Clear Sky Science · en
Naturally occurring dinactin targets cpsA protein and kills Mycobacterium tuberculosis by disrupting the proton motive force
A New Hope Against a Persistent Lung Killer
Tuberculosis remains one of the deadliest infectious diseases on the planet, and strains that resist multiple antibiotics are spreading. This study introduces a naturally occurring compound called dinactin, produced by soil bacteria, that can kill Mycobacterium tuberculosis, including drug‑resistant and dormant forms that are especially hard to eliminate. By probing how dinactin works and how it cooperates with existing medicines, the researchers outline a promising new strategy to shorten and strengthen TB treatment.
Finding a Hidden Weapon in Nature
To search for new TB drugs, the team screened more than 6,000 natural extracts from plants and microorganisms, testing their ability to stop the growth of TB bacteria in whole cells rather than in isolated enzymes. Among the many candidates, dinactin stood out. It belonged to a family of ring‑shaped molecules known as macrotetrolides and showed strong activity against standard laboratory TB strains at very low doses, without damaging red blood cells. When the scientists compared dinactin with closely related molecules, dinactin proved to be both more potent and more selective, making it the best candidate for deeper investigation. 
Powerful Action in Tough‑to‑Reach Bacteria
TB bacteria can hide in several hard‑to‑treat states: actively dividing in the lungs, lying dormant with slowed metabolism, or sheltering inside immune cells. Dinactin attacked them in all three conditions. It killed actively growing TB with a steep drop in viable bacteria and was also effective against nutrient‑starved, non‑replicating cells that often survive standard therapy. In a human macrophage model, dinactin penetrated the host cells and reduced the number of internalized TB bacteria by about one hundredfold. In infected wax moth larvae, an in vivo model, dinactin alone improved survival and lowered bacterial burden, and its benefits were even greater when combined with existing TB drugs.
Teaming Up With Existing Medicines
Because TB treatment relies on drug combinations, the researchers tested how dinactin interacted with current antibiotics such as rifampicin, isoniazid, bedaquiline, and others. Using checkerboard assays, they found that dinactin strongly boosted the effects of most of these drugs, especially rifampicin and isoniazid: adding dinactin allowed much lower doses of these standards to work effectively. Notably, when applied to clinical multidrug‑resistant TB isolates, dinactin restored much of their sensitivity to rifampicin and isoniazid. In stationary‑phase cultures that mimic persistent infection, combinations of dinactin with rifampicin or isoniazid killed many more bacteria than any drug alone, suggesting that dinactin‑based cocktails could help clear stubborn infections more quickly.
How Dinactin Undermines Bacterial Energy
