Clear Sky Science · en
Structural basis of Neisseria meningitidis quinol dependent nitric oxide reductase activation by dimerization
How Bacteria Outsmart Our Immune Defense
When harmful bacteria invade the body, our immune cells unleash toxic molecules such as nitric oxide to kill them. Some pathogens, however, have evolved molecular tricks to neutralize this attack and survive. This study reveals, in atomic detail, how a key bacterial enzyme reconfigures itself into pairs to become far better at detoxifying nitric oxide, and why understanding this shape-shifting behavior could open new paths to antibiotic design.
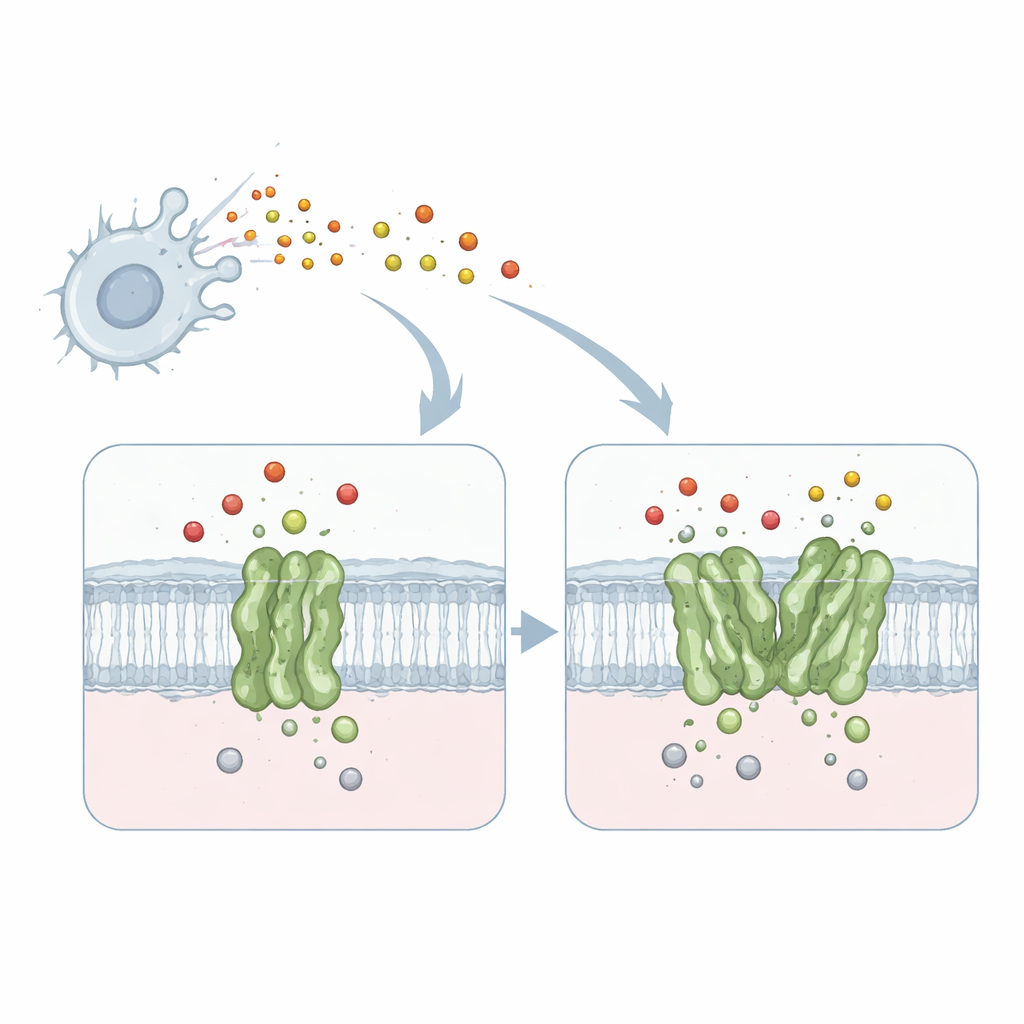
A Tiny Machine That Disarms a Toxic Gas
The work focuses on a membrane protein called quinol-dependent nitric oxide reductase, or qNOR, from the meningitis-causing bacterium Neisseria meningitidis. qNOR sits in the bacterial cell membrane and converts nitric oxide—a toxic gas produced by our immune system—into less harmful products. This detoxification allows the microbe to keep breathing and growing even inside hostile environments such as human macrophages. Because respiration and nitric oxide removal are vital for the pathogen’s survival, qNOR is an attractive target for new antimicrobial drugs.
When One Is Good but Two Are Better
Previous biochemical studies showed that qNOR can exist either as a single unit (a monomer) or as a pair (a dimer), and that the dimer works two to four times faster at neutralizing nitric oxide. Until now, it was unclear why simply pairing two copies of the same enzyme had such a dramatic impact on performance. To answer this, the researchers used high-resolution single-particle cryo–electron microscopy to capture detailed three-dimensional structures of both monomeric and dimeric qNOR from the same bacterial strain. They obtained near-atomic resolution views—1.89 angstroms for the dimer and 2.25 angstroms for the monomer—sharp enough to locate individual amino-acid side chains, metal ions, and many water molecules.
A Flexible Helix Becomes a Stabilizing Brace
Surprisingly, the catalytic heart of the enzyme—the metal center where nitric oxide is converted—looked almost identical in both forms. Instead, the key differences appeared some distance away, in a transmembrane helix called TM10 and in how it contacts another helix (TM2) when two qNOR molecules pair up. In the monomer, TM10 is loosely anchored and can sway and bend substantially, as revealed by computational analysis of the cryo-EM data. In the dimer, TM10 from each partner locks against its neighbor to form a four-helix bundle, greatly restricting its motion. This stabilization subtly repositions a crucial amino acid, a glutamate named Glu563, bringing it close to another glutamate (Glu494) near the active site and shaping a more focused route for incoming protons needed for the reaction.
