Clear Sky Science · en
S9 protease WprP2 catalyzes uniform cleavage on the precursor peptide in RiPP biosynthesis
Why tiny cutting tools in cells matter
Inside bacteria, specialized molecular scissors cut long chains of amino acids into smaller, active pieces that often become antibiotics or other useful natural products. This study uncovers a previously unknown cutting tool—a protease called WprP2—that trims its target peptide with unusual precision. Understanding how this enzyme works could help scientists design new bioactive peptides and next‑generation antibiotics.
From raw chains to finished mini‑medicines
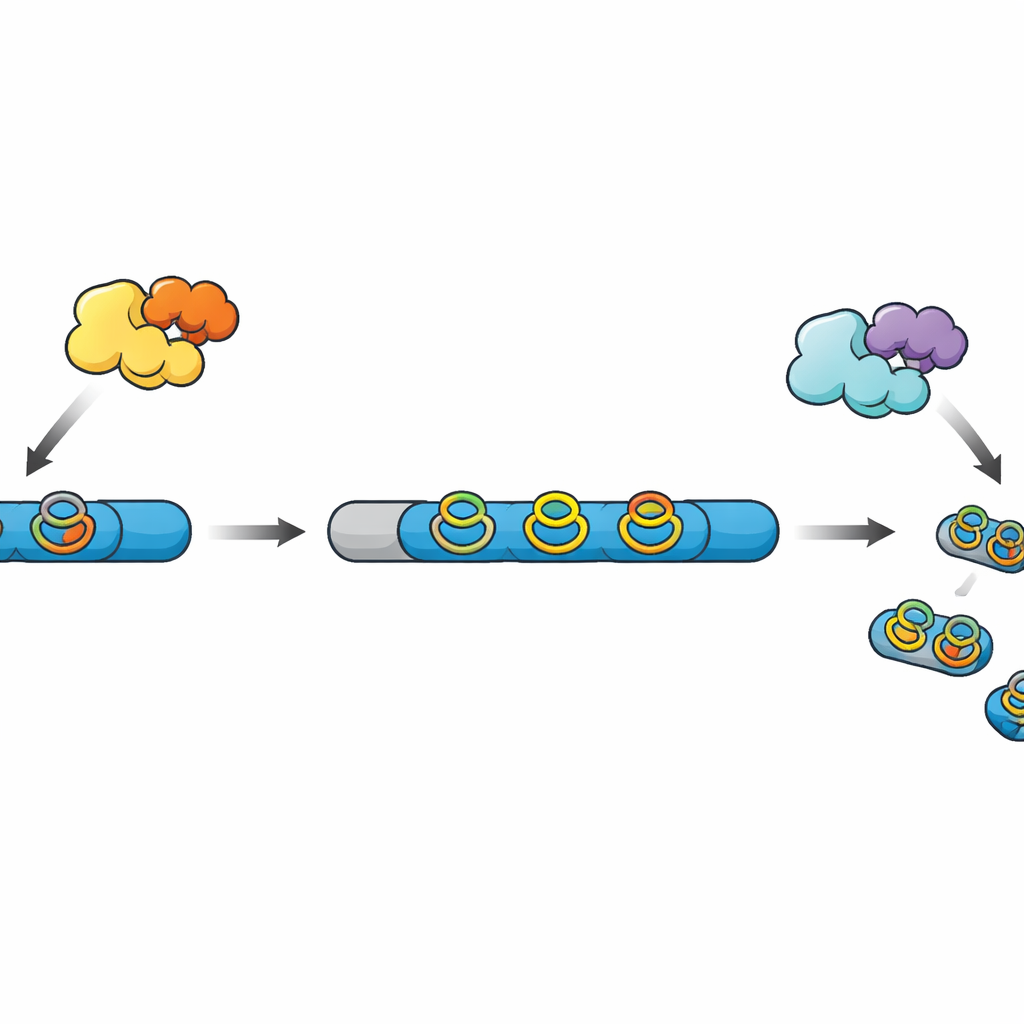
Many modern peptide‑based natural products, known collectively as RiPPs, start as simple chains made on the ribosome. These chains contain a leader region that acts as a handle and a core region that will become the final product after a series of chemical upgrades. Other enzymes add crosslinks and rings that make the peptide more stable and more active, and finally a protease cuts away the leader and releases the mature molecule. In a recently discovered family of RiPPs, radical SAM enzymes build compact “cyclophane” rings involving aromatic amino acids, but in most known gene clusters the matching protease is missing, leaving the final processing steps mysterious.
Tracking down a missing partner enzyme
The authors previously studied a radical SAM enzyme called WprB1 that installed identical crosslinks at three repeating WPR segments in a precursor peptide. In the new work, they searched bacterial genomes for relatives of this enzyme and looked nearby for any genes that could encode a protease. In Streptomyces venezuelae they found a promising cluster that contained a similar crosslinking enzyme, its helper protein, a matching precursor peptide with three WPR repeats, and a candidate serine protease of the S9 family, which they named WprP2. This made the system an ideal testbed for uncovering how the peptide is finally cut to release the active fragments.
Watching the peptide being sliced
To probe WprP2, the team produced the precursor peptide in bacteria, both with and without the crosslinking enzyme, purified it, and then mixed it with the protease in controlled in‑vitro reactions. Mass spectrometry revealed that WprP2 first snips the chain immediately after each WPR segment, and then performs a second cut at a fixed distance before a particular proline residue further along the sequence. These two steps create a series of smaller, uniform fragments. Remarkably, WprP2 carried out this pattern of cuts whether or not the crosslinking enzyme had already installed the cyclophane rings, showing that the rings are not required for recognition. When compared with commercial trypsin, which cannot cut at certain crosslinked sites, WprP2 still functioned, underscoring its unusual robustness.
