Clear Sky Science · en
Selective binding of divalent cations reshapes nucleosome mechanics and unlocks histone tail dynamics
How tiny ions help manage our DNA
Inside every cell, meters of DNA must be folded, packed, and yet still accessible on demand. This paper explores how two common mineral ions in our cells, magnesium and calcium, subtly reshape the way DNA is wrapped around proteins, influencing how tightly our genetic material is packaged and how easily it can be read. By simulating these interactions in atomic detail, the authors reveal that shifts in ion levels can stiffen or relax local DNA structures and change the behavior of floppy protein tails that help organize chromosomes.
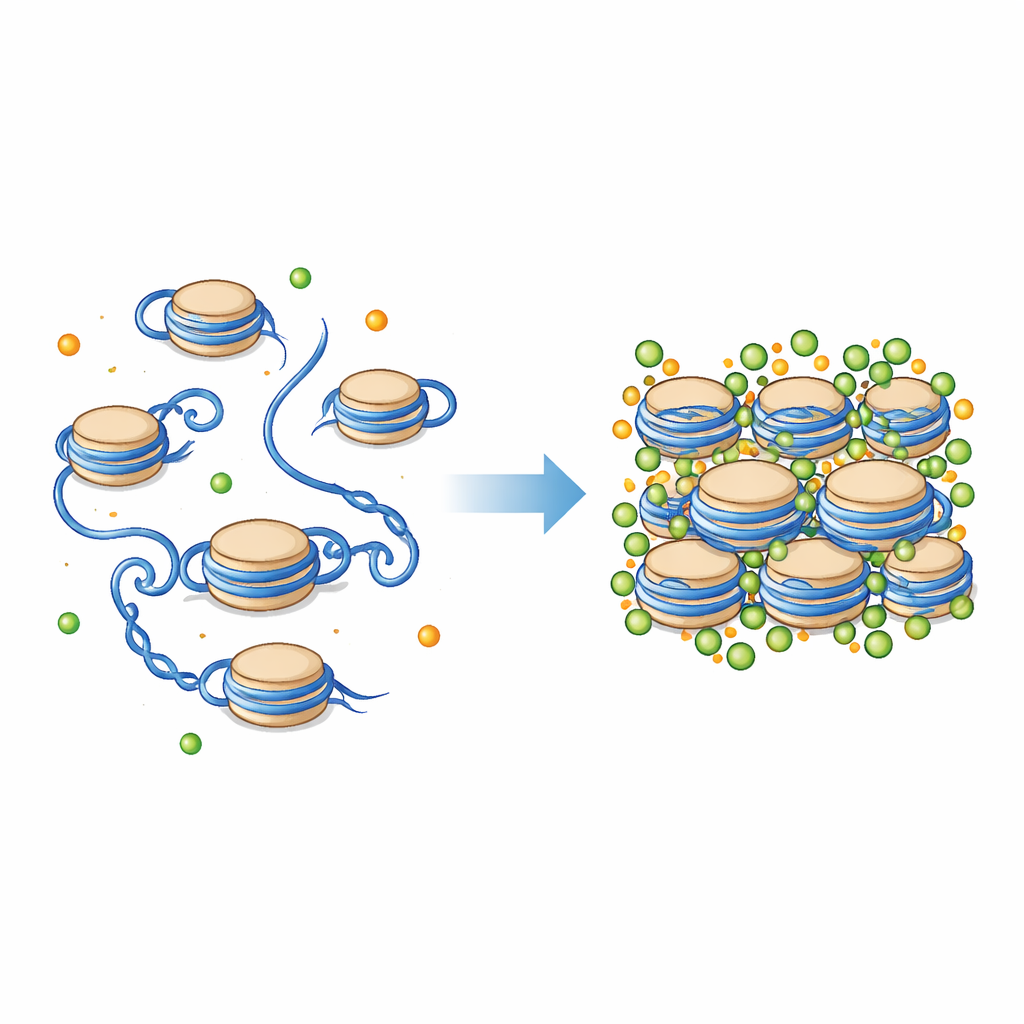
The bead at the heart of chromosome packing
DNA in our cells is not left loose; it is wound around protein spools called nucleosomes. Each nucleosome is a short stretch of DNA wrapped around a cluster of histone proteins, with flexible tails that extend outward. Together, these units form the basic "beads on a string" structure of chromatin, which can then fold into more compact fibers. The environment around nucleosomes is full of charged particles, including magnesium (Mg²⁺) and calcium (Ca²⁺). These ions are known to help chromosomes condense, but exactly how they influence the tiny mechanical details of nucleosomes and their histone tails has been difficult to see directly.
Simulating a crowded atomic world
To uncover these details, the authors ran 81 microseconds of large-scale computer simulations in which every atom of DNA, protein, water, and ions was explicitly modeled. They systematically varied magnesium and calcium levels, tested different ways of describing ion behavior in the simulations, and studied nucleosomes with and without their histone tails. By comparing the simulated ion binding patterns with experimental measurements, they identified a refined model that best matched how real nucleosomes attract these ions, where magnesium prefers DNA grooves and calcium more often grips the DNA backbone and certain acidic protein sites.
