Clear Sky Science · en
Optimizing CRISPR precision in mouse embryos via microhomology-mediated end joining-dominant targeting
Why making sharper gene-edited mice matters
Gene-editing tools like CRISPR have made it remarkably easy to create mice that model human diseases, but there’s a hidden problem: the genetic changes in the very first generation of animals are often messy and mixed. That makes experiments slower, less reliable, and uses more animals. This study introduces a way to steer CRISPR cuts in mouse embryos toward highly predictable outcomes, so that most founder mice are born with the same, well-defined mutation—bringing cleaner biology and better ethics to gene-editing research.
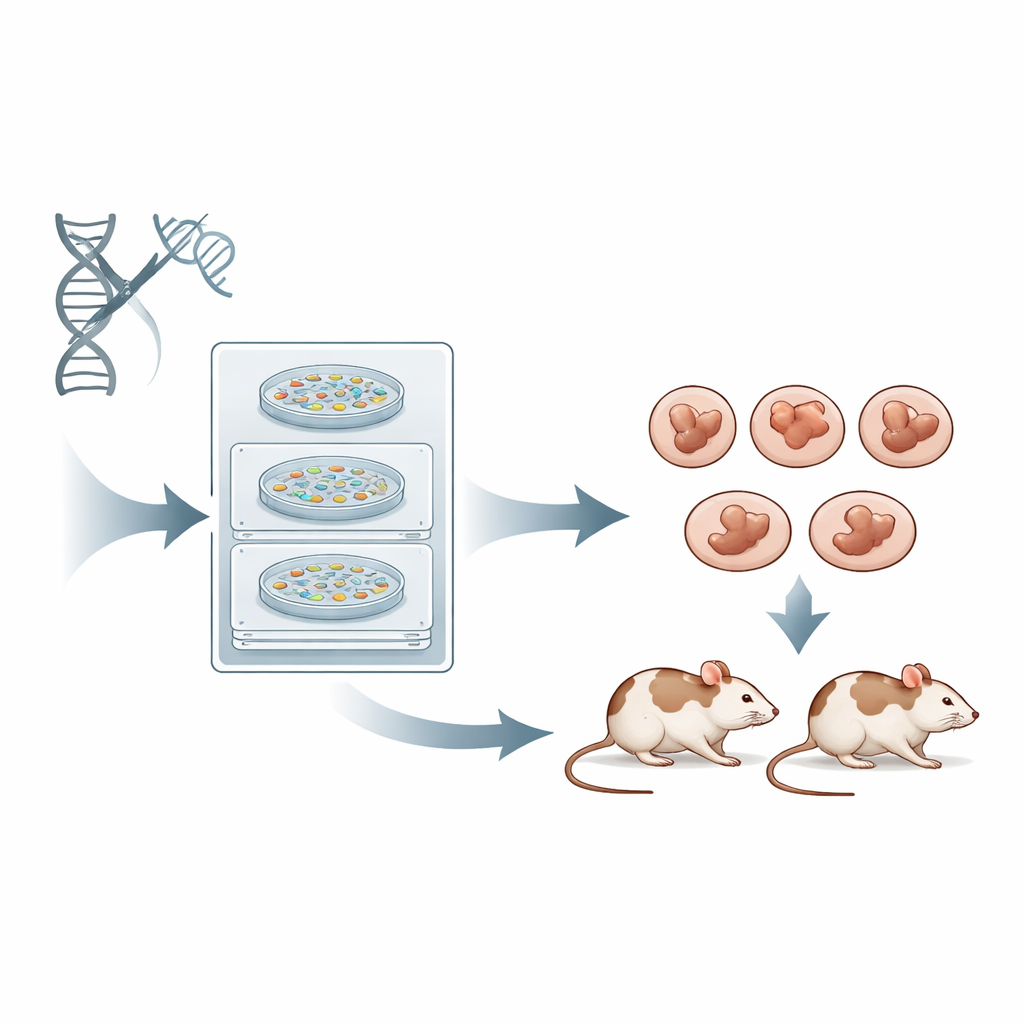
The challenge of messy DNA repairs
When CRISPR cuts DNA, the cell must patch the break using its own repair systems. The most common pathway, called non-homologous end joining, is quick but imprecise, producing a jumble of small insertions and deletions at the cut site. Another pathway, microhomology-mediated end joining, tends to delete chunks of DNA in stereotyped ways using short matching sequences as guides. Both are much more efficient than the precise but sluggish homology-directed route. In standard CRISPR experiments, scientists mostly focus on how strongly a guide RNA can cut and how few off-target sites it hits, and pay much less attention to which repair pathway will be favored or what exact mutation will result. The outcome is that many founder mice carry a patchwork of different mutations in different cells, forcing researchers to breed to the next generation before they can work with a clean, uniform genotype.
