Clear Sky Science · en
Ultrafast and specific miRNA quantification via single-molecule fluorescence quenching kinetics
Why Speedy Disease Tests Matter
Diagnosing diseases like cancer or viral infections often comes down to spotting tiny traces of genetic material in blood or other body fluids. Today’s gold-standard tests can be highly accurate but may be slow, expensive, or struggle to detect very rare signals. This study introduces a new microscope-based method, called Q‑FISH, that can read out these genetic clues in less than a second. If translated into clinical tools, such speed and precision could make it easier to catch diseases earlier, track how treatments are working, and tailor care to each patient.
A New Way to See Tiny Genetic Messages
The work focuses on microRNAs—short snippets of RNA that help control how our genes are used and that are strongly linked to many cancers, heart disease, infections, and brain disorders. Because microRNAs are so small and often differ by only one or two building blocks, standard methods like PCR and next‑generation sequencing can have trouble telling closely related types apart, especially when they are present at very low levels. Recent single‑molecule imaging approaches improved this specificity by tracking how individual probes bind and unbind, but they were still relatively slow, taking around ten minutes to analyze a single target.
Watching Light Flicker to Find the Right Target
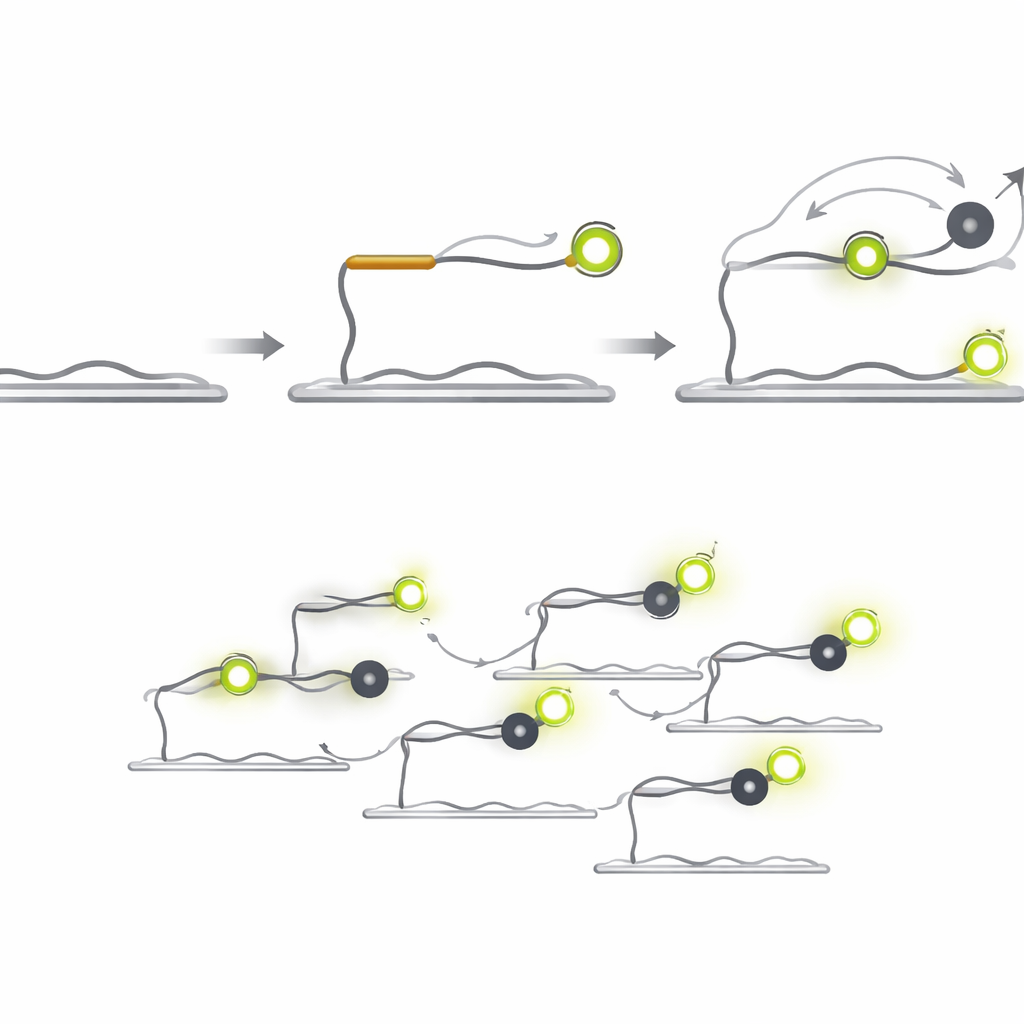
Q‑FISH turns the problem on its head by exploiting the brief flickers of light from single molecules. The method uses two short DNA probes that recognize neighboring spots on a target microRNA. One probe carries a fluorescent dye that glows under a laser, while the other carries a “quencher” that soaks up the light when it comes close. The fluorescent probe binds to the target and stays on long enough to be seen, whereas the quencher probe is designed to bind and fall off very quickly. Each time the quencher lands next to the dye, the light suddenly drops; when it leaves, the light jumps back up. By recording these rapid on‑off flashes from individual molecules and analyzing how long the bright and dim periods last, the system can decide whether a real target is present.
From Minutes to Milliseconds
Because the quencher probe does not emit its own light, it can be used at much higher concentrations than fluorescent probes without creating background glare. Shortening the quencher probe increases how quickly it falls off, and boosting its concentration increases how often it lands. Together, these design choices lead to a dramatic jump in speed. In tests with a cancer‑related microRNA called let‑7a, Q‑FISH reached more than 70% of its maximum detection efficiency in just one second of observation. Comparable single‑molecule methods needed tens to hundreds of seconds to achieve similar performance, making Q‑FISH over 600 times faster in practical terms.
Sorting Nearly Identical Signals and Measuring Real Samples
The researchers also showed that Q‑FISH can distinguish between several members of the let‑7 microRNA family, which share almost the same sequence but have different roles in controlling cancer‑linked genes. They used two multiplexing strategies. In one, different quencher probes were introduced one after another, each tuned to a slightly different microRNA; in another, the probes were labeled with different colors and imaged at the same time. In both cases, reading out the blinking patterns allowed the team to correctly identify which microRNA was which within about one second. Finally, they applied the method to total RNA extracted from human liver and lung tissues. By adding known amounts of synthetic microRNAs and counting the resulting spots, they built calibration curves and then read off the natural levels present in the tissues, revealing clear differences between organs.
What This Could Mean for Future Tests
Taken together, the study shows that Q‑FISH can pick out specific microRNAs with very high accuracy, even when they are almost identical to one another, and can do so at speeds that greatly surpass earlier single‑molecule methods. Although the experiments were done on prepared samples using a specialized microscope, the underlying idea—using fast light‑quenching events instead of slow binding alone—could be adapted to many kinds of genetic markers, including tumor DNA fragments in blood. With further engineering and streamlined sample preparation, this approach could help bring rapid, highly multiplexed, and extremely sensitive molecular diagnostics closer to everyday clinical use.
Citation: Kim, J., Hohng, S. Ultrafast and specific miRNA quantification via single-molecule fluorescence quenching kinetics. Commun Biol 9, 432 (2026). https://doi.org/10.1038/s42003-026-09714-8
Keywords: microRNA detection, single-molecule imaging, molecular diagnostics, fluorescence quenching, liquid biopsy