Clear Sky Science · en
Network hierarchy entropy for quantifying graph dissimilarity
Why Tiny Differences in Networks Matter
From social media friendships to airline routes and protein structures, many systems around us can be represented as networks of nodes and links. But telling when two such networks are meaningfully different is surprisingly hard, especially when they look similar at first glance. This paper introduces a new way to measure how different two networks really are, by paying attention not just to individual points (nodes) but also to the connections (edges) and how they work together. The method, called network hierarchy entropy, can spot subtle structural changes that other tools miss and even helps distinguish enzyme proteins from non-enzymes.
Looking at Networks Layer by Layer
To understand a network, the authors first consider how far each node is from every other node in terms of steps along links. Around any chosen node, other nodes can be grouped into layers: immediate neighbors, neighbors of neighbors, and so on. This “hierarchy” around a node describes how influence or an infection might spread outward through the network. The twist is that two very different networks can share the same node-level layering, so this view alone can fail to tell them apart. The paper shows that classic examples, like the Desargues and Dodecahedral graphs, share identical node hierarchies even though their internal wiring is different.
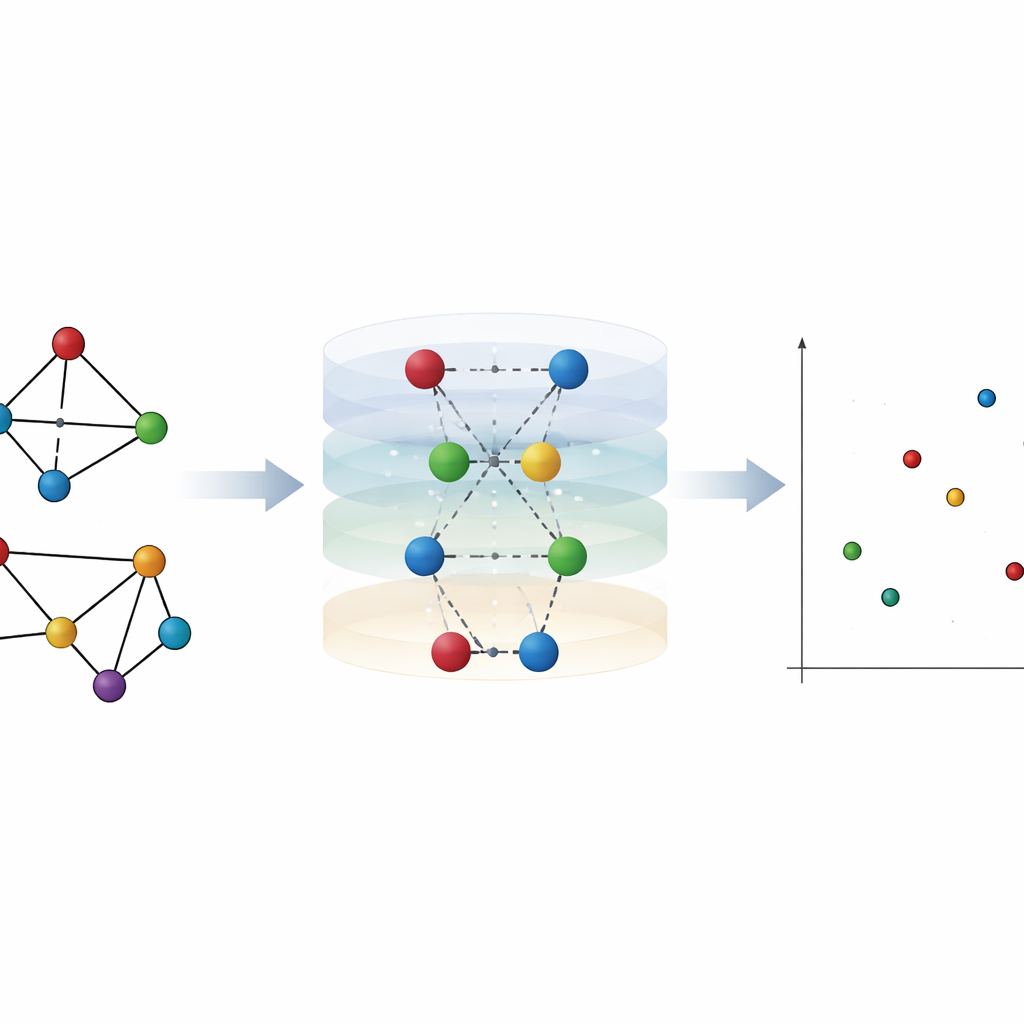
Letting Links Speak: Shrinking Node Pairs
To capture what node-level views miss, the authors focus on edges—the links between nodes—and how they reshape distances across the network. They introduce a simple but powerful idea: "node pair shrinking." Here, two connected nodes are temporarily merged into a single new node, while still keeping their combined neighbors. This reveals how close every other node is to the pair compared with each endpoint alone. In spreading terms, it mimics the effect of infecting both ends of a link at once rather than starting from just one node. From these layered distance patterns, they define "hierarchy centrality" for both nodes and edges, which turns out to correlate strongly with how well nodes or edges act as spreaders in simulated epidemics on real-world networks.
Measuring Information Loss with Entropy
Building on these centralities, the authors define two kinds of hierarchy entropy. Edge hierarchy entropy asks: how much information do we lose if we try to approximate an edge’s importance just by averaging the importance of the two nodes it connects? Node hierarchy entropy asks the reverse question for nodes and their surrounding edges. Both quantities are normalized so that they do not depend on the overall network size. Together they form a two-number fingerprint for any network. The distance between two networks is then simply the geometric distance between their fingerprints. This new metric obeys the standard rules expected for a distance and matches intuitive ideas, such as giving larger penalties when a change breaks a network apart.
