Clear Sky Science · en
Structural basis for human RegⅢα filament formation
How Our Guts Fight Germs
Our intestines are constantly exposed to billions of bacteria, many of which are harmless or even helpful. But when dangerous microbes appear, our bodies need fast, precise ways to attack them without damaging our own tissues. This study looks closely at one such natural defense protein, called RegIIIα, and reveals how it changes shape to kill bacteria and then switches into a safer, less harmful form. Understanding this shape-shifting act could help scientists better grasp how we keep gut microbes in balance and what goes wrong in disease.
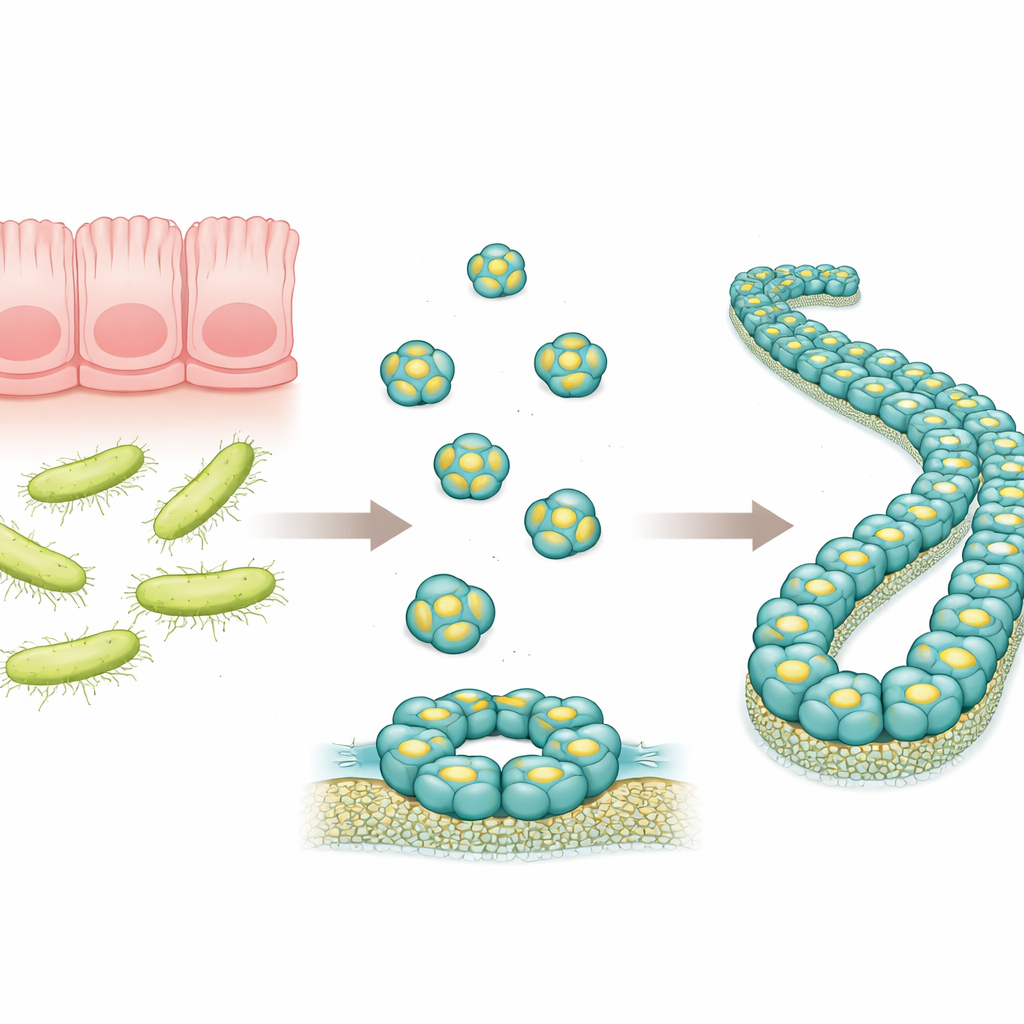
A Tiny Defender in the Intestine
RegIIIα is a protein made in the digestive tract that targets certain bacteria, especially those with thick outer walls known as Gram-positive bacteria. Earlier work suggested that RegIIIα can punch holes in bacterial membranes by forming ring-like structures, allowing the contents of the microbe to leak out and leading to its death. Researchers had also seen that, over time, these ring structures can stack into long filaments, which appear to reduce the protein’s killing power. However, the exact arrangement of RegIIIα molecules in these larger assemblies was not known in detail, making it hard to understand how the protein works and how its activity is controlled.
