Clear Sky Science · en
Minimized sample consumption for time-resolved serial crystallography applied to the redox cycle of human NQO1
Watching Enzymes in Action with Fewer Precious Samples
Modern biology often relies on powerful X-ray lasers to see how life’s molecules move and change shape, but these experiments normally devour huge amounts of painstakingly prepared protein. This study introduces a new way to “feed” tiny protein crystals into an X-ray free-electron laser far more efficiently, cutting the material needed by as much as 97%. It also shows that this leaner method can still capture early steps in the activity of a medically relevant human enzyme, opening the door to more routine movies of proteins at work. 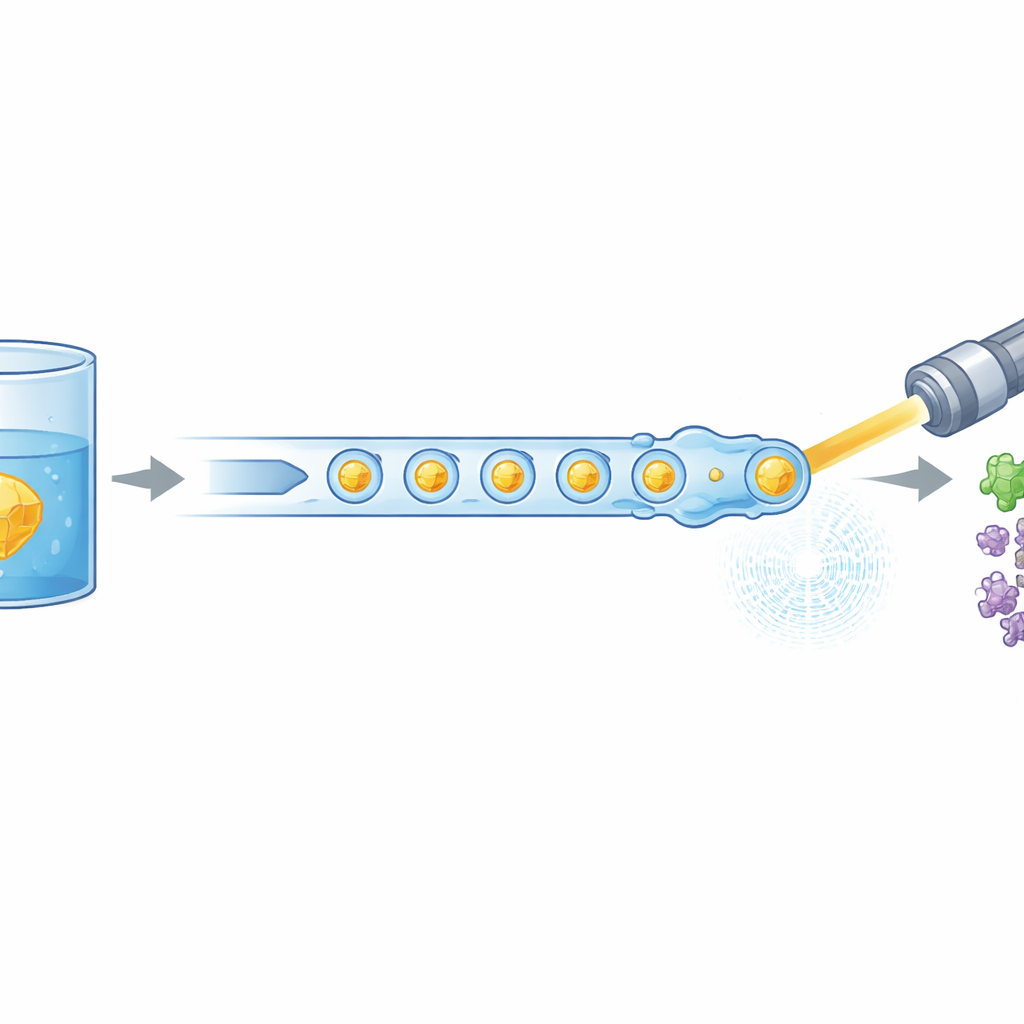
Why Taking Molecular Movies Is So Costly
To understand how proteins actually do their jobs, scientists increasingly turn from static snapshots to time-lapse “movies” that follow reactions in real time. One leading approach, time-resolved serial crystallography, sprays millions of microscopic crystals of a protein across an ultra-bright X-ray beam. Each crystal is hit only once, providing a damage-free glimpse of the structure, and thousands of such images are combined into a full picture. The catch is that every moment along a reaction—tenth of a second, a full second, and so on—requires a fresh batch of crystals. Because protein preparation can be slow and expensive, sample consumption has become a major bottleneck, especially at cutting-edge facilities like the European XFEL, where X-ray pulses arrive in rapid trains at megahertz rates.
