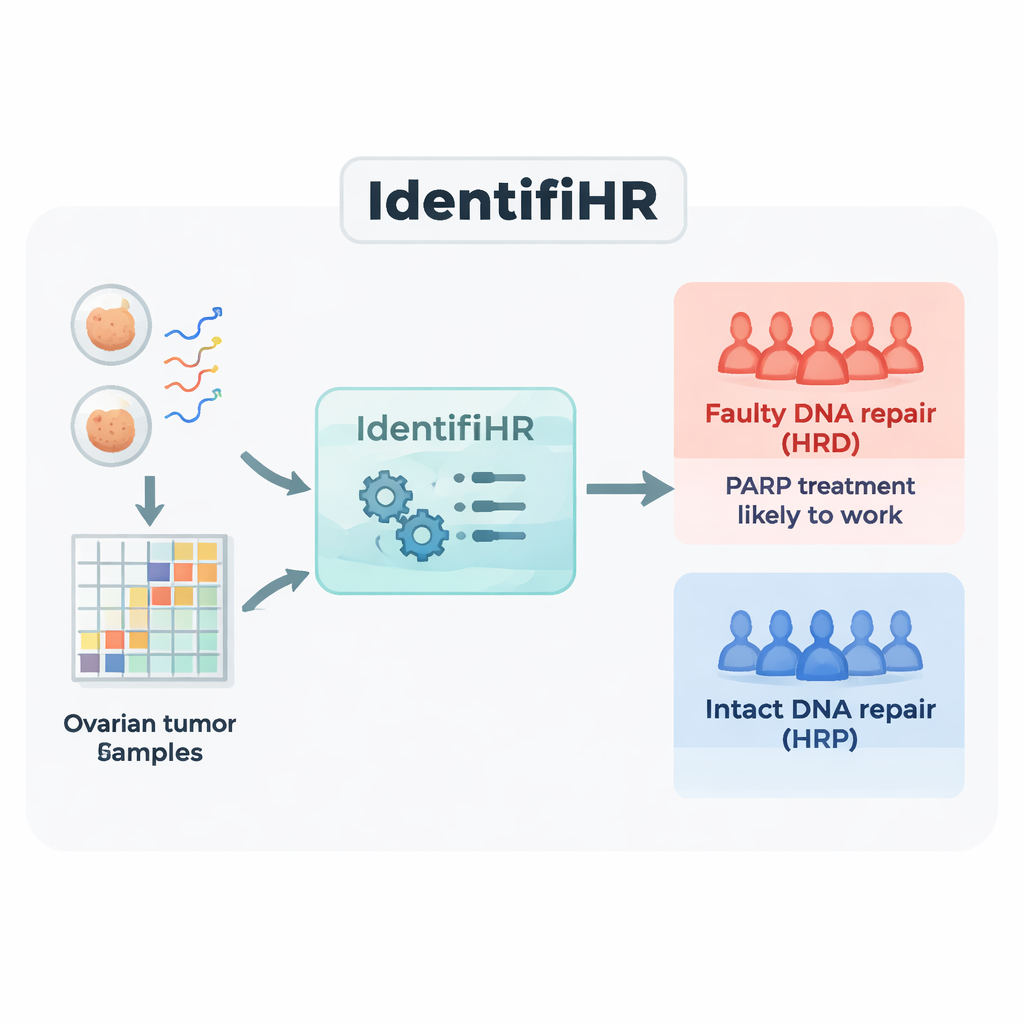
Building a gene-based predictor, IdentifiHR
The team started with RNA sequencing data from 361 ovarian tumors in a large public resource, The Cancer Genome Atlas. RNA sequencing measures which genes are active, and to what extent, in each sample. They divided the tumors into a training group and a test group, labeling each case as either repair-deficient (HRD) or repair-proficient (HRP) using the current DNA-based standard that combines several measures of genomic scarring. In the training tumors, they identified 2,604 genes whose activity consistently differed between HRD and HRP cancers. Many of these genes sat in regions of the genome already known to be repeatedly gained or lost in repair-defective tumors, showing that the gene activity signal was echoing the underlying DNA damage.
A 209‑gene signature that tracks repair status
Next, the researchers used a machine-learning approach known as penalized logistic regression to compress this list of 2,604 genes down to the most informative set. The resulting model, which they named IdentifiHR, relies on the activity of just 209 genes to estimate how likely a tumor is to be repair-deficient. Interestingly, only one of these genes is a classic DNA-repair gene; most are ordinary genes whose activity is altered because of broader changes in chromosome structure. IdentifiHR does not simply output a yes-or-no label—it produces a probability score that tracks smoothly with the underlying DNA-based damage score, reflecting the idea that repair deficiency exists on a spectrum rather than as a strict on/off state.
