Clear Sky Science · en
Model selection in preclinical nucleic acid therapeutics research
Turning Genes into Medicines
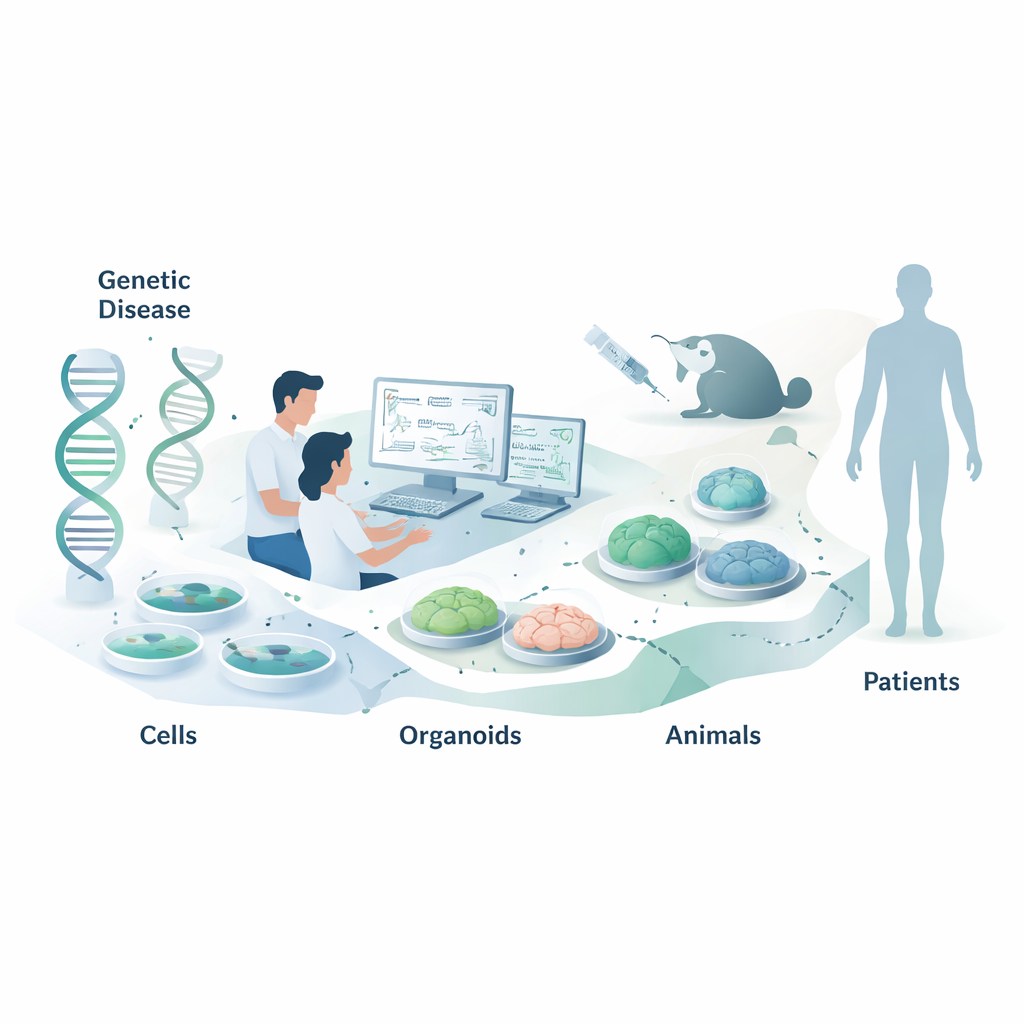
Nucleic acid therapies are a new kind of medicine that work not by blocking proteins, like most drugs do, but by going one step earlier to the level of our genetic messages. This review explains how scientists choose the right laboratory and animal models to test these RNA‑targeting drugs before they reach patients. A lay reader might care about this because smart model choices can mean faster, safer, and more effective treatments for genetic diseases, including rare childhood disorders and more common conditions like heart disease.
What Makes These Genetic Medicines Different?
Nucleic acid therapeutics (NATs) include short strands called antisense oligonucleotides (ASOs) and small interfering RNAs (siRNAs). Rather than relying on the shape and chemistry of protein targets, these drugs recognize their targets by base‑pairing rules—the same A‑T and G‑C matching that underlies DNA. That makes them programmable: once you know the RNA sequence you want to change, you can often design many candidate drugs quickly without years of traditional chemistry. The hard part is no longer “Can we make an active compound?” but “How do we measure whether it truly works in a realistic biological setting?” Because even a single letter difference in sequence can make or break activity, choosing the right test systems becomes critical.
Simple Cell Tests: Fast but Not the Whole Story
The first stop for most NATs is a panel of relatively easy‑to‑grow cell lines, like HeLa or HEK293 cells, that already make the target RNA. Researchers add large libraries of candidate ASOs or siRNAs and then measure how much the target RNA and its protein drop, using techniques such as PCR and Western blotting. They may also work with cells from patients, which better capture a person’s exact genetic background and allow testing of “allele‑selective” drugs that silence only the faulty copy of a gene. When the natural gene is not conveniently expressed, scientists sometimes introduce artificial mini‑genes or reporter plasmids that glow when the RNA is cut or correctly spliced. These systems are powerful for comparing many designs quickly, but they can miss important details, such as natural RNA structure or cell‑type‑specific processing, so results must be confirmed in more realistic contexts.
