Clear Sky Science · en
Homologous recombination deficiency in primary ER-positive and HER2-negative breast cancer
Why this matters for breast cancer patients
Most breast cancers fall into a large group called estrogen‑receptor positive and HER2‑negative. These tumors are usually treated with hormone‑blocking drugs, sometimes combined with chemotherapy. Yet many patients still relapse, and doctors lack precise tools to decide who really needs intensive treatment or could benefit from new targeted drugs. This study explores a specific weakness in some tumors—called homologous recombination deficiency, or HRD—that makes cancer cells bad at repairing broken DNA, potentially opening the door to more personalized treatment.
A hidden weakness in only some tumors
Homologous recombination is one of the cell’s main systems for fixing dangerous DNA breaks. When this system fails—often because of faults in well‑known genes such as BRCA1, BRCA2, PALB2, or RAD51C—cells accumulate mutations and may become especially sensitive to certain drugs, including platinum chemotherapies and PARP inhibitors. HRD is common in an aggressive form of breast cancer called triple‑negative, but its role in the more common estrogen‑receptor positive, HER2‑negative group has been uncertain. To clarify this, researchers analyzed 502 such tumors from the Swedish SCAN‑B study using whole‑genome sequencing, along with matched data on gene activity, DNA methylation, treatments received, and long‑term outcomes.
How the study was done
All tumors were collected at the time of surgery, before any drug treatment, providing a clear snapshot of their original biology. The team used a sophisticated pattern‑recognition tool, HRDetect, to read the “mutational signatures” left in cancer DNA when homologous recombination is broken. Tumors were labeled as HRD if they crossed a strict probability threshold. The scientists also compared different ways of detecting HRD—from other DNA‑based scores to an RNA‑based gene expression test—to see how reliably different methods agree. In addition, they examined broader features such as overall mutation load, patterns of chromosomal gains and losses, immune‑related gene activity, and chemical marks on DNA (methylation) that can switch genes on or off.
How common is HRD, and what causes it?
The researchers found that only 8.4% of estrogen‑receptor positive, HER2‑negative tumors in their sequenced group showed strong evidence of HRD—far lower than the roughly 60% seen in triple‑negative disease. By combining these data with national registry information and other studies, they estimated that around 1 in 20 tumors in this clinical subgroup, and about 1 in 9 breast cancers overall in Western Europe/Nordic populations, are HRD. In the HRD tumors they could often pinpoint a likely cause: about 70% had clear damage to BRCA1, BRCA2, RAD51C, or PALB2, through inherited mutations, tumor‑specific mutations or deletions, or epigenetic silencing of gene promoters. Strikingly, about one‑third of HRD cases were due to promoter hypermethylation—an added chemical coating that turns off DNA repair genes without changing their sequence. However, roughly 30% of HRD tumors showed no obvious single hit, suggesting additional, yet‑to‑be‑discovered routes to this repair failure.
What HRD tumors look like under the microscope and in the clinic
HRD tumors in this common breast cancer group tended to show hallmarks of more aggressive disease: they often had higher cell‑division rates, lower levels of hormone‑receptor staining, and more complex, mutation‑rich genomes than HR‑proficient tumors. They appeared across nearly all major molecular subtypes, but were rare in the more indolent Luminal A class and relatively enriched in a smaller basal‑like subset. Yet when the researchers looked at overall gene activity and DNA methylation patterns, HRD tumors did not fall into a single, easily recognizable profile. Instead, their expression patterns were diverse, and any differences within specific subtypes were modest. Some HRD tumors, especially in the Luminal B and basal‑like groups, showed higher signals of immune activity and increased expression of PD‑L1, hinting that they might be more visible to the immune system and potentially more responsive to immunotherapy.
