Clear Sky Science · en
Structure and polydispersity of single lipid vesicles by small-angle X-ray scattering at European XFEL
Why tiny bubbles in water matter
Lipid vesicles are microscopic bubbles made from the same kind of fatty molecules that build our cell membranes. They are key players in drug delivery, cosmetics, and in how cells ship hormones and neurotransmitters. Yet, because each vesicle is only tens of billionths of a meter across and sits in water, it is surprisingly hard to see its detailed structure. This study shows how to examine individual vesicles one by one with intense X-ray flashes, revealing not just their average structure but also how much they differ from each other—information that is crucial for both biology and nanotechnology.
From blur to single-particle clarity
For decades, scientists have used a method called small-angle X-ray scattering to study soft materials like proteins, nanoparticles, and lipid vesicles in solution. In a typical experiment, a thin beam of X-rays passes through a sample containing astronomically many copies of the same kind of particle. The beam is scattered, and the resulting pattern encodes their overall size and internal structure. The catch is that this approach returns only averages over trillions of particles, all in random orientations and with slightly different sizes and shapes. Much of the interesting detail—such as how broad that size distribution actually is, or how much each particle deviates from a perfect sphere—gets washed out.
Freezing motion with ultrafast X-ray pulses
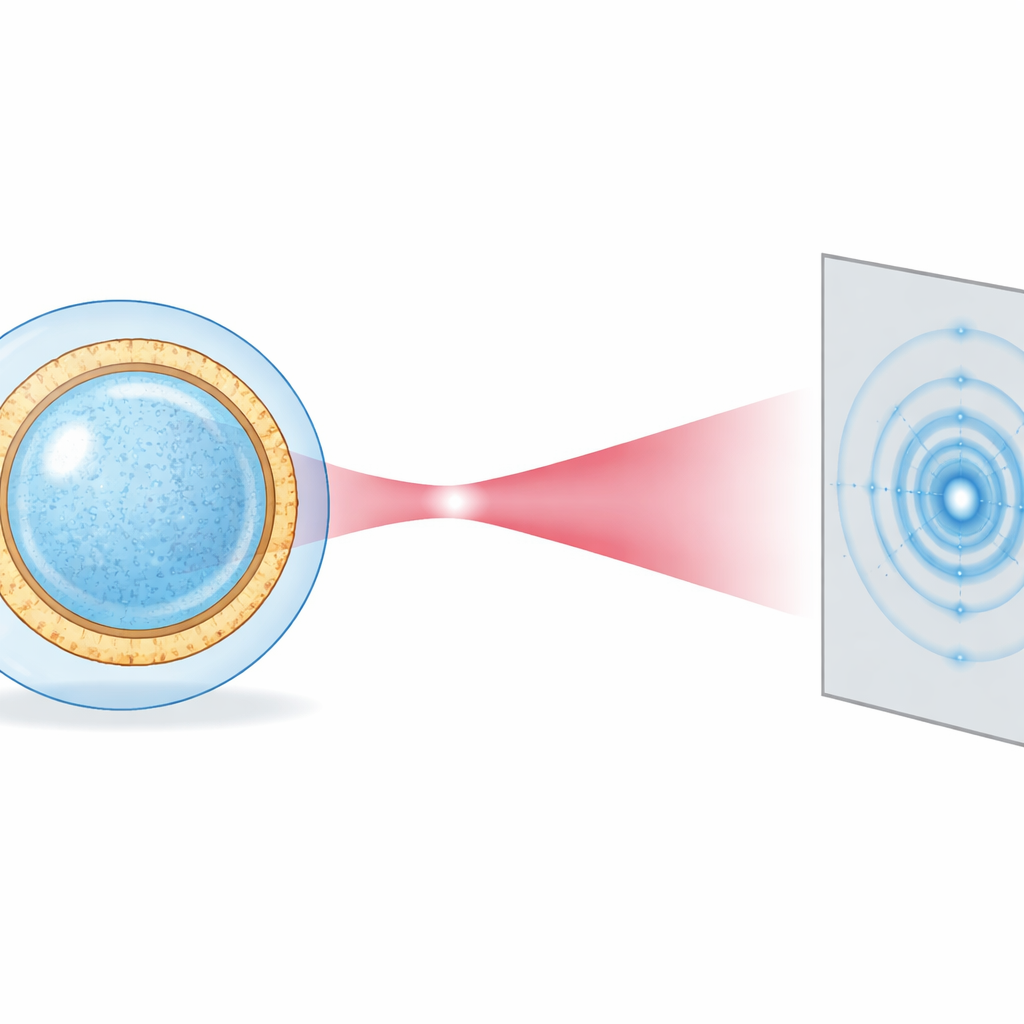
To move beyond averages, the authors turn to an X-ray free-electron laser (XFEL) at the European XFEL facility. This machine produces ultrashort, extremely bright X-ray pulses lasting just a few quadrillionths of a second. In that instant, a single vesicle can be probed before the intense radiation has time to damage it, a concept known as “diffract-before-destroy.” The team uses an aerosol injector to spray individual vesicles from water into vacuum, where the droplets rapidly cool and vitrify, leaving intact vesicles wrapped in a thin layer of water. A nano-focused X-ray beam, only a few hundred nanometers wide, hits one vesicle at a time, and a large-area detector records the resulting diffraction pattern.
