Clear Sky Science · en
Structural basis for bat receptor recognition by SARS-CoV-2 and bat SARS2-like coronaviruses
Why this bat–virus story still matters
The COVID-19 pandemic began more than six years ago, yet scientists are still piecing together how the virus that causes it, SARS-CoV-2, became so good at infecting people. A central puzzle is how this virus, and its close cousins found in bats, latch onto a protein called ACE2 on the surface of cells—a first and crucial step for infection. Understanding these microscopic lock-and-key interactions can clarify where the virus came from, how it adapted, and what might happen if related viruses cross into humans in the future. 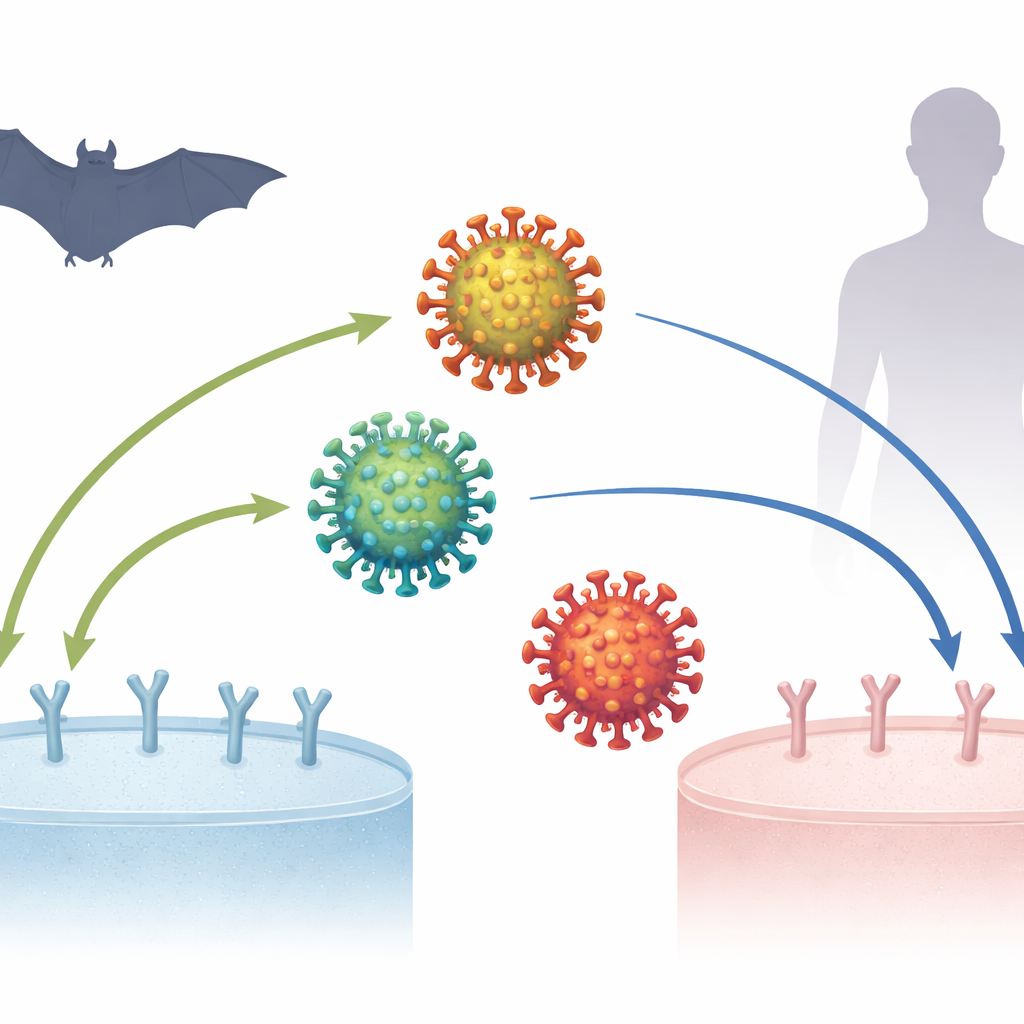
Looking at the viral handshake
The study focuses on the spike protein, the crown-like structure on the virus surface that grabs ACE2. A small region of the spike, the receptor-binding domain (RBD), acts like a fingertip that grips the ACE2 “doorknob.” Earlier work showed that when coronaviruses jump between species—say, from bats to civets to humans—the RBD often picks up mutations that improve its grip on the new host’s ACE2. But SARS-CoV-2 posed a riddle: its original form already bound human ACE2 extremely well, even before much time had passed for adaptation, and some bat viruses related to SARS-CoV-2 seemed to bind human ACE2 better than bat ACE2. This led some to wonder whether SARS-CoV-2 might be an exception to the usual evolutionary rules.