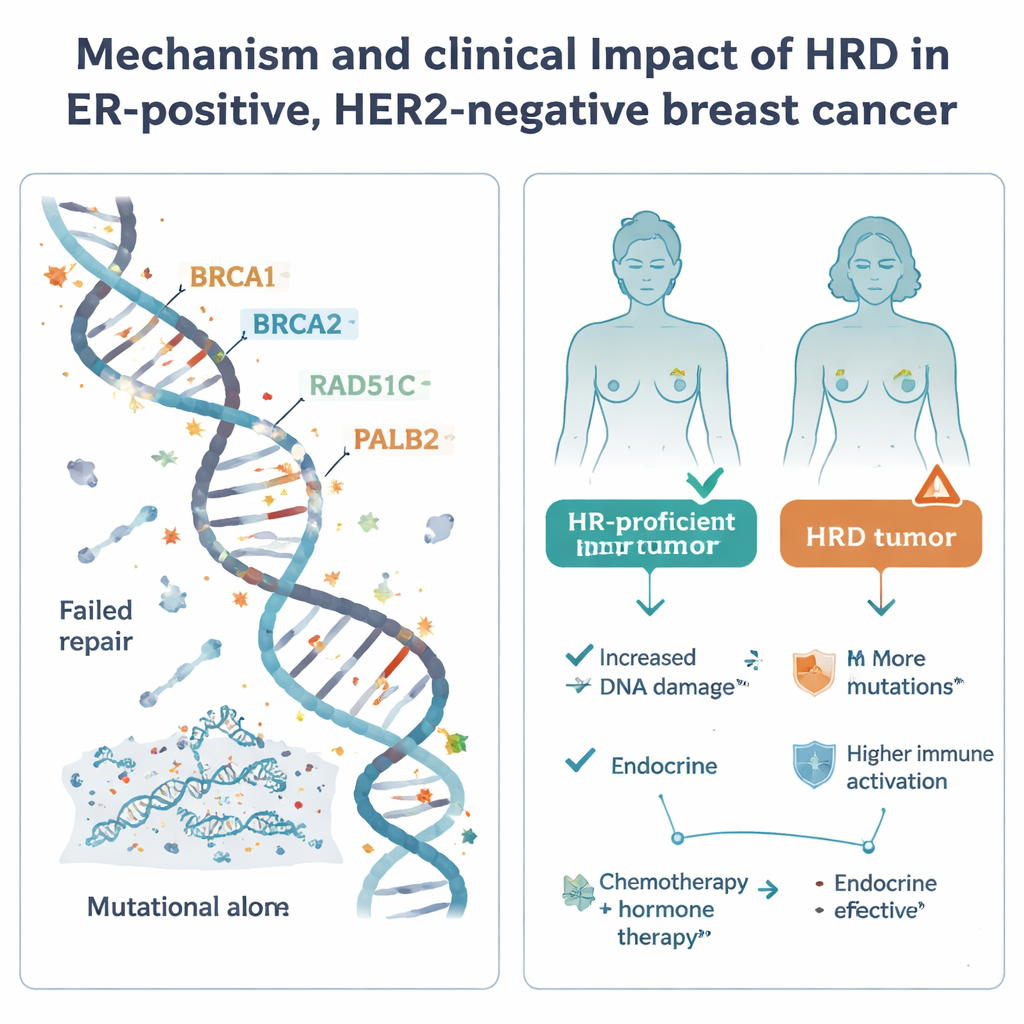
Does HRD change patient outcomes?
The study then examined how HRD status related to real‑world outcomes under standard care. In patients treated with hormone therapy alone after surgery, HRD tumors showed a trend toward worse distant‑relapse‑free survival, though the number of HRD cases was small and the result did not reach formal statistical significance. This pattern, together with their aggressive molecular features, suggests that relying only on endocrine therapy may be risky for patients whose tumors are HRD. In contrast, among patients who received both chemotherapy and hormone therapy, HRD status was not clearly linked to better or worse outcomes; all groups had broadly similar relapse rates, and chemotherapy may be mitigating some of the added risk in HRD tumors.
What this means for future treatment choices
For a general reader, the key message is that only a minority of estrogen‑receptor positive, HER2‑negative breast cancers carry this particular DNA‑repair weakness, but when present it tends to mark more aggressive disease that may not be well controlled by hormone therapy alone. The work supports the idea that whole‑genome sequencing can reliably flag HRD and clarify its underlying causes, beyond just testing for inherited BRCA1 or BRCA2 mutations. Although the study does not yet prove that changing treatment based on HRD status improves survival, it provides early evidence that patients with HRD tumors might benefit from chemotherapy and, in future trials, from PARP inhibitors or immunotherapies. In short, HRD testing could become one piece of a more refined toolkit to tailor therapy intensity—and novel targeted options—to the biology of each patient’s breast cancer.
Citation: Davies, H.R., Black, D., Kvist, A. et al. Homologous recombination deficiency in primary ER-positive and HER2-negative breast cancer. Commun Med 6, 118 (2026). https://doi.org/10.1038/s43856-026-01385-0
Keywords: breast cancer, DNA repair, BRCA genes, genome sequencing, targeted therapy