Clear Sky Science · en
Tbr2-dependent parallel pathways regulate the development of distinct ipRGC subtypes
How Our Eyes Tell Time and Shape
Beyond forming images, our eyes also quietly keep our body clock on schedule, control how wide our pupils open, and help us sense overall brightness. A small population of special retinal nerve cells, called intrinsically photosensitive retinal ganglion cells (ipRGCs), handle much of this work. They respond to light using a pigment called melanopsin and send signals deep into the brain. This study asks a basic question with big implications: how does one early group of developing nerve cells split into several distinct ipRGC types, each wired for a different job?
One Master Switch, Many Light-Sensing Cell Types
In the mouse retina, all six known ipRGC subtypes arise from an early population of ganglion cells that switch on a gene called Tbr2. This gene acts like a master switch, turning on programs needed to make ipRGCs and to keep the melanopsin gene, Opn4, active. But a single master switch cannot by itself explain how six different ipRGC varieties emerge, each with unique shapes, light responses, and brain targets. The authors used RNA sequencing of developing mouse retinas to search for genes whose activity depends on Tbr2. Among a small set of candidates, two stood out: Irx1 and Tbx20, both known regulators of cell identity in other parts of the body. 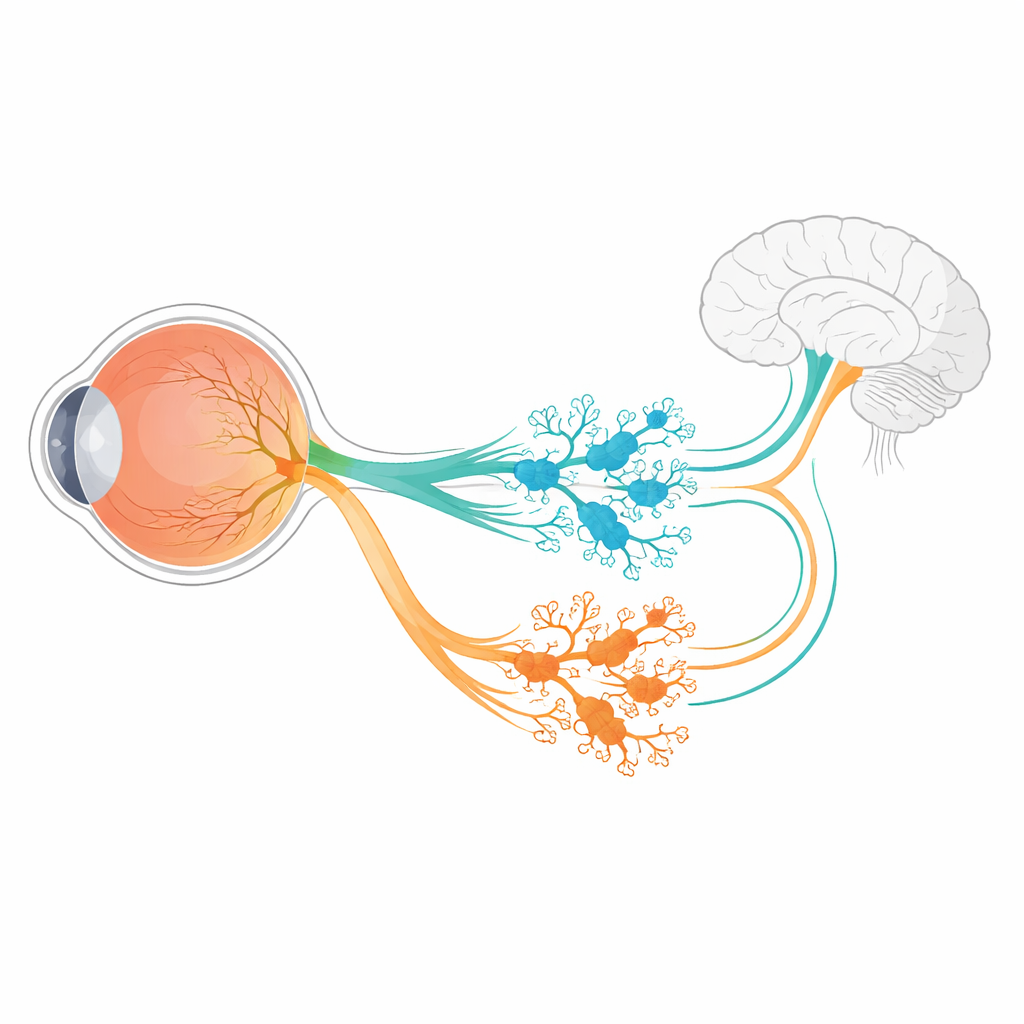
Two Branching Paths Inside the Retina
Using newly engineered mouse lines that tag Irx1- and Tbx20-producing cells with fluorescent or enzyme markers, the researchers traced where and when these factors are turned on. They found that Irx1 and Tbx20 appear in overlapping groups of young retinal ganglion cells around mid-gestation, but quickly sort into almost completely separate sets by adulthood. Irx1 is mostly found in three ipRGC types labeled M3, M4, and M5, whereas Tbx20 is concentrated in M1, M2, and M6 cells, with only minor overlap in some M3 and M5 cells. Detailed imaging, electrical recordings, and brain tracing showed that each factor marks a distinct collection of ipRGCs with characteristic dendritic layouts, light responses, and projection patterns to specific brain regions involved in clock setting, reflexes, and visual processing.