Turning patterns into shapes and shells
Each vesicle produces a faint, ring-like pattern that depends on its radius, its deviation from a perfect sphere, and the detailed layering of electron-rich lipid head groups and more diffuse fatty tails in the membrane. Instead of trying to reconstruct a full image pixel by pixel—a process that needs many identical copies—the researchers fit each pattern directly with a physically motivated model borrowed from conventional solution scattering. The vesicle is treated as a slightly squashed sphere surrounded by a smooth water shell, and the membrane is described by simple mathematical bell-shaped curves. By azimuthally averaging each pattern (turning it into a one-dimensional curve) and performing least-squares fits, they extract, for every vesicle, its radius, its ellipticity (how stretched or flattened it is), and an estimate of the membrane’s internal density profile.
Mapping real-world variability
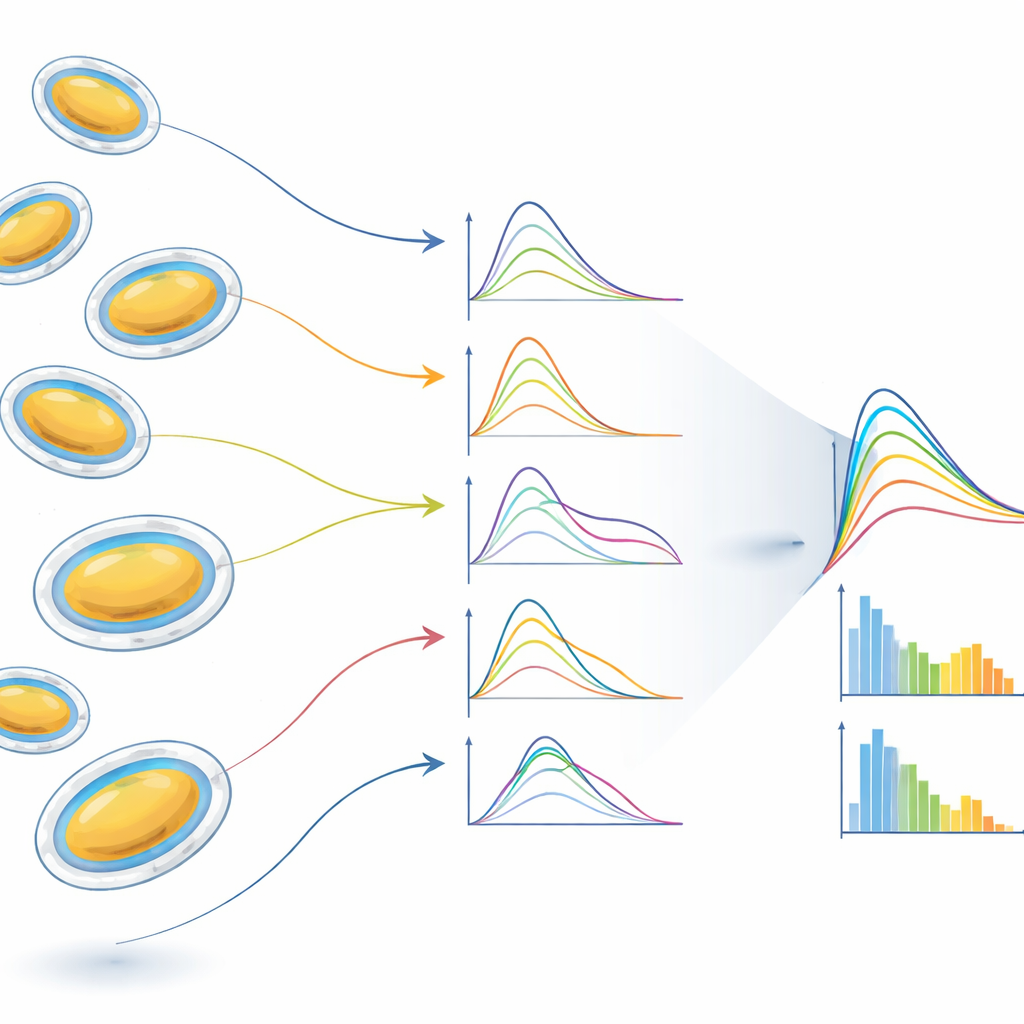
Because the experiment runs at high repetition rate, the team collects more than a million images per run. Automated “hit-finding” routines select those that truly contain a single vesicle rather than multiple particles or empty shots. From thousands of such hits, the researchers build histograms of vesicle radius and shape. They find that vesicles, prepared to be spherical, often become slightly flattened ellipsoids during aerosolization, likely because water slowly leaves the interior while the membrane remains hydrated on the outside. The data also reveal how strongly variations in size smear out the characteristic wiggles of the scattering curves, and how selecting subsets of vesicles with similar radii or shapes—an “in silico purification”—restores clearer structural signals of the membrane bilayer and its thin surrounding water layer.
A new window on soft nanostructures
By combining XFEL pulses, single-particle delivery, and model-based analysis, this work effectively brings traditional small-angle X-ray scattering down to the level of individual vesicles. Instead of one average curve for a huge ensemble, researchers can now obtain structural parameters for each vesicle separately and then deliberately regroup them to study well-defined subpopulations. This makes it possible to both reduce the blurring caused by polydispersity and to measure that polydispersity itself in detail. The approach is broadly applicable to fragile biological and soft-matter systems that are heterogeneous by nature—from drug-carrying liposomes and proteo-liposomes to more complex cellular compartments—opening the door not only to better static structure measurements but eventually also to real-time movies of structural changes triggered by light or other stimuli.
Citation: Neuhaus, C., Stammer, M.L., Alfken, J. et al. Structure and polydispersity of single lipid vesicles by small-angle X-ray scattering at European XFEL. Commun Phys 9, 93 (2026). https://doi.org/10.1038/s42005-026-02551-5
Keywords: lipid vesicles, X-ray free-electron laser, small-angle X-ray scattering, single-particle imaging, nanobiotechnology