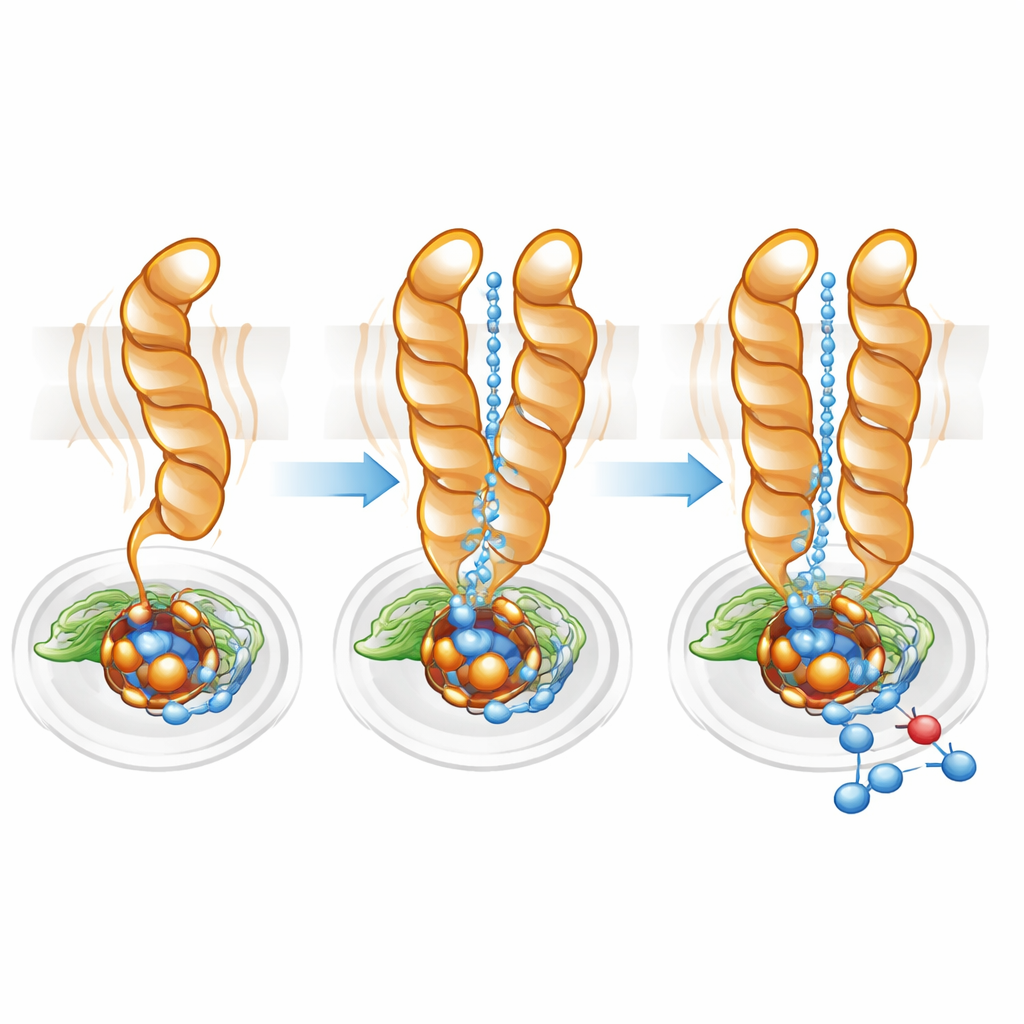
Fine-Tuning the Proton Highway
Protons—positively charged hydrogen ions—must be delivered efficiently from the cell interior to the buried active site for nitric oxide reduction to proceed. Using pathway-mapping software, the team found a hydrophilic cavity connecting the cytoplasm to the active site in both monomer and dimer. However, in the monomer this channel was wider and more diffuse, consistent with the greater mobility of TM10. In the dimer, TM10’s locked position and the favorable orientation of Glu563 helped define a more direct, better organized pathway. Mutating Glu563 to amino acids that cannot form the same interactions slashed the enzyme’s activity to under 10% of normal and also destabilized the dimer, while mutations at related positions reduced the amount of essential non-heme iron at the active site. Together, these results show that dimer formation and the precise positioning of Glu563 and Glu494 are tightly linked to both structural integrity and catalytic efficiency.
Turning Structural Insights into New Treatments
The findings reveal a clear line from protein pairing at the membrane surface to fine adjustments deep inside the active site that control how effectively qNOR removes nitric oxide. In simple terms, when two copies of the enzyme clasp together, they straighten and stiffen a flexible segment that helps steer protons into the reaction center, boosting the enzyme’s output. For drug development, this suggests an unconventional strategy: instead of blocking the active site directly, one could design molecules that pry the dimer apart or disrupt the Glu563–Glu494 interaction, forcing the enzyme into its sluggish monomeric state. Since qNOR and related enzymes are crucial for the survival of several dangerous, drug-resistant pathogens, such structure-guided approaches could contribute to the next generation of targeted antimicrobials.
Citation: Gopalasingam, C.C., Egami, H., Shigematsu, H. et al. Structural basis of Neisseria meningitidis quinol dependent nitric oxide reductase activation by dimerization. Commun Biol 9, 433 (2026). https://doi.org/10.1038/s42003-026-09754-0
Keywords: nitric oxide detoxification, bacterial respiration, membrane enzymes, cryo electron microscopy, antimicrobial drug targets