Switches That Tune Light Sensitivity and Cell Survival
The team then asked what happens when each factor is removed. When Irx1 was knocked out in developing retina, the number and basic structure of Irx1-marked ipRGCs remained largely intact, but their melanopsin levels dropped sharply. In other words, Irx1 proved essential for fully turning on the Opn4 light-sensing program in M3, M4, and M5 cells, but not for forming those cells in the first place. Tbx20 behaved differently. Deleting Tbx20 not only lowered Opn4 expression but also cut the number of Tbx20-positive ipRGCs roughly in half and later reduced the survival of these cells. This indicates that Tbx20 helps both build and maintain particular ipRGC subtypes, especially M1, M2, and M6, while also supporting their melanopsin-based light sensitivity. 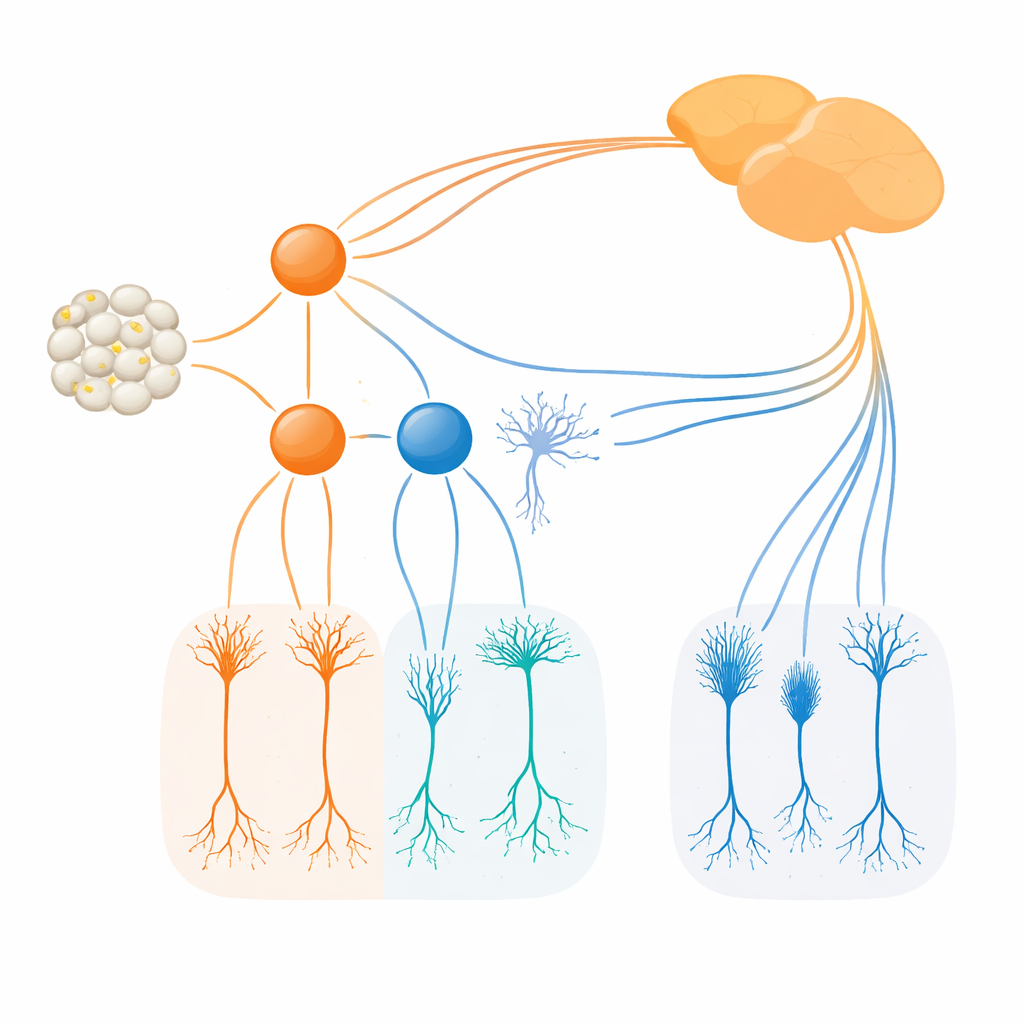
Timing the Split Into Distinct Cell Families
By briefly activating genetic reporters at precise embryonic days, the authors could “time-stamp” cells that expressed Irx1 or Tbx20 and follow their fates into adulthood. These experiments showed that Tbx20-marked cells commit to their final ipRGC identities slightly earlier than Irx1-marked cells, with most decisions locked in just before birth. During a short window, both Irx1 and Tbx20 can be co-expressed in some cells, and double knockout mice lacking both genes showed a much stronger loss of melanopsin-positive ipRGCs than either single mutant. This suggests that the two pathways work together transiently, then diverge to guide different subsets of ipRGCs along separate developmental tracks.
What This Means for Understanding Vision and Health
To a non-specialist, the main message is that a single early gene, Tbr2, does not act alone. Instead, it feeds into two parallel control paths, one centered on Irx1 and the other on Tbx20. These paths decide which kind of ipRGC a young cell will become, fine-tune how strongly it senses light, and determine whether it connects to brain regions for body-clock regulation, pupil reflexes, or other visual roles. The work offers a clearer wiring diagram for how our internal “light meters” are built, and highlights specific genetic steps that could be involved when these cells are lost or malfunction, with potential relevance for sleep disorders, seasonal mood changes, and diseases that damage the retina.
Citation: Kiyama, T., Chen, CK., Altay, H.Y. et al. Tbr2-dependent parallel pathways regulate the development of distinct ipRGC subtypes. Commun Biol 9, 347 (2026). https://doi.org/10.1038/s42003-026-09645-4
Keywords: retinal ganglion cells, melanopsin, neural development, circadian rhythms, transcription factors