To understand how dinactin kills TB bacteria, the team examined its effects on the cell envelope and energy systems. Dinactin acts as an ion carrier, shuttling potassium and sodium across the bacterial membrane. This extra ion flow makes the membrane leakier and more fluid, visibly wrinkling the bacterial surface and allowing dyes to penetrate more easily. By moving ions, dinactin collapses the proton motive force—the electrical and chemical gradient that bacteria use like a tiny battery to fuel ATP production. Measurements showed that both components of this gradient, the voltage across the membrane and the proton difference, were lost after dinactin treatment. As a result, ATP levels inside the cells plunged, even though their oxygen consumption machinery kept running, indicating that energy production had been uncoupled from respiration. Dinactin also disturbed the balance between the reduced and oxidized forms of a key metabolic cofactor (NADH/NAD+) and triggered a burst of reactive oxygen species inside the bacteria, further damaging cellular components. 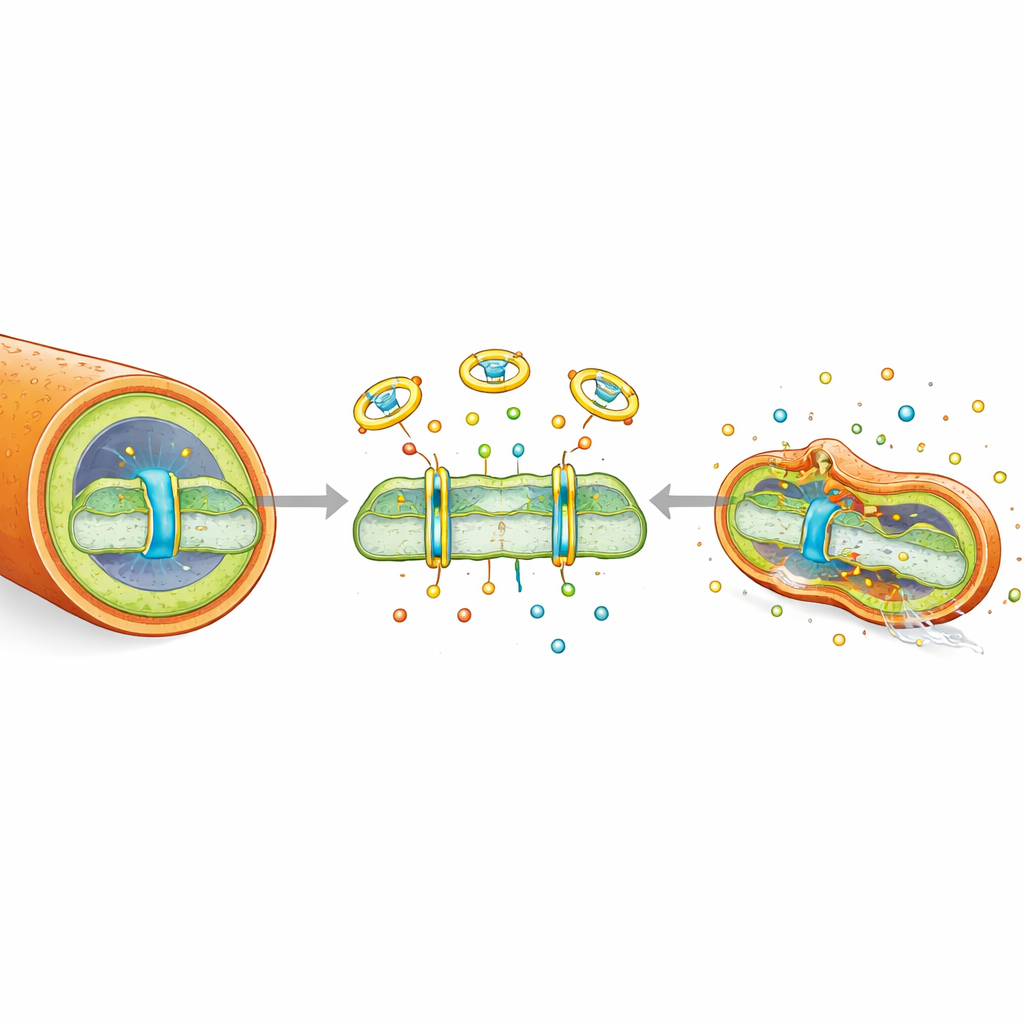
Targeting a Key Cell‑Wall Builder
To pinpoint a specific molecular target, the researchers isolated rare TB relatives that had spontaneously become less sensitive to dinactin and sequenced their genomes. Most of these mutants carried the same change in a gene called cpsA, which encodes a member of the LytR‑Cps2A‑Psr (LCP) protein family involved in attaching major cell‑wall components together. When cpsA or its partner protein was overproduced, bacteria became more tolerant to dinactin; deleting cpsA made cells more resistant on plates, but also highlighted that dinactin likely has additional targets. Using structural modeling and binding experiments, the team showed that dinactin binds tightly to the cpsA protein at a specific site, and that the resistance‑associated mutation greatly weakens this interaction. Because LCP proteins are widespread in Gram‑positive bacteria and are absent from most Gram‑negative species, this targeting helps explain why dinactin preferentially attacks TB and related organisms.
What This Could Mean for Future TB Treatment
For non‑specialists, the central message is that dinactin is a natural compound that strikes TB bacteria at their weakest point: their energy supply and cell‑wall assembly. It behaves like a tiny ion shuttle that drains the bacterial “battery,” starves the cells of ATP, scrambles their internal chemistry, and interferes with a crucial cell‑wall‑building protein. At the same time, it works hand‑in‑hand with frontline TB drugs, making them more effective against resistant and dormant bacteria. While much work remains—especially safety testing and trials in mammalian models—this study positions dinactin and related molecules as promising building blocks for the next generation of TB therapies.
Citation: Wang, G., Dong, W., Bai, Y. et al. Naturally occurring dinactin targets cpsA protein and kills Mycobacterium tuberculosis by disrupting the proton motive force. Commun Biol 9, 417 (2026). https://doi.org/10.1038/s42003-026-09654-3
Keywords: tuberculosis, dinactin, antibiotic resistance, bacterial energy metabolism, cell wall proteins