A smarter way to pick CRISPR guides
The authors set out to flip this script by designing guides not only for strength and safety, but also for predictability. They began with inDelphi, a machine-learning tool trained on huge datasets of CRISPR-induced mutations in cultured cells. inDelphi does not merely say how often a site will be edited; it predicts the full menu of possible insertions and deletions and how frequently each one will appear, with special attention to microhomology-driven events. The team scanned the mouse tyrosinase (Tyr) gene, where loss of function makes animals albino, and selected guide RNAs predicted to favor strong, repeatable microhomology-mediated deletions while keeping off-target risks low. They then edited mouse embryos and measured the resulting mutations with deep sequencing. Overall, inDelphi’s favorite genotype for each guide appeared at similar frequencies in embryos as predicted, and guides with stronger microhomology features indeed produced more uniform mutation patterns.
Using stem cells as a rehearsal stage
Yet prediction alone was not enough. When the team compared inDelphi forecasts with actual editing patterns, they found only moderate agreement. To bridge this gap, they introduced a practical intermediate step: testing each guide in mouse embryonic stem cells that share many features with very early embryos. After transfecting these cells with CRISPR components, they sorted edited cells and sequenced the target sites. The mutation patterns in stem cells matched those in embryos much more closely than the computer model did. Guides that produced a single dominant deletion in stem cells typically did the same in blastocysts and later-stage embryos. By combining inDelphi’s ranking with this stem-cell “dress rehearsal,” the researchers could reliably pick guides that drive microhomology-mediated repair and minimize the diversity of mutant alleles.
From eye color to missing limbs
The authors put their pipeline to the test in living animals. For the Tyr gene, they chose three guides representing high, medium, and low predicted precision and transferred edited embryos into foster mothers. At day 11.5 of development, they examined eye pigmentation and sequenced each embryo individually. The highly microhomology-favoring guide produced embryos that were mostly albino and carried one dominant small deletion, often in both copies of the gene, with very little variation. A less-optimized guide produced a mixture of pigment loss and partial pigmentation tied to a more complex set of mutations. They then applied the same approach to the Fgf10 gene, where loss of function yields limbless embryos. Selecting a guide predicted—and confirmed in stem cells—to give a specific four-base deletion with a high chance of disrupting the gene, they generated embryos at day 15.5 that were uniformly limbless and carried a strongly enriched set of the expected deletions. Across both genes, the same few mutation types dominated in inDelphi predictions, stem cells, early embryos, and later-stage embryos.