Decoding what the enzyme recognizes
The researchers next asked what features of the peptide WprP2 actually reads. By systematically changing individual amino acids around the cutting sites, they found that altering many nearby residues had little effect, but changing a key arginine within the WPR segment or the glutamine or proline at the second cutting site could block that particular step. Experiments with a related precursor peptide carrying different flanking residues showed that the protease still cut after WPR repeats but only carried out the second cut when a glutamine‑proline pair was present. Computer‑generated structural models using AlphaFold3 supported this picture: in the predicted complexes, the WPR‑based and glutamine‑proline sites each lined up in turn with the enzyme’s catalytic center, consistent with a sequential two‑step mechanism.
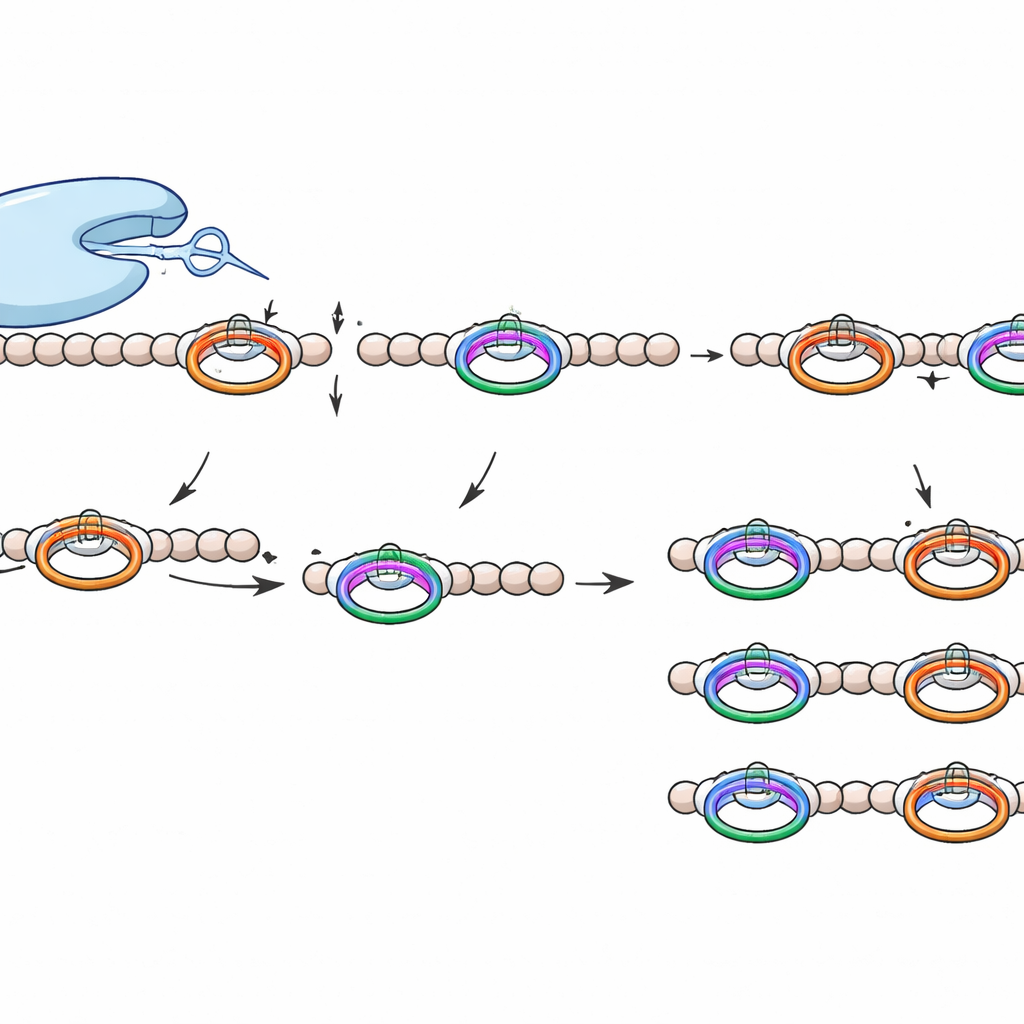
What makes this cutting tool special
WprP2 belongs to a wider S9 family of proteases that already includes medically relevant enzymes, but it stands out in several ways. It cuts on the N‑terminal side of a proline residue rather than after it, it acts on a repeating motif to generate evenly sized fragments across the entire precursor chain, and it shows broad tolerance to the amino acids that lie between its two recognition sites. This combination of precision and flexibility has not been seen before in RiPP‑related proteases.
How this discovery could be used
By revealing a protease that recognizes simple sequence cues and performs uniform, repeated cuts, this work adds a powerful new tool to the growing toolkit for peptide engineering. In practical terms, WprP2 could be harnessed to release or reshape designer peptides, including those with complex crosslinks that resist standard enzymes such as trypsin. For non‑specialists, the key takeaway is that scientists have discovered a new kind of molecular scissors that cuts long peptide chains into precise, evenly sized pieces, opening the door to more controlled production of potential antibiotics and other bioactive molecules.
Citation: Haedar, J.R., Khan, A.H., Ma, S. et al. S9 protease WprP2 catalyzes uniform cleavage on the precursor peptide in RiPP biosynthesis. Commun Chem 9, 108 (2026). https://doi.org/10.1038/s42004-026-01915-w
Keywords: RiPP peptides, serine protease, WprP2, cyclophane natural products, peptide engineering