Seeing the Filament in Atomic Detail
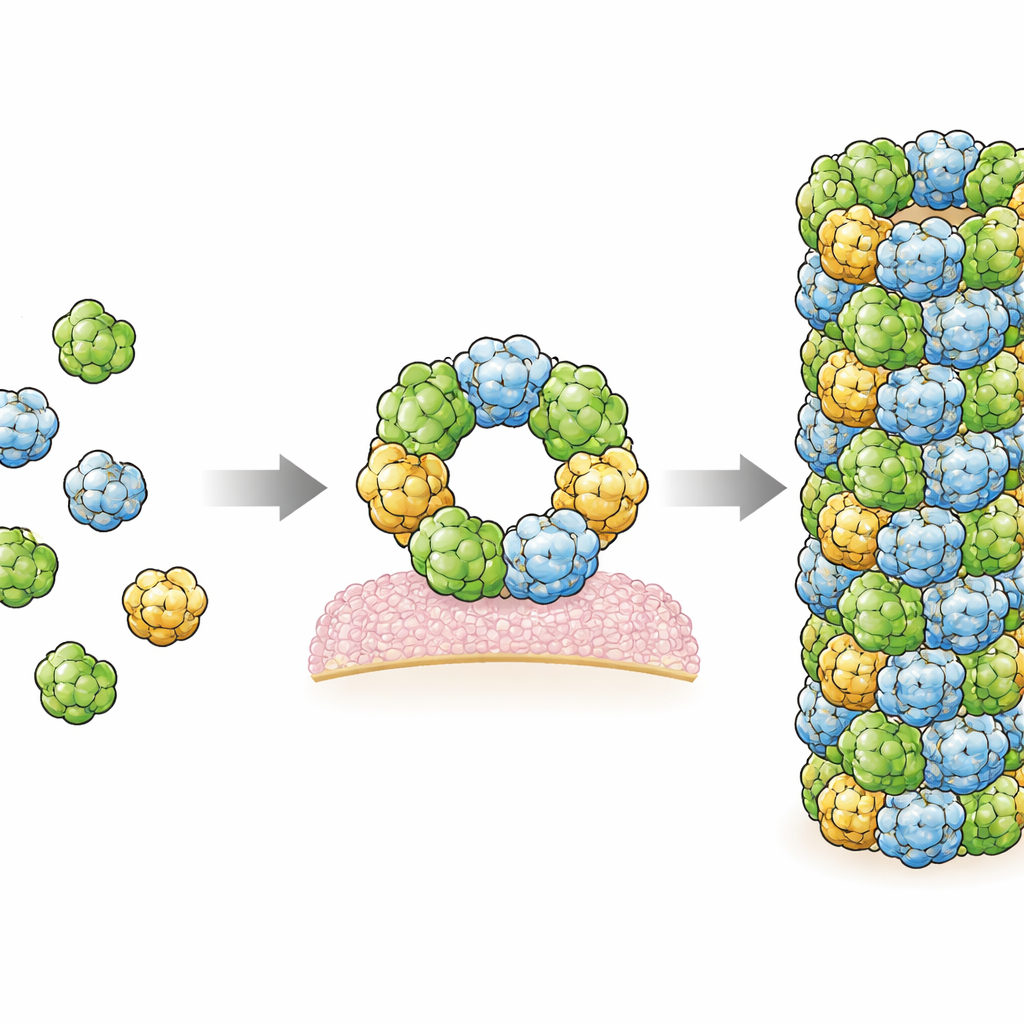
In this study, the authors produced human RegIIIα in bacteria and then triggered it to assemble into filaments under controlled conditions. They used a powerful imaging method called cryo-electron microscopy, which can visualize biological molecules frozen in a thin layer of ice at near-atomic resolution. They found that the filaments are built from repeating sets of three identical RegIIIα units, called trimers, which stack in a regular helical pattern. Each individual RegIIIα molecule looks very similar to what had been seen before in crystal structures, but their orientation within the filament is different from that in an earlier, lower-resolution model. This corrected orientation changes how scientists think the protein interacts with bacterial membranes and with its own regulatory segment.
How Proteins Link Up and Why Lipids Matter
By examining the structure closely, the researchers identified two main contact regions, or interfaces, where neighboring RegIIIα molecules touch and hold the filament together. They then altered specific amino acids at these interfaces and showed that these changes prevented filament formation, confirming that these contact sites are essential. Interestingly, in both interfaces, the cryo-EM maps showed extra fuzzy regions that likely represent small, elongated molecules wedged between protein surfaces. Their shape and chemical environment suggest they may be lipids, the same kinds of oily molecules that make up cell membranes. This supports earlier hints that lipids help promote RegIIIα assembly, possibly by acting like molecular glue that stabilizes the connections between subunits.
The On–Off Switch: pH and a Hidden Segment
The team also investigated what controls when RegIIIα can assemble. Before it becomes active, the protein carries a short “pro-segment” that must be cut off by digestive enzymes. In their high-resolution filament structure, there is no room in the center of the assembly to fit this extra piece, implying that the pro-segment blocks the ability of RegIIIα to form the pore-forming rings and filaments in the first place. They found that the tendency to form filaments also depends on acidity: filaments form readily at a pH similar to that of the small intestine but poorly at a more acidic pH. A single amino acid, histidine 50, appears to act as a pH-sensitive switch; when it is changed to another residue, filament formation becomes much less sensitive to pH. Together, these features—enzyme cutting, lipid binding, and pH—create a layered control system that tunes when and where RegIIIα is active.
What This Means for Gut Health
Overall, this work provides the most detailed picture yet of how RegIIIα molecules assemble into larger structures that relate to both their antibacterial action and their self-limiting behavior. The same interfaces that build the filaments also build the hexameric rings thought to form pores in bacterial membranes, hinting at a delicate balance between a potent killing form and a safer, turned-down state. Mapping disease-linked mutations onto this structure shows that many lie at key contact points, suggesting that disruptions in assembly may underlie certain gut or pancreatic disorders. By clarifying how this natural antibiotic is activated, guided by lipids, and shaped by local pH, the study lays groundwork for future efforts to modulate our innate defenses—either by boosting them against infection or calming them when they become misregulated.
Citation: Han, J., Cao, Q. Structural basis for human RegⅢα filament formation. Commun Chem 9, 117 (2026). https://doi.org/10.1038/s42004-026-01921-y
Keywords: RegIIIα, intestinal immunity, cryo-electron microscopy, protein filaments, antibacterial peptides