Clear Sky Science · en
Reusability Report: Evaluating the performance of a meta-learning foundation model on predicting the antibacterial activity of natural products
Searching for New Antibiotics, Faster
Antibiotic resistance is rising, yet discovering new drugs is painfully slow and often depends on trial and error in the lab. This study explores whether a powerful kind of artificial intelligence, originally trained on huge collections of drug data, can be quickly adapted to predict which plant-derived natural compounds might fight bacteria—using only small amounts of new experimental data. If successful, such tools could help scientists focus precious lab time on the most promising candidates and speed up the search for the next generation of antibiotics.
Why Plant Chemicals Matter
Many of our best antibiotics started their lives as natural products from plants and microbes. These molecules can stop bacterial growth, but finding new ones in nature is a bit like looking for needles in a haystack. Researchers must test many compounds against many bacterial strains, and each test is costly. Worse, large, carefully labeled datasets—which modern deep learning methods need to perform well—are rare in this area. That makes antibiotic discovery a prime testing ground for “foundation models”: large, general-purpose AI systems that can be fine-tuned for specific tasks with only a handful of new examples.
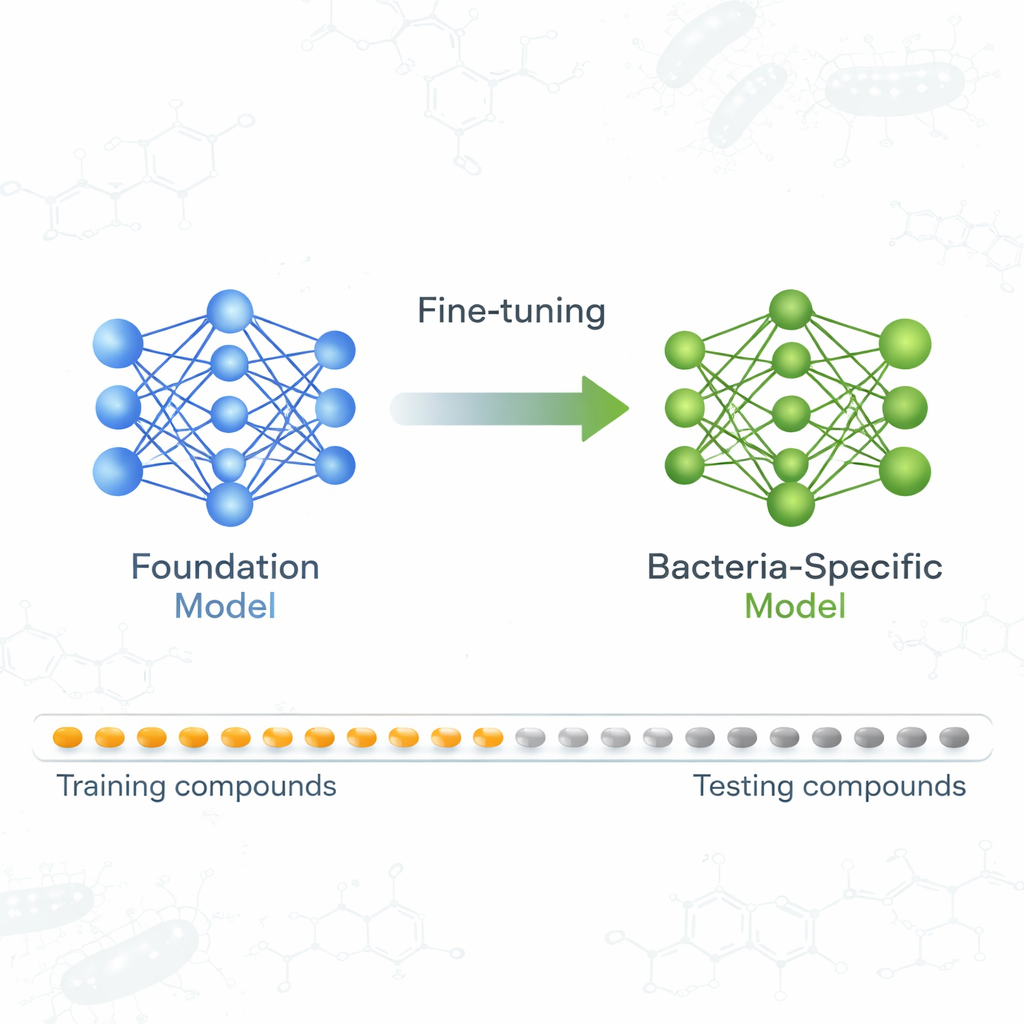
A Foundation Model Learns About Germ Killers
The team focused on a foundation model called ActFound, originally trained to predict how strongly different chemicals affect biological targets, using vast datasets from resources like the ChEMBL and BindingDB databases. Instead of predicting a single number for each compound, ActFound learns by comparing pairs of compounds within the same experiment and estimating which is more active. This “pairwise” learning, combined with a training strategy known as meta-learning, is designed to help the model quickly adapt to new prediction tasks when only a small number of labeled examples are available—exactly the situation in many antibiotic screens.
