Clear Sky Science · en
Orthologue inference-based enzyme mining for diversification of the anti-cancer evodiamine scaffold
Turning Plant Chemicals into Better Medicines
Many of today’s medicines began their lives as defensive chemicals in plants. These natural molecules often show powerful effects against cancer, infections, or pain, but they are rarely “perfect” drugs. Chemists would love to tweak their structures in precise ways to make them safer and more effective, yet those molecules are often so complex that even small changes are difficult. This study shows how scientists can enlist plant enzymes as tiny molecular tools to make targeted changes to an anticancer compound called evodiamine, potentially opening new routes to improved therapies.
Why This Plant Molecule Matters
Evodiamine is a natural compound found in the fruits of a tree used in traditional Chinese medicine. It belongs to a family of ring-shaped molecules that already underpin important drugs for cancer, high blood pressure, and infections. Evodiamine itself has shown anticancer, anti-inflammatory, pain-relieving, and antimicrobial properties, and some of its modified versions look especially promising as multitarget cancer leads. The challenge is how to attach new chemical “handles,” such as oxygen-containing groups, at very specific spots on this crowded scaffold without using harsh chemicals or multi-step syntheses.
Letting Enzymes Do the Hard Chemistry
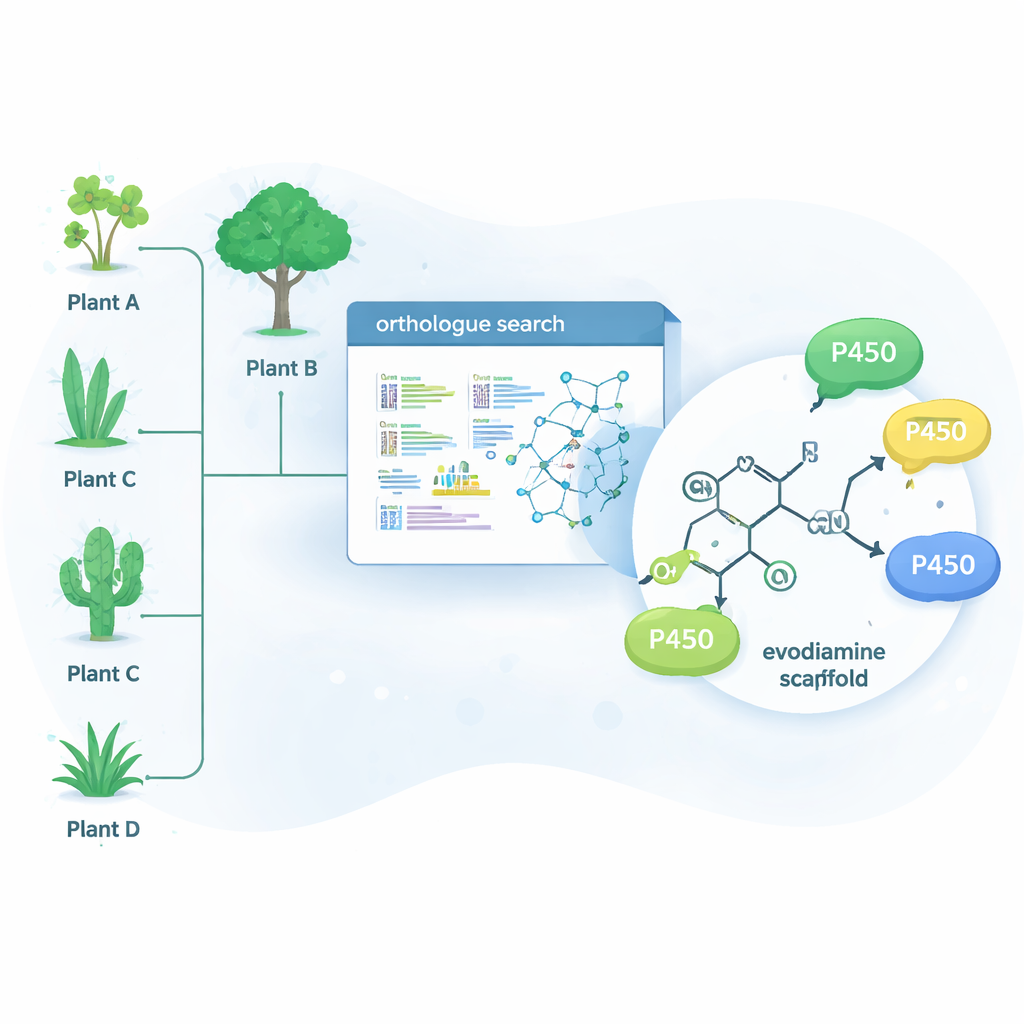
Nature has already solved many of these hard chemistry problems using enzymes—proteins that catalyze specific reactions. One large enzyme family, called cytochrome P450s, excels at adding oxygen to otherwise unreactive carbon–hydrogen bonds. That single step can dramatically change how a molecule behaves in the body and can also create a starting point for further chemical modification. Instead of searching only in plants that naturally make evodiamine, the researchers used a bioinformatics tool called OrthoFinder to scan genetic data from 15 different alkaloid-producing plants. They looked for P450 enzymes that are close “orthologues” of known alkaloid-modifying enzymes, reasoning that relatives of these enzymes might also be able to fine-tune related drug-like molecules.