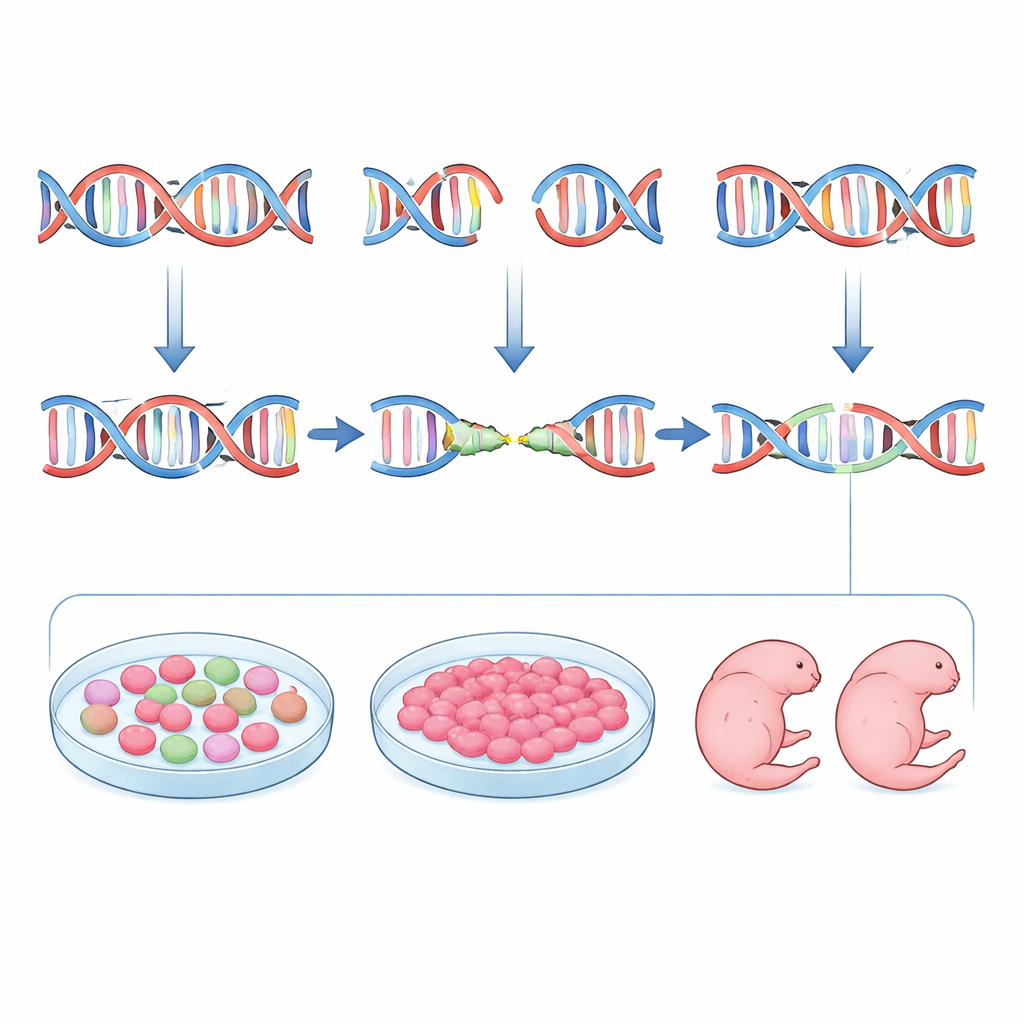
Cleaner genetics with fewer animals
In practical terms, the study offers a new template for designing CRISPR experiments in mice. Instead of rushing directly from a computer-designed guide to embryo editing, the authors advocate an integrated pipeline: use inDelphi and off-target tools to pick guides that are likely to favor microhomology-mediated deletions and frameshifts, test those guides in embryonic stem cells to confirm both efficiency and uniformity of mutations, and advance only the best performers into embryo work. This strategy yields founder mice whose cells overwhelmingly share the same, well-characterized mutation, making them immediately useful for modeling human diseases—especially those caused by recurrent deletion-type changes—while cutting down on the number of animals that must be bred and screened. The result is sharper, more reproducible genetics and a more ethical path to powerful disease models.
Citation: Lkhagvadorj, K., Okamura, E., Taki, T. et al. Optimizing CRISPR precision in mouse embryos via microhomology-mediated end joining-dominant targeting. Commun Biol 9, 371 (2026). https://doi.org/10.1038/s42003-026-09771-z
Keywords: CRISPR, mouse models, genome editing, DNA repair, disease modeling