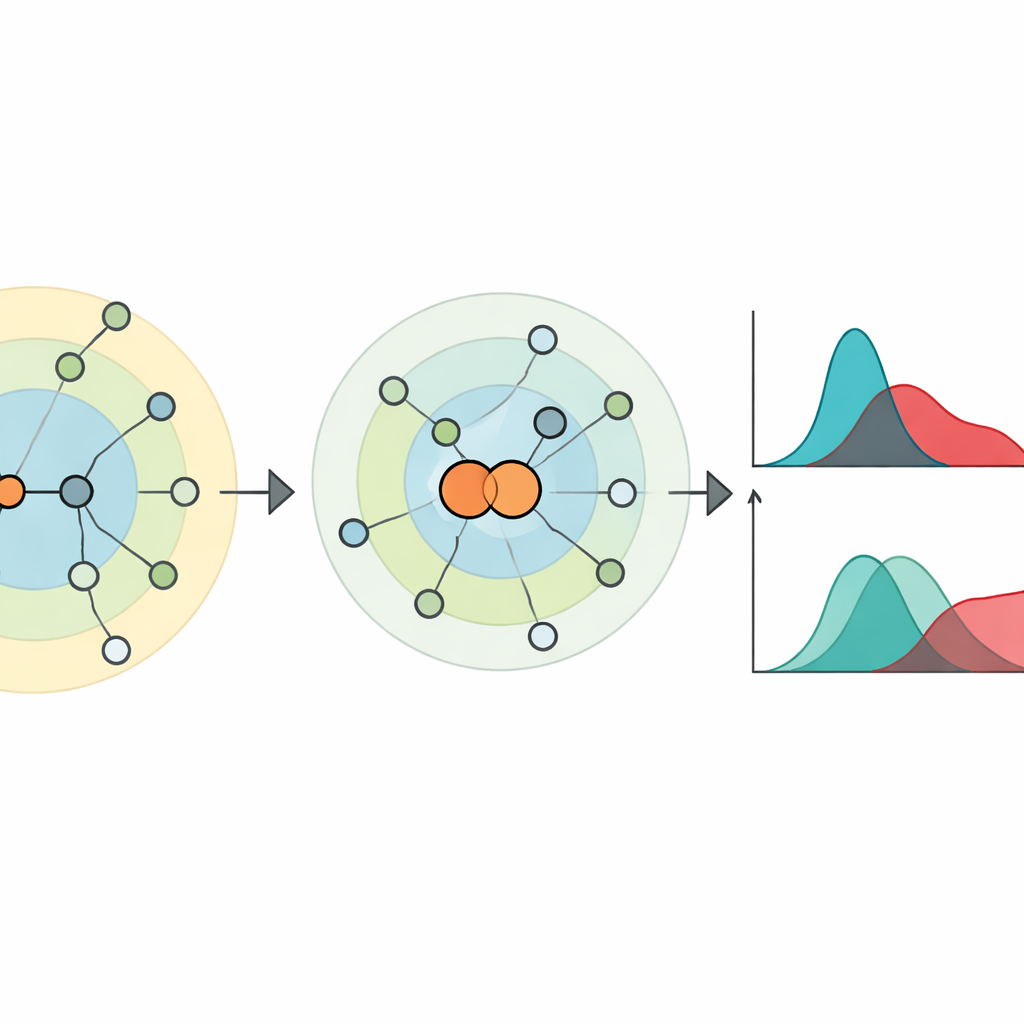
Seeing Finer Structure and Change Over Time
The authors test their measure on both artificial and real networks. In synthetic benchmarks that mimic social or technological systems, the new metric can track how networks evolve as model parameters change, and it clearly separates networks with strong communities from those where communities are weaker, even when competing methods struggle. In controlled experiments where networks are carefully reshuffled to preserve many common statistics—like degree sequences and even distance distributions—the hierarchy entropy distance still detects differences that other popular measures treat as negligible. It also excels at grouping randomized versions of the same network into the right categories, indicating a sharp sensitivity to higher-order structure that goes beyond simple counts of links and paths.
Real-World Uses: Mobility and Proteins
To show practical value, the authors apply their distance measure to daily mobility networks between hundreds of Chinese cities during the first months of COVID-19. Using early January as a baseline, hierarchy entropy reveals how travel patterns shift through the Lunar New Year travel rush, the onset of strict quarantine, and the gradual recovery period, aligning well with known policy changes and mobility community patterns. In another application, they treat protein structures as networks of amino acids connected when they are close in space. Without any learning or handcrafted features, clustering proteins by the new distance reaches about 75% accuracy in separating enzymes from non-enzymes—competitive with modern supervised neural-network approaches.
What This Means in Simple Terms
In essence, this work shows that paying attention to how nodes and links jointly shape distances in a network provides a much sharper "fingerprint" than looking at nodes alone. By quantifying how much is lost when we try to replace edges with their endpoints—or nodes with their surrounding edges—the proposed hierarchy entropy distance highlights subtle structural differences that strongly affect spreading, mobility, and biological function. For scientists and analysts working with any kind of network data, this offers a practical, general-purpose tool for comparing complex systems in a way that is both mathematically sound and closely tied to how processes actually unfold on those networks.
Citation: Mou, J., Wang, L., Zhang, C. et al. Network hierarchy entropy for quantifying graph dissimilarity. Commun Phys 9, 83 (2026). https://doi.org/10.1038/s42005-026-02523-9
Keywords: network similarity, complex networks, entropy measures, epidemic spreading, protein structure networks