A New Way to Deliver Tiny Droplets on Cue
The researchers tackled this problem by redesigning how crystals are delivered to the X-ray beam. Instead of a continuous liquid stream, they generate a carefully spaced train of microscopic droplets, each containing protein crystals, separated by an oil phase. A miniature 3D-printed device combines two streams—protein crystals and a dissolved reaction partner—into tiny mixed volumes, which are then pinched off into droplets. These droplets are guided through a gas-focused nozzle that forms a narrow jet compatible with the vacuum and speed of the XFEL. Crucially, the droplet production is electrically synchronized with the timing of the X-ray pulse trains so that nearly every useful pulse hits a droplet, rather than empty liquid. 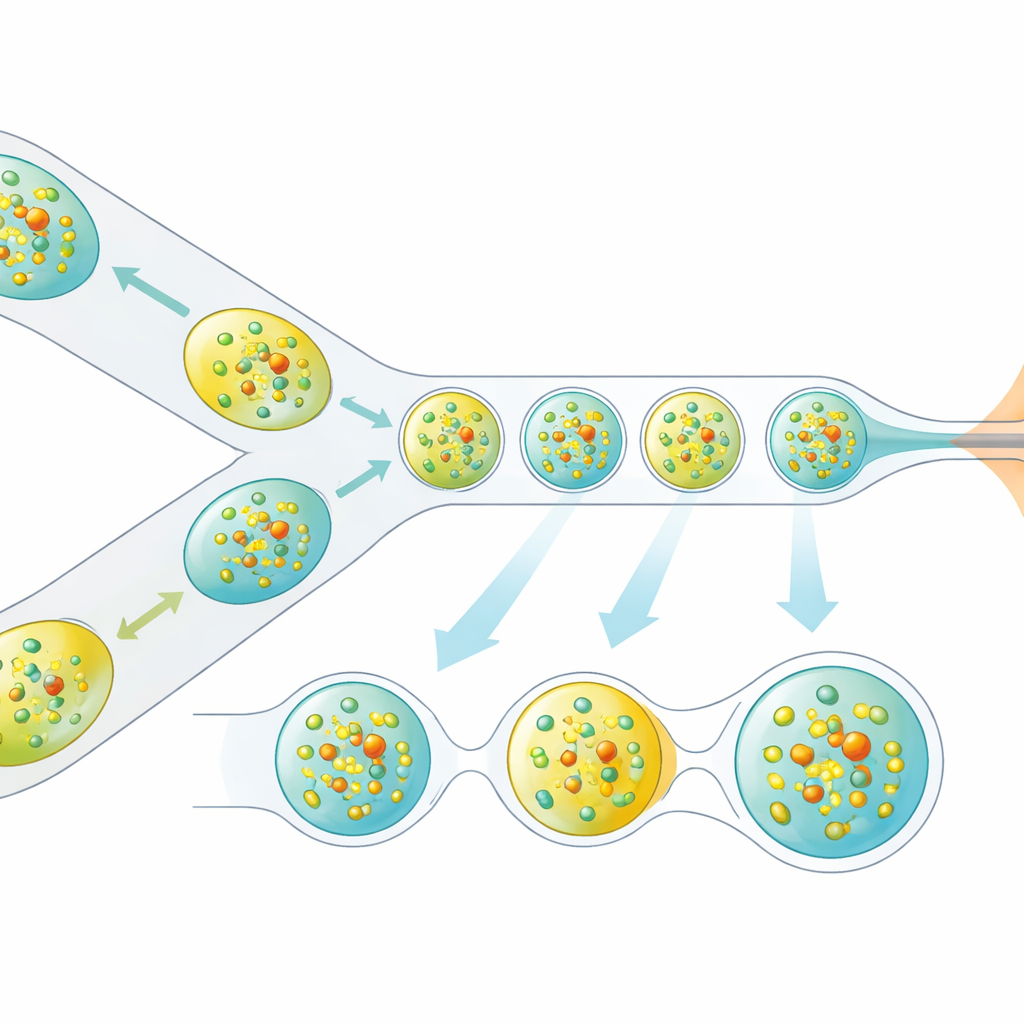
Putting the Method to the Test with a Human Enzyme
To prove that this droplet-based approach works for real biological questions, the team studied NAD(P)H:quinone oxidoreductase 1 (NQO1), a human enzyme involved in managing the cell’s redox balance and linked to disease. They mixed microcrystals of NQO1 with its natural cofactor, NADH, and probed the system at two early time points: 0.3 seconds and 1.2 seconds after mixing. By examining the resulting X-ray diffraction patterns, they reconstructed three-dimensional structures for the enzyme at each delay. Despite using dramatically less protein than conventional continuous-flow experiments, the data quality was sufficient to reveal subtle electron density features in the enzyme’s active sites, consistent with NADH molecules beginning to bind at low occupancy.
What the Early Structural Snapshots Reveal
The structures show that shortly after mixing, not all active sites in the crystal behave the same way. At 0.3 seconds, signs of NADH appear clearly in three out of four active sites in the unit cell, and often in more than one shape, suggesting that the cofactor explores several positions before settling. By 1.2 seconds, some of these features sharpen into a dominant binding mode, but the overall picture remains that of a flexible and only partially occupied interaction. This uneven, shifting behavior aligns with earlier biochemical evidence that NQO1’s two partner units in a dimer do not act in perfect unison, but instead show “half-of-the-sites” activity, where one side of the pair engages first while the other lags behind. The time-resolved data thus offer an early structural glimpse of how this asymmetry plays out in real space.
Saving Sample While Keeping the Science
From a practical standpoint, the segmented droplet method slashed protein use by about sixfold for the 0.3-second measurements and by up to 97% for the 1.2-second experiments, compared with traditional continuous jets at similar conditions. Yet it still delivered reliable structural information at room temperature, compatible with the demanding pulse structure of the European XFEL. For non-specialists, the takeaway is that researchers can now watch enzymes like NQO1 begin their work in near-real time, while using only milligrams of precious sample per time point. This makes it far more realistic to survey many reaction times and many different proteins, ultimately helping to reveal how medically important enzymes move, flex, and cooperate as they carry out the chemistry of life.
Citation: Doppler, D., Grieco, A., Koh, D. et al. Minimized sample consumption for time-resolved serial crystallography applied to the redox cycle of human NQO1. Commun Chem 9, 107 (2026). https://doi.org/10.1038/s42004-026-01908-9
Keywords: serial crystallography, X-ray free-electron laser, droplet microfluidics, enzyme dynamics, NQO1