Clear Sky Science · en
Unravelling the molecular mechanisms causal to type 2 diabetes across global populations and disease-relevant tissues
Why understanding diabetes roots matters
Type 2 diabetes affects hundreds of millions of people worldwide, yet we still know surprisingly little about which molecular switches in the body truly cause the disease rather than just accompany it. This study digs deep into our DNA and into multiple organs to pinpoint which genes and proteins actually push blood sugar toward diabetes, and which ones help protect against it. By including people from diverse ancestries and examining several key tissues, the researchers move us closer to more precise prevention strategies and therapies that can work for many populations, not just those of European descent.
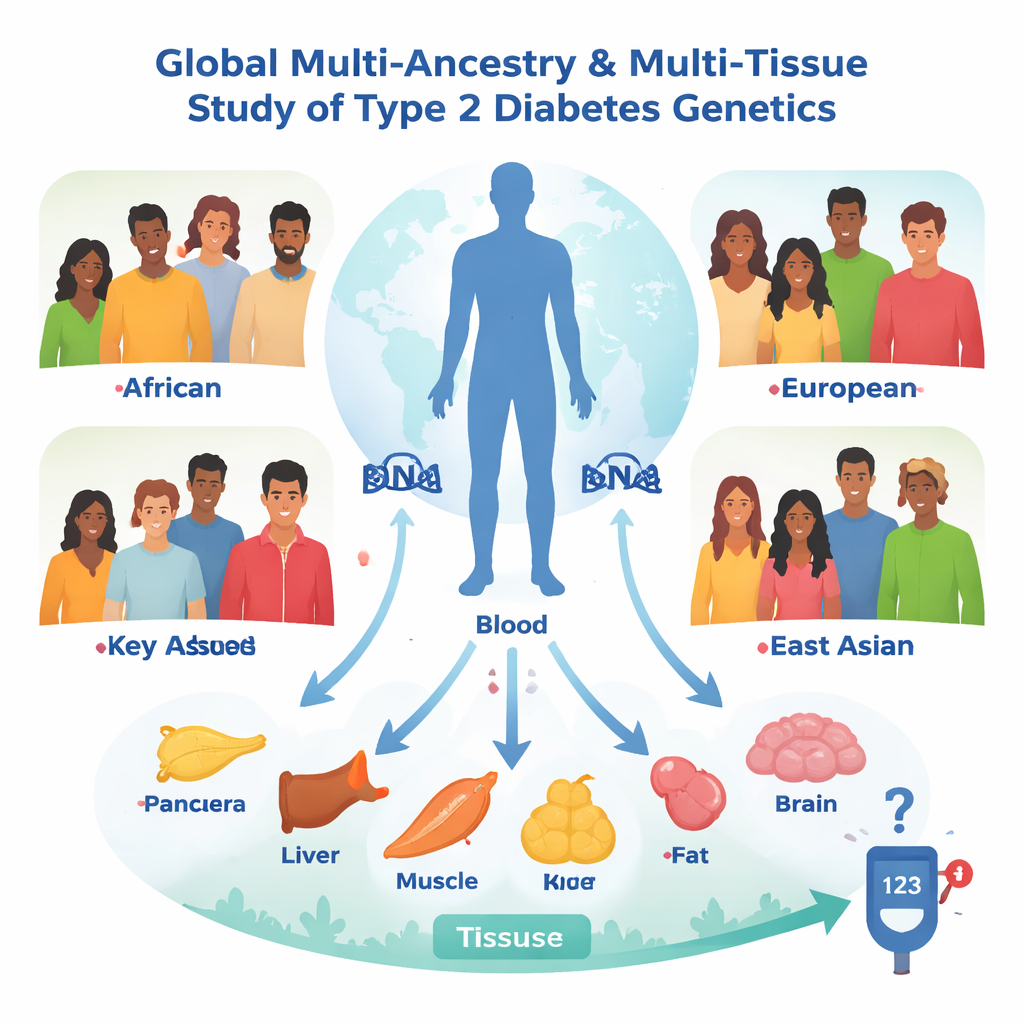
Looking across the globe and inside the body
The team began with genetic data from more than 2.5 million people gathered by the Type 2 Diabetes Global Genomics Initiative. Instead of just asking which DNA variants are linked to diabetes, they asked a more powerful question: which variants change the activity of specific genes or proteins in the body, and do those changes in turn alter diabetes risk? To do this, they used a statistical approach called Mendelian randomization, which treats naturally occurring genetic differences as a kind of built-in randomized trial. They analysed over 20,000 gene activity measures and more than 1,600 blood proteins in people from four ancestry groups—European, African, admixed American and East Asian—and then repeated the analyses in seven tissues that are central to blood sugar control, including the pancreas, insulin-producing islets, liver, muscle and different fat depots.
Finding molecular levers that raise or lower risk
By tracing these genetic pathways, the researchers identified 335 genes and 46 blood proteins whose genetically predicted levels have a causal impact on type 2 diabetes risk, and they confirmed many of these findings in independent cohorts. Some of the molecular levers they found were already known suspects, such as MTNR1B, a gene involved in insulin release from pancreatic islet cells, and BAK1, which influences cell death in the pancreas and fat. Others were new or less appreciated players, including CPXM1, a protein tied to fat tissue development and insulin resistance, and HIBCH, a gene involved in mitochondrial function. Overall, they catalogued 923 genes and 46 proteins with evidence that shifting their activity in at least one tissue can change the odds of developing diabetes.
