Clear Sky Science · en
Predicting and interpreting cell-type-specific drug responses in the small-data regime using inductive priors
Why this research matters for future medicines
When a new drug is tested, one of the biggest unknowns is how differently it will affect the many kinds of cells in our bodies. A compound that helps one cell type might do little in another or even cause harm. Generating this information experimentally for thousands of drugs and countless cell types is far too slow and expensive. This paper introduces a computer-based approach, called PrePR-CT, that learns to predict how individual cell types respond to drugs, even when only limited data are available. The work points toward faster, cheaper and more precise ways to explore potential medicines in silico before committing to costly lab and clinical studies.
Looking inside cells instead of just at drugs
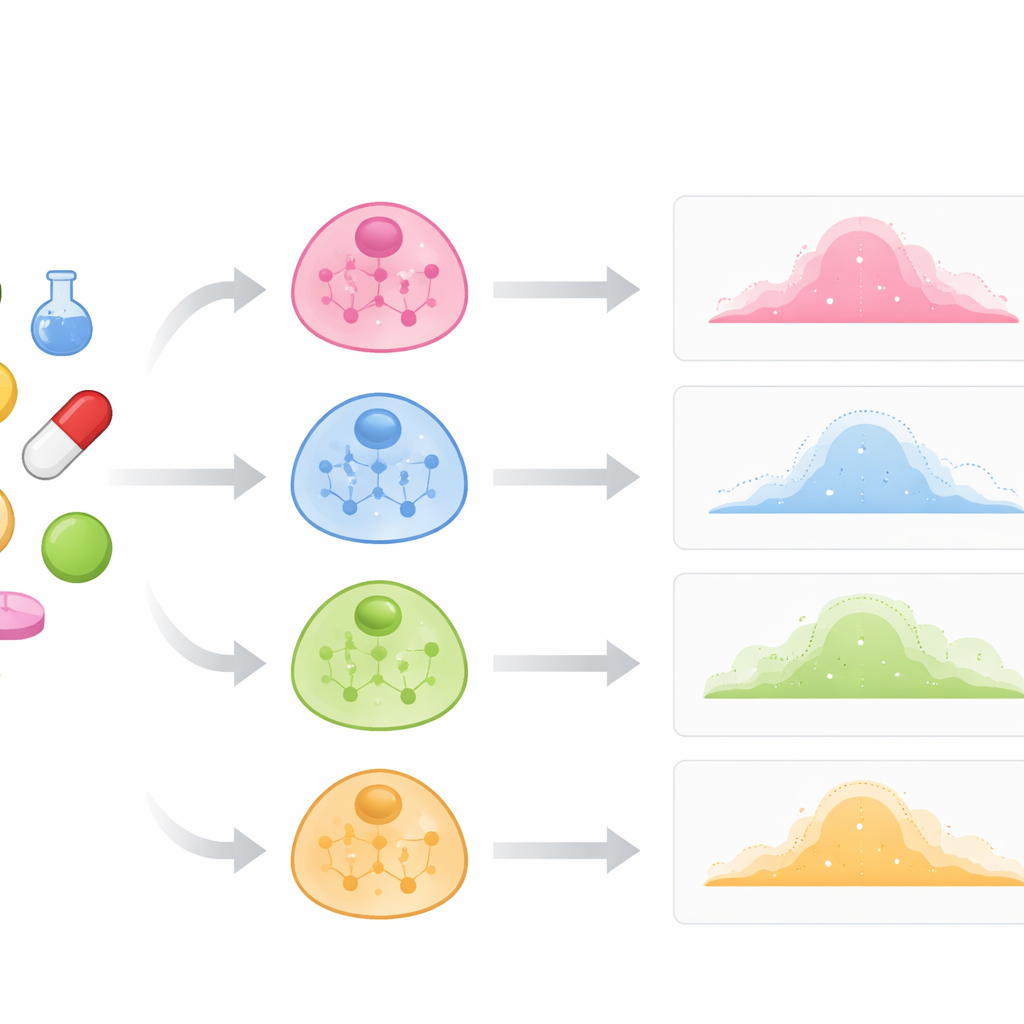
Traditional drug screens often treat cells as if they were all the same and focus mainly on bulk averages. In reality, immune cells, liver cells and cancer cells can react very differently to the same compound. The authors argue that to predict these differences, a model must understand the internal wiring of each cell type: which genes tend to be active together and how those patterns define the cell’s identity. They build cell-type "maps" by examining which genes in unperturbed (control) cells rise and fall in concert. Each map is represented as a network, where nodes stand for genes and links reflect strong co-activity. These networks serve as prior knowledge about how a given cell type is organized before any drug is added.
A network-aware learning engine
PrePR-CT combines three ingredients: the gene-activity network of a cell type, the baseline gene expression of that cell type, and a compact description of the drug’s chemical structure. The model uses a class of neural networks designed for graphs to digest the cell’s gene network and extract a summary that captures its characteristic patterns. In parallel, it turns each drug into a numerical fingerprint derived from its molecular structure. These pieces are fed into a downstream prediction module that learns, from available experiments, how a given drug will shift the distribution of gene activity in that cell type. Rather than producing a single number per gene, the method estimates both the average change and how variable the response is across individual cells, which is crucial for understanding subtle and strong effects alike.