Finding New Enzymes in Unexpected Plants
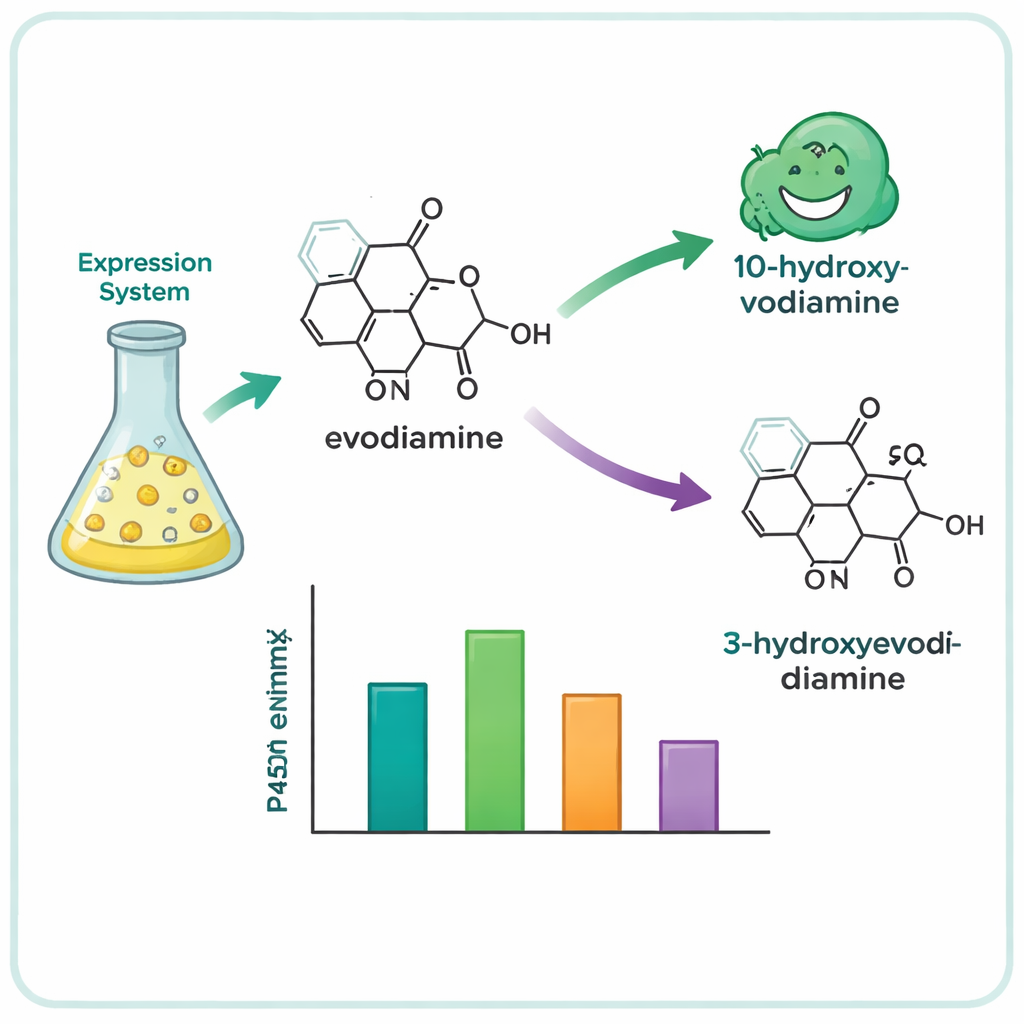
From hundreds of candidate genes, the team narrowed the list to 15 promising P450s and expressed them in yeast cells, which served as miniature enzyme factories. They then fed these yeasts a collection of complex plant molecules and analyzed which ones were chemically altered. Four enzymes—three from the tree Camptotheca acuminata and one from the shrub Tabernaemontana elegans—turned out to act on evodiamine, even though neither plant is known to produce it. These enzymes selectively inserted an oxygen atom at one of two positions on the evodiamine ring system, generating two main products: 10‑hydroxyevodiamine and 3‑hydroxyevodiamine. Such oxygenated versions are easier to further transform into water-soluble or more potent drug candidates, using gentler chemistry than traditional synthetic routes.
Peeking Inside the Molecular Tool Kit
To understand why these closely related enzymes behaved differently, the researchers built three-dimensional models of the most active enzyme and its relatives using modern protein-structure prediction tools. They then used computer docking to see how evodiamine might sit inside the enzymes’ active pockets, near the P450’s iron-containing center where the reaction happens. The models highlighted several bulky, water-repelling amino acids—especially phenylalanine residues—positioned near evodiamine’s aromatic rings. By carefully mutating these positions, the team showed that changing the size and shape of this pocket could weaken activity, alter how well the substrate fits, or even switch the favored oxygenation site from one ring of the molecule to another. In one case, a single mutation flipped the enzyme’s preference from making the 10‑hydroxy product to making the 3‑hydroxy product.
What This Means for Future Medicines
For non-specialists, the key takeaway is that the authors have demonstrated a practical roadmap for discovering and tuning plant enzymes that can perform precise, “surgical” edits on complex drug-like molecules. By combining large-scale genetic mining with enzyme testing and structural modeling, they uncovered a set of biocatalysts that can selectively remodel the evodiamine scaffold at positions that are hard to reach with standard chemistry. This not only offers a cleaner, more sustainable way to make advanced versions of evodiamine—such as water-soluble anticancer leads—but also shows that useful enzymes can be found in plants that never make the target compound themselves. The same strategy could now be applied to many other natural products, accelerating the design of next-generation plant-derived medicines.
Citation: Kwan, B.D., Kim, T., Nguyen, H.H. et al. Orthologue inference-based enzyme mining for diversification of the anti-cancer evodiamine scaffold. Commun Chem 9, 73 (2026). https://doi.org/10.1038/s42004-025-01876-6
Keywords: evodiamine, plant enzymes, cytochrome P450, biocatalysis, drug discovery