From Molecular Fixes to Real Cell Behavior
Many NATs aim not just to destroy faulty RNA but to repair it by altering splicing, the process that stitches together pieces of the gene message. Carefully designed ASOs can cause cells to skip a harmful exon, restore a missing piece, or prevent insertion of a “pseudo‑exon” that breaks the code. To see whether such changes actually improve cell function, researchers move beyond simple readouts of RNA and protein levels. In patient‑derived cells, they test whether enzyme activity returns, whether ion transport in lung cells normalizes, or whether signaling pathways in immune cells respond correctly. Increasingly, they use three‑dimensional organoids—mini‑tissues grown from stem cells of patients—that can mimic heartbeats, brain network activity, or other complex behaviors. These 3‑D models are closer to real organs but are harder, slower, and more expensive to run, so teams must balance realism with throughput.
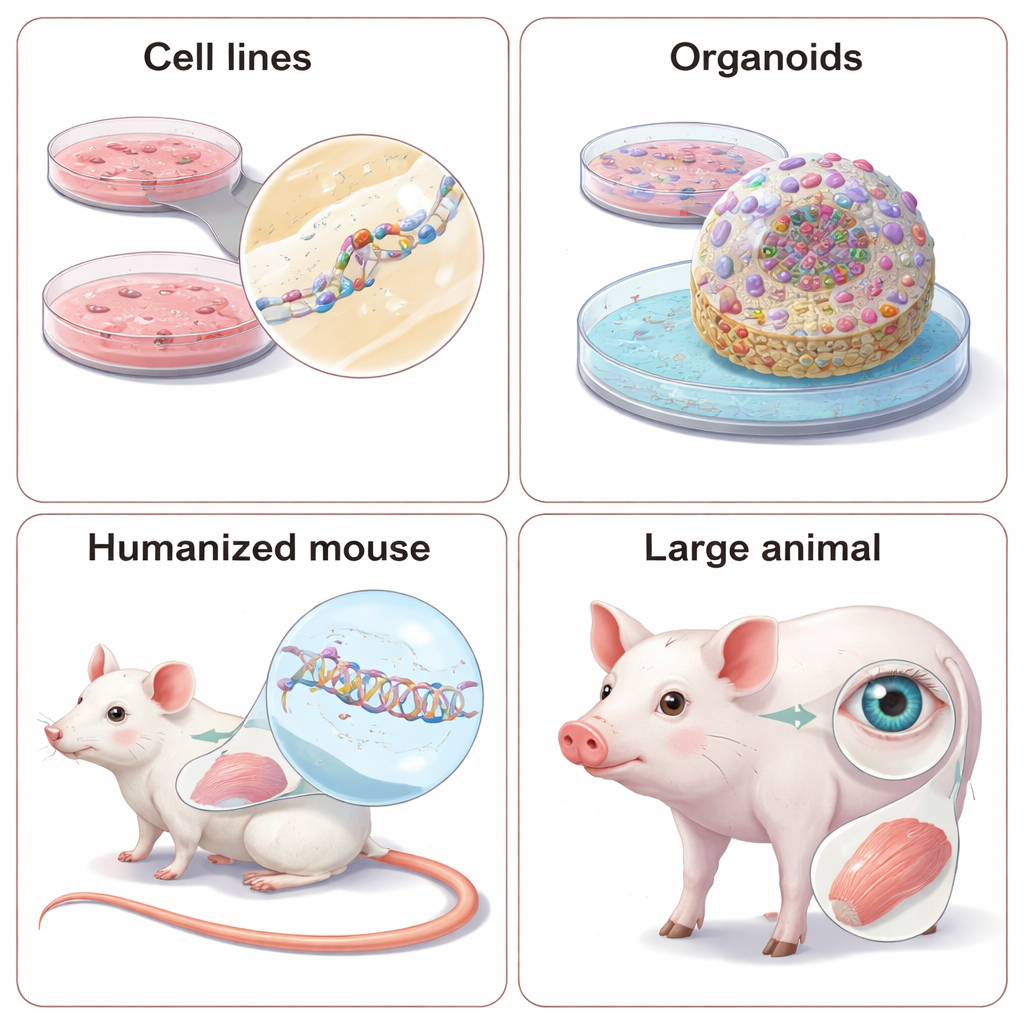
Why Animals Still Matter—and How to Humanize Them
Some disease features, such as whole‑body metabolism, organ‑to‑organ interactions, or behavior, simply cannot be captured in dishes. For this, NATs are tested in living animals. Because these drugs care about exact RNA sequences, scientists face a choice: design “surrogate” drugs that match the animal’s version of the gene, or re‑engineer the animal to carry human genetic sequences. Surrogate ASOs and siRNAs can reveal whether lowering a gene improves symptoms in well‑established mouse models, but they do not perfectly mimic the human‑targeting drug. Humanized models—mice or even mini‑pigs with sections or entire copies of human genes—allow testing of the real clinical candidate but require significant time and cost, and subtle differences in gene regulation between species can still lead to surprises. The review also highlights growing use of large animals, especially for diseases of the eye and muscle, where anatomy more closely resembles that of humans.
Looking Ahead: Smarter Models and Less Guesswork
The authors conclude that there is no single “best” model for nucleic acid drugs; instead, each project needs a tailored path that starts with fast, simple assays and advances toward more complex, human‑relevant systems. As regulators and funders push to reduce animal use, sophisticated organoids, organ‑on‑a‑chip devices, and computer models are expected to carry more of the load, especially when combined with machine learning tools that predict which sequences and chemistries will work best. Ultimately, by understanding the strengths and blind spots of each model—and by sharing best practices and data across the field—researchers can more reliably turn programmable genetic ideas into safe, effective medicines for patients.
Citation: Oliver, P.L., Hill, A.C. Model selection in preclinical nucleic acid therapeutics research. Commun Biol 9, 200 (2026). https://doi.org/10.1038/s42003-026-09650-7
Keywords: nucleic acid therapeutics, antisense oligonucleotides, siRNA, preclinical models, gene therapy