Clear Sky Science · en
DANST enables cell-type deconvolution in spatial transcriptomics using deep domain adversarial neural networks
Seeing Cells in Their Neighborhoods
Human tissues are crowded cities of many different cell types, each playing its own role. New “spatial transcriptomics” technologies can measure which genes are active across a tissue slice, but each measurement often mixes signals from several neighboring cells. This paper introduces DANST, a smart computational method that teases apart those mixtures. By telling us which cell types are present, and where, in organs and tumors, it helps scientists better understand how tissues are built, how diseases spread, and where treatments might best be aimed. 
The Challenge of Untangling Cell Crowds
Modern gene-reading tools can either look very precisely at single cells or capture the full layout of a tissue, but rarely both at once. Popular spatial technologies record gene activity in relatively large “spots” that can cover several cells. The result is like hearing a choir without being able to tell who sings which notes. To make sense of this, researchers need “deconvolution” methods that estimate how much each cell type contributes to every spot. Many existing approaches use single-cell data as a reference, but they struggle because the two data types are collected by different experiments and do not perfectly match in quality, noise, or resolution.
Building a Bridge Between Data Worlds
DANST tackles this mismatch by creating a bridge between single-cell data and spatial data. First, it uses the detailed single-cell profiles to simulate many artificial mixed spots with known cell-type proportions. At the same time, it groups the real spatial spots according to both their location in the tissue and their gene patterns, and uses the distances to these groups to assign each simulated spot a “pseudo” position. This step creates a linked map where artificial and real spots share a common spatial frame, allowing the method to learn how mixed signals should look in specific neighborhoods of the tissue. 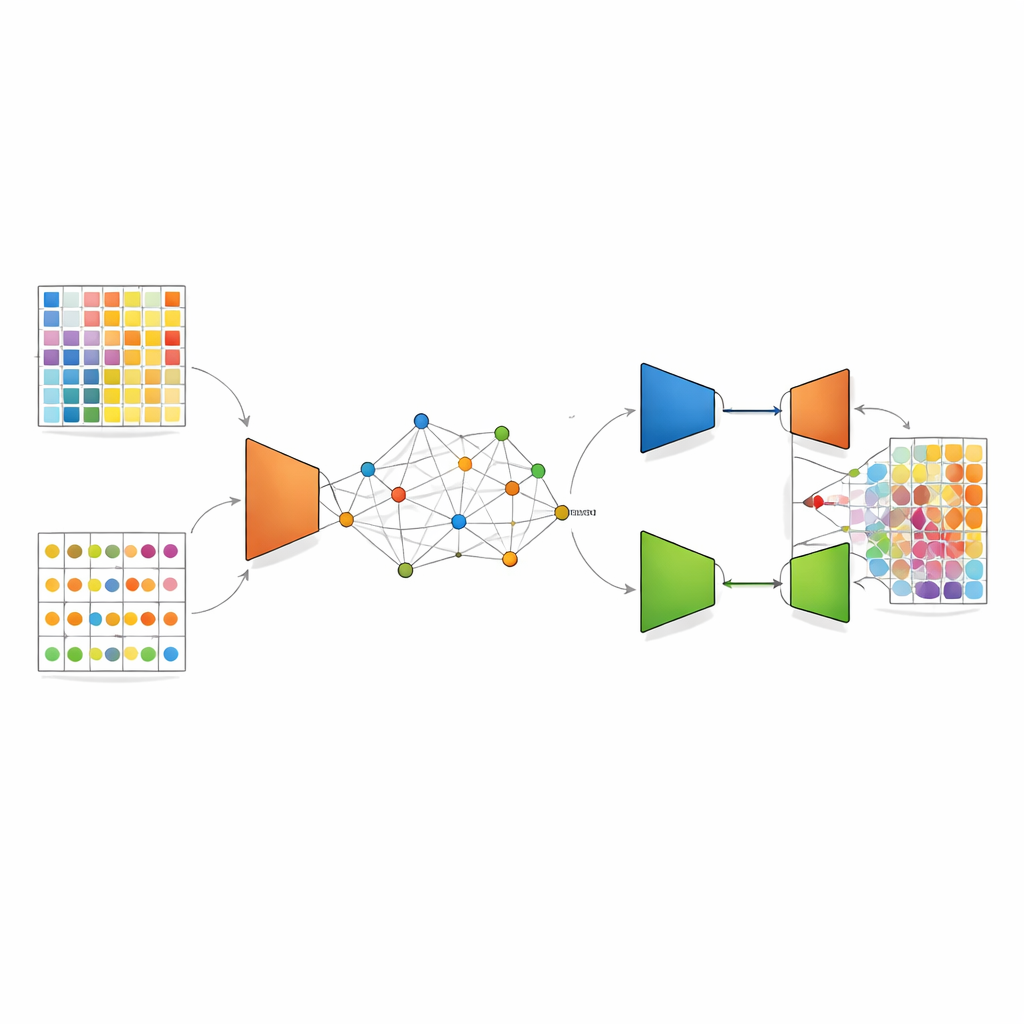
Cleaning Signals and Aligning Domains
Once this joint map is in place, DANST applies a type of deep learning called a variational autoencoder. This network compresses gene patterns from both real and simulated spots into a refined internal representation and then tries to reconstruct them, effectively denoising the data and emphasizing important features. On top of this, the authors add an adversarial component: a second network tries to tell whether a refined pattern came from real spatial data or simulated data, while the feature extractor learns to fool it. This “tug of war” pushes the model toward features that work well for both data sources, so that knowledge gained from simulated spots with known cell-type fractions can be reliably transferred to real tissue spots whose composition is unknown.
Testing on Hearts, Brains, and Tumors
The team tested DANST on both artificial benchmarks and real biological samples from mice and humans. Compared to several leading methods, DANST more accurately recovered the proportions of cell types in synthetic datasets and held its advantage across very different tissues and platforms. In a mouse brain dataset, it clearly reconstructed the layered organization of the cortex and matched expert-defined anatomical regions. In a separate mouse brain slice, it captured fine patterns in areas such as the hippocampus. Most notably, in human breast cancer tissue, DANST traced how various immune cells, supporting cells, and hormone-sensitive luminal cells are arranged within and around tumor regions. These maps aligned with known biology and suggested clinically relevant features, such as hormone dependence and potentially poorer prognosis where certain immune cells were scarce.
What This Means for Biology and Medicine
To a non-specialist, DANST can be viewed as a powerful interpreter that turns blurry, overlapping signals into a clear picture of which cells live where in a tissue. By reliably separating cell types in space, it gives researchers a sharper view of how healthy organs are organized and how disease reshapes that organization. In cancer, this can reveal how tumor cells and immune cells interact in specific regions, which may guide targeted therapies and help predict patient outcomes. As more spatial and single-cell datasets become available, tools like DANST are poised to become essential for decoding the cellular neighborhoods that underlie health and disease.
Citation: Zhang, X., Wu, Z., Wang, T. et al. DANST enables cell-type deconvolution in spatial transcriptomics using deep domain adversarial neural networks. Commun Biol 9, 388 (2026). https://doi.org/10.1038/s42003-026-09659-y
Keywords: spatial transcriptomics, cell type deconvolution, deep learning, tumor microenvironment, single-cell RNA sequencing