How ions squeeze DNA and stiffen the core
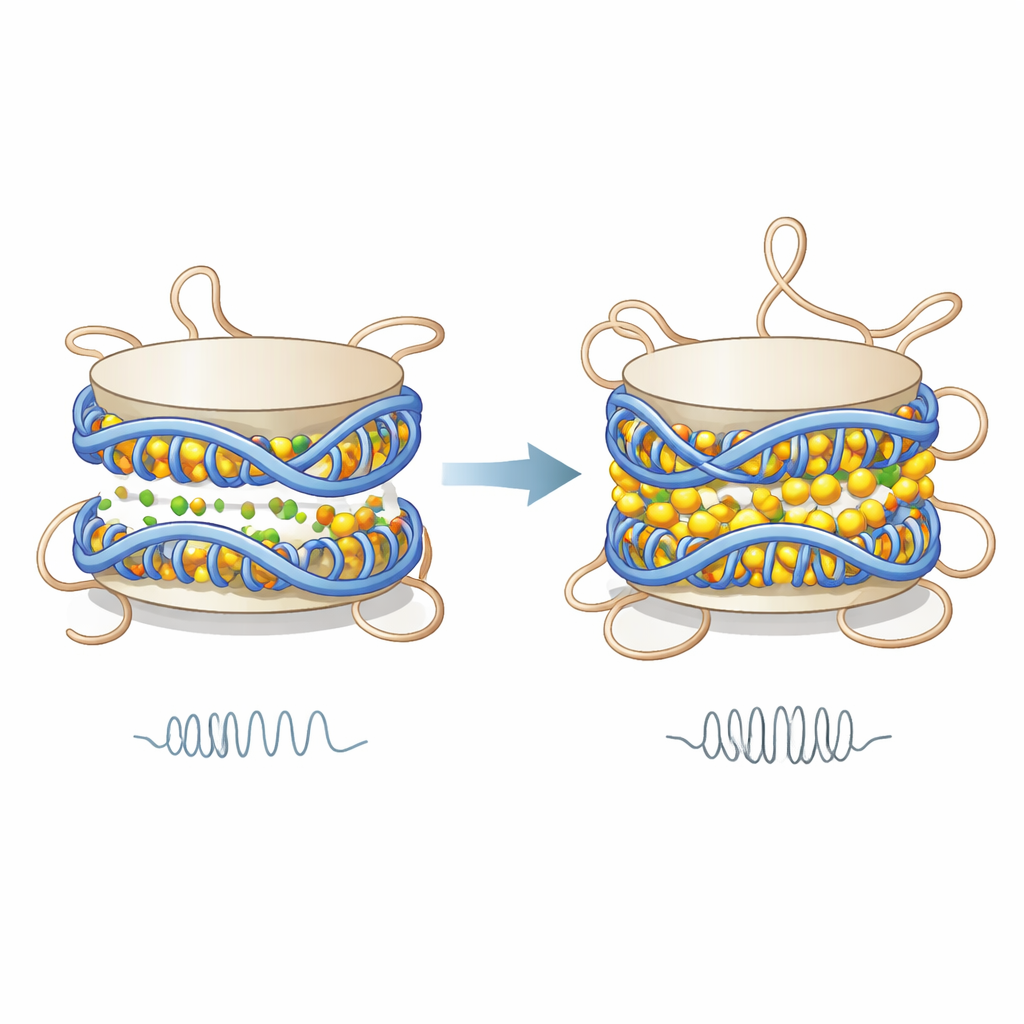
The simulations show that divalent ions cluster along the DNA surface and in the narrow space between the two turns of DNA that wind around each nucleosome. By partially neutralizing DNA’s strong negative charge, these ions reduce the electrical repulsion between neighboring DNA turns. The result is that the two DNA coils are drawn closer together, the nucleosome becomes slightly shorter, and its mechanical stiffness increases, as measured by a model that treats the nucleosome like a tiny elastic cylinder. This added stiffness does not come from DNA falling off or major unwrapping; instead, it arises from subtle changes in spacing and more coordinated motions of specific DNA regions, which move together like a more rigid unit.
Unlocking the floppy protein tails
While the DNA core becomes more compact and rigid, the flexible histone tails respond in the opposite way. Under low-ion conditions, positively charged amino acids in these tails cling to the negatively charged DNA, forming many stable contacts. When magnesium or calcium bind along the DNA, they screen those charges, weakening the attraction between tails and DNA. The authors find that contacts between tails—especially those from the H3 histone—and DNA drop, residence times on DNA become shorter, and tails sample a wider range of positions. This means the tails interact with DNA in more frequent but briefer encounters, becoming more dynamic and potentially freer to reach neighboring nucleosomes.
What this means for gene control and chromatin state
Together, these findings outline a dual role for magnesium and calcium in chromatin. By tightening the wrapping of DNA and increasing nucleosome stiffness, they promote more compact, less flexible chromatin. At the same time, by loosening histone tail–DNA contacts and boosting tail motion, they expose key sites that can be chemically modified or recognized by regulatory proteins. Because ion concentrations in the nucleus fluctuate with signals and energy usage, this work suggests that small chemical changes in the cellular environment can tune both the mechanical feel of nucleosomes and the accessibility of their regulatory tails, helping to open or close windows of opportunity for gene activity.
Citation: Hu, G., Zhang, H., Xu, W. et al. Selective binding of divalent cations reshapes nucleosome mechanics and unlocks histone tail dynamics. Commun Biol 9, 365 (2026). https://doi.org/10.1038/s42003-026-09648-1
Keywords: chromatin, nucleosome mechanics, magnesium and calcium ions, histone tails, gene regulation