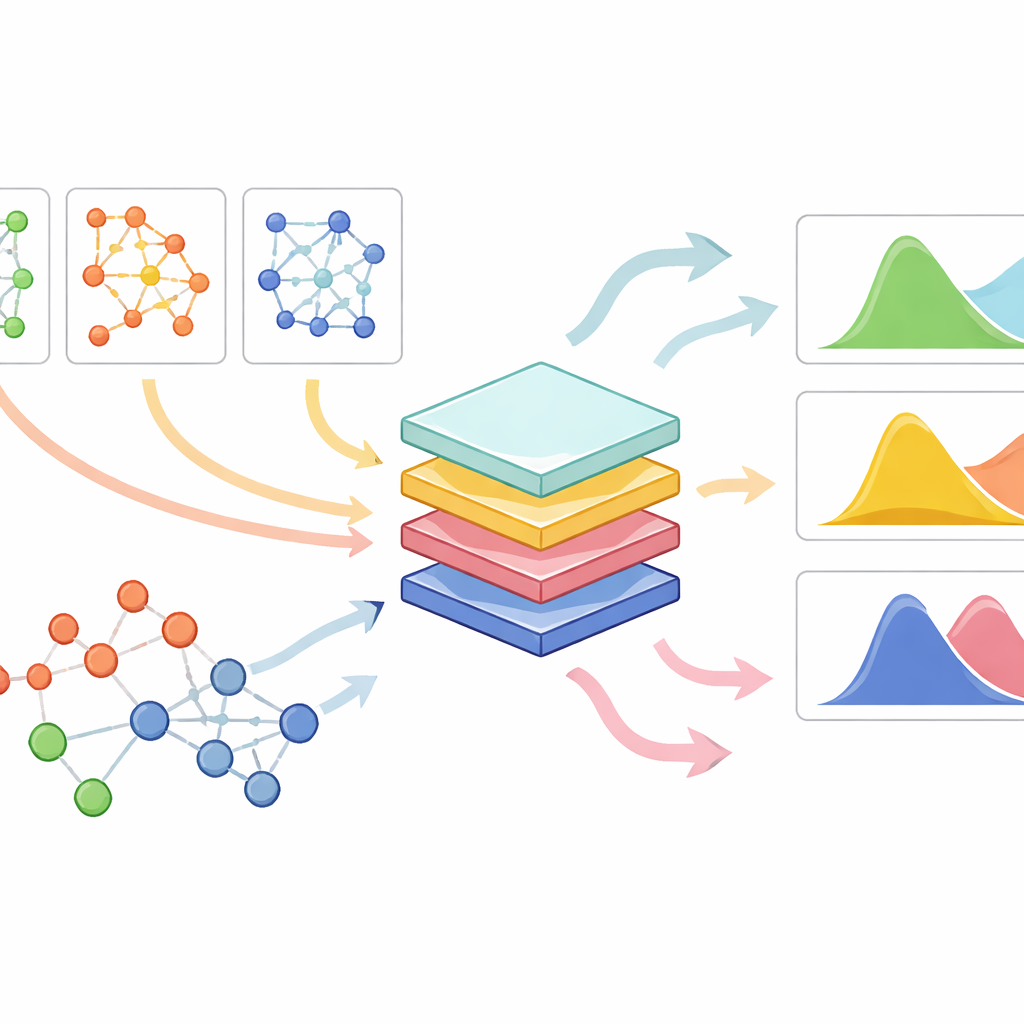
Working across many cell types, drugs and small datasets
The researchers tested PrePR-CT on a wide collection of datasets, including human blood cells exposed to immune signals, multiple cancer cell lines treated with various compounds, mouse liver cells exposed to a pollutant and large-scale drug screens from public resources. In challenging scenarios where an entire cell type was held out during training, the model could still predict how that new cell type would respond to a familiar drug, often with accuracy exceeding earlier generative models. Likewise, when a new drug but familiar cell type was held out, the method successfully anticipated its impact using only its chemical fingerprint. Importantly, the model remained effective when trained on relatively small numbers of cells, a setting where many deep learning approaches struggle.
From black box to clues about mechanism
Beyond raw prediction, the authors wanted to know whether their model could offer insight into which genes and pathways drive a cell’s response. The graph-based architecture includes an attention mechanism that highlights genes the model deems especially influential in each cell type. Many of these "high-attention" genes were not the usual suspects flagged by standard differential expression analysis, yet they clustered in immune-related pathways consistent with the biology of the tested drugs. When the researchers deliberately disrupted these influential genes in the model’s input, prediction quality dropped, especially for the most responsive genes, suggesting that the attention scores point to meaningful mechanistic players rather than noise.
What this means for designing better drugs
In plain terms, this work shows that giving artificial intelligence models a structured view of how each cell type is wired—its internal gene network—greatly improves their ability to forecast how drugs will reshape those cells, even when only modest data are available. PrePR-CT does not replace experiments, but it can help narrow down which compounds and cell types are worth testing and hint at why certain cells react as they do. As datasets grow and additional cellular features are incorporated, such approaches could become key tools for tailoring therapies to specific tissues or patient cell types, reducing trial-and-error in the lab and bringing more precise medicines closer to reality.
Citation: Alsulami, R., Lehmann, R., Khan, S.A. et al. Predicting and interpreting cell-type-specific drug responses in the small-data regime using inductive priors. Nat Mach Intell 8, 461–473 (2026). https://doi.org/10.1038/s42256-026-01202-2
Keywords: drug response prediction, single-cell transcriptomics, graph neural networks, drug discovery, cell-type specificity