Testing the Model on Real-World Plant Data
To see how reusable ActFound really is, the authors fine-tuned it on a curated dataset of plant-derived natural products tested for their ability to inhibit the growth of various bacteria. Each bacterial strain was treated as its own task, and the model was adapted using only 8 to 128 compounds per strain, or fixed percentages of the available data. They also compared ActFound to simpler meta-learning and transfer-learning models that do not use pairwise comparisons. Across these tests, ActFound did not reach the accuracy it had shown in earlier work on other types of drug data. However, when very little data was available—roughly only a handful of compounds per strain—ActFound and its transfer-learning variant generally matched or outperformed alternative methods.
When Similarity Helps—and When It Hurts
ActFound assumes that similar molecules behave similarly, which works well when datasets are built around groups of related chemicals. The natural product dataset, however, was chemically diverse and often lacked closely related “families” of compounds. This diversity, while scientifically valuable, undermined the pairwise learning strategy: when compounds within an experiment are very different from each other, the model has trouble learning stable comparisons. The authors also found that a simple diagnostic, proposed in the original ActFound paper to predict in advance how well the model would perform on a new task, did not hold up for these natural product data, highlighting an important limitation when moving to new chemical spaces.
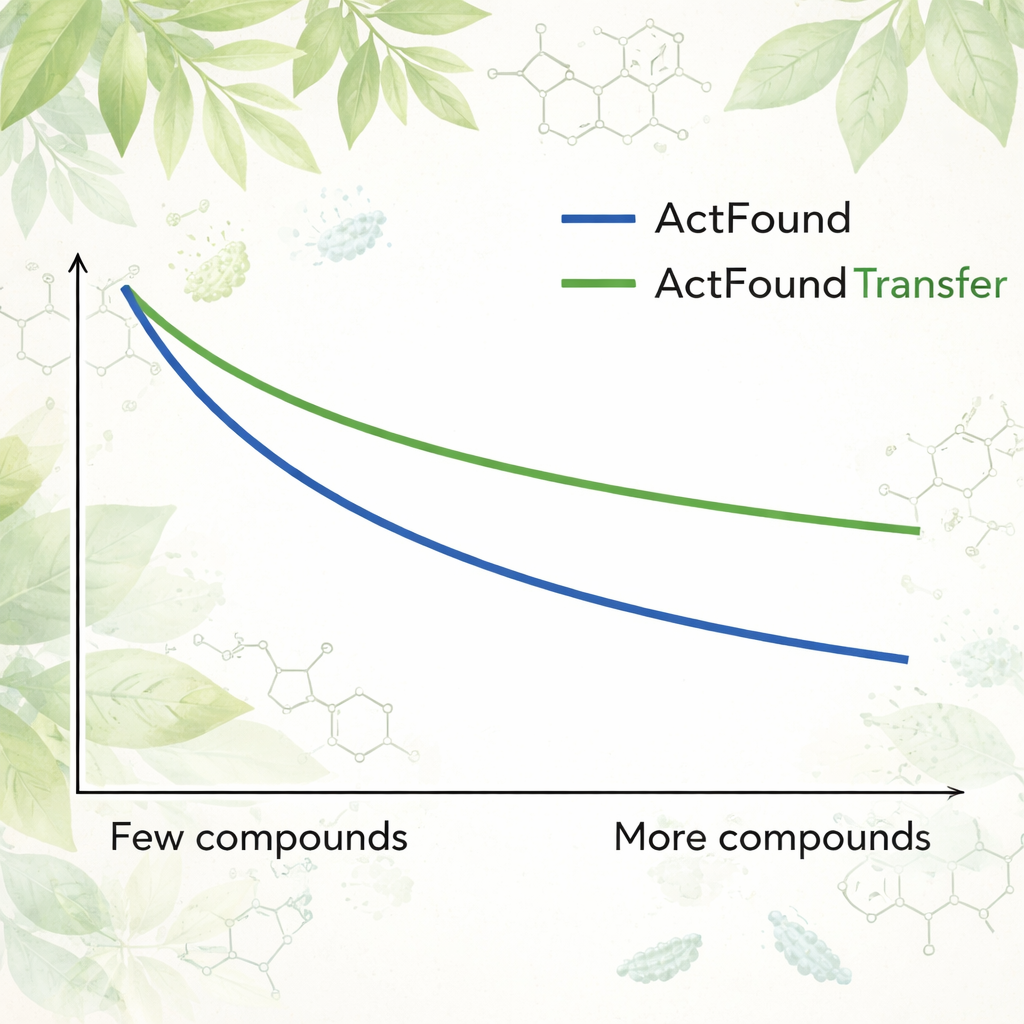
What This Means for Future Drug Discovery
For non-specialists, the takeaway is that foundation models like ActFound are promising tools for drug discovery when data are scarce, but they are not magic bullets. In this study, ActFound and its transfer-learning version often did as well as or better than competing methods when only a few plant compounds were available for training, yet they struggled on this highly diverse set of natural products. The work suggests that these AI models are most useful when the data include many chemically similar compounds—such as in focused studies of structure–activity relationships—but they remain less reliable for predicting how entirely new kinds of molecules will behave. In other words, AI can help narrow the search, but the hardest part of exploring truly new chemical territory still lies ahead.
Citation: Butt, C.M., Walker, A.S. Reusability Report: Evaluating the performance of a meta-learning foundation model on predicting the antibacterial activity of natural products. Nat Mach Intell 8, 270–275 (2026). https://doi.org/10.1038/s42256-026-01187-y
Keywords: antibiotic discovery, natural products, deep learning, meta-learning, drug screening