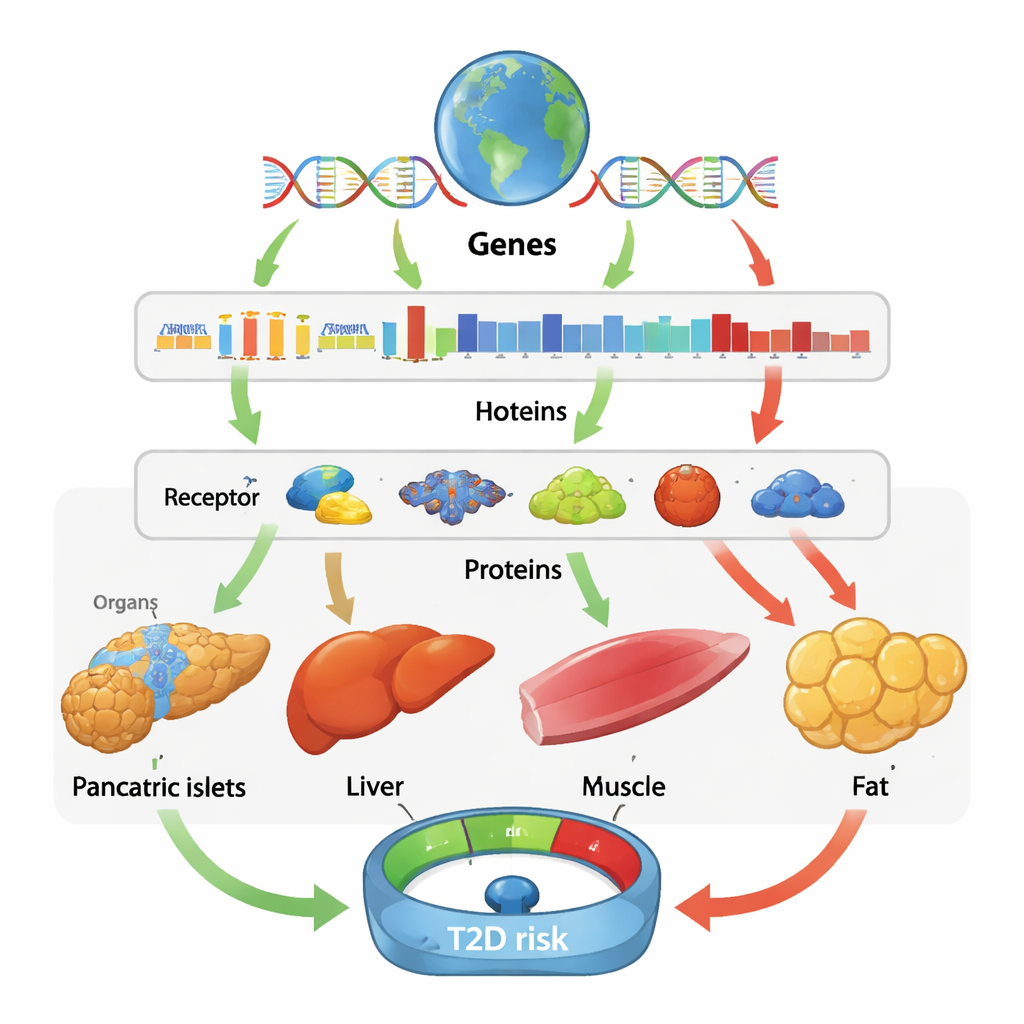
Same genes, different stories across tissues and peoples
One striking lesson is that the effect of a gene is often highly tissue-specific. For example, higher BAK1 activity in the pancreas and islets appears to increase diabetes risk, likely by contributing to the loss of insulin-producing cells, whereas higher BAK1 in fat and muscle seems to be protective. HIBCH showed a similarly mixed pattern: in some tissues, more activity lowered diabetes risk, while in others it raised it. These findings show that looking only at blood can miss crucial biology happening inside organs, and that the same molecule can be beneficial in one tissue yet harmful in another. In contrast, when the team compared results across ancestries, they saw relatively little difference in effect sizes, suggesting that many of the underlying causal mechanisms are shared globally, even though certain signals—such as specific protective or risk proteins in East Asian or African groups—were only detectable thanks to non-European data.
Connecting new findings to known diabetes biology
To check whether their approach was making biological sense, the authors cross-referenced their causal genes with curated lists of diabetes-related genes from human studies and mouse experiments. Genes with the strongest prior evidence for involvement in diabetes were far more likely to show causal effects in their analyses than genes chosen at random. Moreover, the tissues in which these causal effects appeared matched known disease mechanisms: genes linked to beta-cell failure tended to matter most in pancreatic islets, while genes tied to metabolic syndrome showed their strongest effects in visceral (deep belly) fat. This alignment supports the idea that the statistical pipeline is successfully homing in on mechanisms, not just correlations.
What this means for future treatment and prevention
For non-specialists, the key takeaway is that this work turns long, impersonal lists of DNA variants into a clearer map of specific genes, proteins and organs that actually drive type 2 diabetes. By distinguishing cause from effect and revealing when the same molecule has opposite roles in different tissues, it gives drug developers more precise targets and warns where a one-size-fits-all therapy might backfire. Importantly, by deliberately including diverse populations, the study helps ensure that any future medicines or risk scores built from these insights have a better chance of working broadly, not just for people of European ancestry.
Citation: Bocher, O., Arruda, A.L., Yoshiji, S. et al. Unravelling the molecular mechanisms causal to type 2 diabetes across global populations and disease-relevant tissues. Nat Metab 8, 506–520 (2026). https://doi.org/10.1038/s42255-025-01444-1
Keywords: type 2 diabetes, genetic mechanisms, multi-ancestry genomics, tissue-specific gene expression, causal inference