Putting bat and human receptors to the test
To probe this mystery, the researchers examined two bat coronaviruses closely related to SARS-CoV-2, known as BANAL-52 and BANAL-236. They compared how strongly the RBDs of these viruses and SARS-CoV-2 attached to ACE2 from humans and from several bat species. Using cell-based binding assays, high-precision biosensor measurements, and infection tests with harmless “pseudoviruses,” they found a clear pattern. BANAL-52’s RBD bound most strongly to ACE2 from one bat species, Rhinolophus sinicus, and slightly less strongly to human ACE2. SARS-CoV-2’s RBD, in contrast, showed a modest preference for human ACE2 over bat ACE2. Overall, BANAL-52’s RBD gripped both bat and human receptors more tightly than SARS-CoV-2’s, but it was most finely tuned to that particular bat ACE2. 
How a single atom-scale tweak tips the balance
The team then turned to X-ray crystallography, which reveals atomic-level structures, to see exactly how the viral “finger” and the ACE2 “doorknob” touch. They focused on two key positions: one on the viral RBD (called residue 498) and one on ACE2 (residue 41). In BANAL-52, both the viral site and the bat ACE2 site use the same building block, histidine, which allows a very snug, layered interaction—stacking like two coins and also forming a hydrogen bond. In human ACE2, the same spot carries a related but slightly different building block, tyrosine, which still stacks well with BANAL-52’s histidine but lacks the extra hydrogen bond. SARS-CoV-2 instead uses glutamine at this viral position, which cannot stack in the same way, resulting in weaker binding to both bat and human ACE2. By deliberately swapping these amino acids in bat ACE2, the researchers confirmed that strengthening or weakening this one contact point could flip which virus or host was favored.
Why human ACE2 is such a welcoming doorway
Beyond this single contact, the authors asked why human ACE2 in general is such an efficient gateway for coronaviruses. Comparing bat and human ACE2 side by side, they pinpointed several human-specific features that improve the viral handshake. Two positions on human ACE2, containing the amino acids histidine (at site 34) and methionine (at site 82), create stronger hydrogen bonds and greasy “patches” that help the viral RBD nestle in more firmly. Another human residue, threonine at site 27, actually makes binding a bit weaker than in the bat version, but overall the added favorable contacts win out. These details fit with earlier work showing that human ACE2 has multiple “hotspots” that naturally make it an attractive receptor for a wide range of coronaviruses.
Reframing the origin story
Putting all the data together, the study argues that SARS-CoV-2 and its bat relatives still follow the usual evolutionary script. BANAL-52’s spike appears best adapted to ACE2 from certain bats, while also being compatible with human ACE2. SARS-CoV-2’s spike, in turn, is better tuned to human ACE2 than to bat ACE2, helped by special contact points on the human receptor. Small changes at just a few positions—particularly at the viral residue 498 and neighboring sites—can shift the balance of which host is favored. For non-specialists, the takeaway is that there is no need to invoke exotic explanations: standard, well-understood structural principles of protein–protein interaction are enough to explain how these viruses recognize bat and human cells, and they support a close evolutionary link between SARS-CoV-2 and related bat coronaviruses.
Citation: Hsueh, FC., Shi, K., Aihara, H. et al. Structural basis for bat receptor recognition by SARS-CoV-2 and bat SARS2-like coronaviruses. Commun Biol 9, 398 (2026). https://doi.org/10.1038/s42003-026-09682-z
Keywords: SARS-CoV-2 evolution, bat coronaviruses, ACE2 receptor, virus host range, spike protein binding