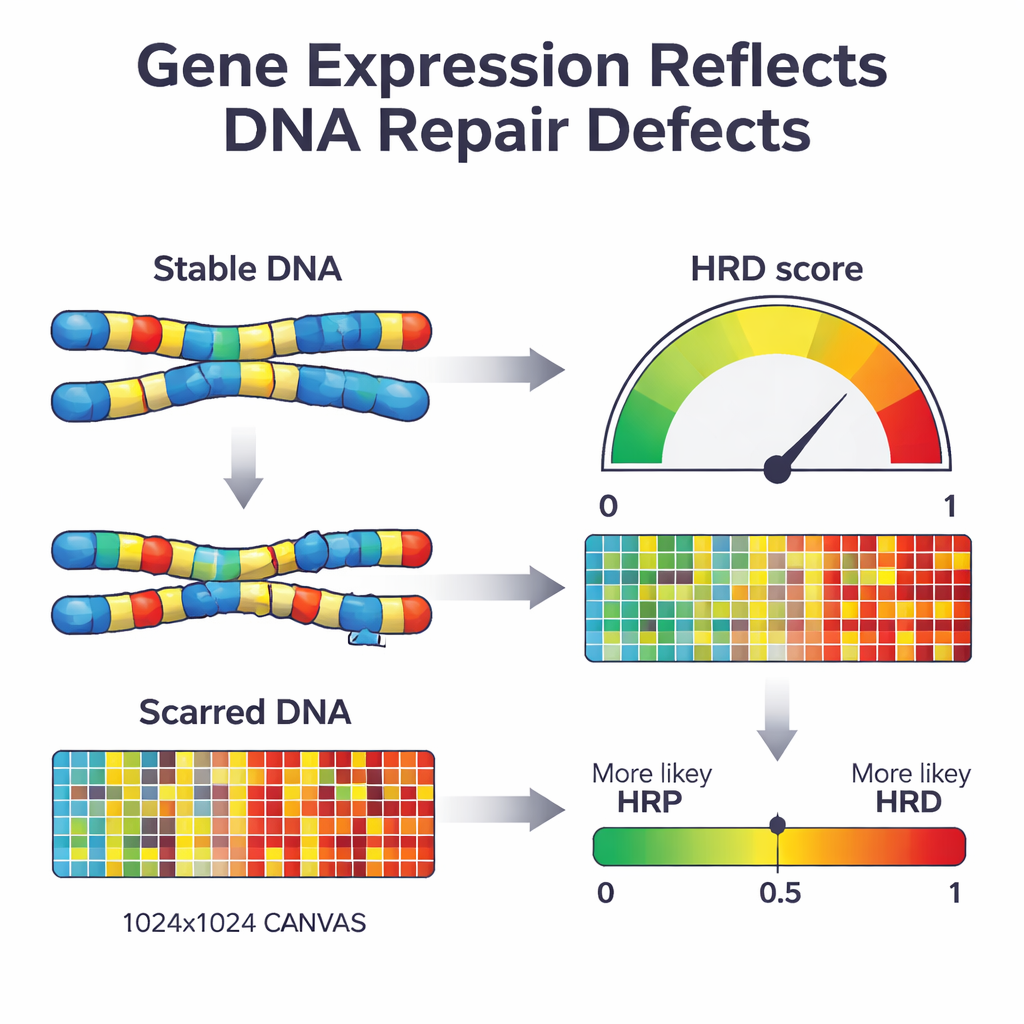
Testing the tool across multiple patient cohorts
The authors rigorously tested IdentifiHR in three independent datasets that had never been used in training. In the held-out subset of The Cancer Genome Atlas, the model correctly distinguished HRD from HRP tumors in about 85% of cases. It performed just as well—around 86% accuracy—in a separate Australian study that included not only primary tumors but also samples taken at autopsy, from fluid in the abdomen (ascites), and from normal fallopian tubes, the likely site where many of these cancers begin. In every normal fallopian tube sample, IdentifiHR correctly predicted intact DNA repair. The tool also worked on “pseudobulked” single-cell data, where thousands of individual cancer cells were combined computationally to mimic a bulk sample, again achieving about 84% accuracy. Across these tests, IdentifiHR matched or exceeded the performance of several existing gene-based methods originally developed for other cancers or for predicting related damage scores.
How this could change research and care
Because IdentifiHR runs on RNA data, which are often cheaper and easier to collect than whole-genome DNA profiles, it offers a practical way for researchers—and potentially, in the future, clinicians—to estimate DNA repair status when only gene expression data are available. The model is released as an open-source R package, so any group with suitable sequencing data can apply it. While it does not yet replace gold-standard DNA tests, and its ability to capture more subtle changes such as repair restoration still needs study, IdentifiHR provides a powerful new lens on which ovarian tumors are most likely to respond to PARP inhibitors and similar drugs. For patients, this line of work moves the field closer to more precise, biology-driven treatment decisions tailored to the actual behavior of their cancer cells.
Citation: Weir, A.L., Lee, S.C., Li, M. et al. IdentifiHR predicts homologous recombination deficiency in high-grade serous ovarian carcinoma using gene expression. Commun Med 6, 119 (2026). https://doi.org/10.1038/s43856-026-01387-y
Keywords: ovarian cancer, DNA repair, homologous recombination deficiency, gene expression, machine learning