Clear Sky Science · en
LncRNA P4HA2-AS1 drives renal interstitial fibrosis via trim32-mediated k63 ubiquitination of ULK1 and autophagic dysregulation
Why Kidney Scarring Matters
Chronic kidney disease affects hundreds of millions of people worldwide and often creeps forward silently until dialysis or a transplant is the only way to stay alive. At the heart of this decline is a process called scarring, where normal kidney tissue is gradually replaced by stiff, non‑functional material. This study asks a simple but crucial question: what molecular switches inside kidney cells push them toward this harmful scarring, and can those switches be turned back?
A Hidden RNA Switch in Kidney Cells
Our kidneys filter blood using elaborate networks of tiny tubules. When these tubules are injured by blocked urine flow, high blood sugar, or a sudden loss of blood supply, they can respond by laying down scar tissue between the cells. The researchers searched the kidneys of mice with several types of injury and compared them to healthy kidneys using RNA sequencing, a technique that reads out which genes are turned on. They discovered that a little‑known molecule called P4HA2-AS1, a long non‑coding RNA that does not make protein, was consistently elevated in damaged, scar‑forming kidneys and in human kidney cells stressed in the laboratory. This RNA was mainly found in the fluid‑filled part of the cells, hinting that it might control other proteins there. 
Dialing Down the RNA to Protect the Kidney
To test whether P4HA2-AS1 is simply a bystander or an active driver of damage, the team reduced its levels in mice and in cultured human kidney cells. In a well‑established mouse model where one ureter is tied off to cause long‑term obstruction, lowering this RNA through a targeted viral delivery eased many hallmarks of scarring: tubules stayed more intact, there was less collagen and other scar proteins between cells, and kidney function markers in the blood improved. In dishes of human kidney cells, shrinking P4HA2-AS1 levels also blunted the response to a potent scarring signal, TGF‑β, leading to less production of fibronectin, collagen, and other pro‑fibrotic factors. These experiments showed that this RNA is not merely a marker of damage but actively fuels the scarring process.
Autophagy: From Housekeeping to Harm
The researchers then asked how this RNA does its damage. Their gene‑expression and imaging data pointed to autophagy, the cell’s built‑in recycling system that breaks down worn‑out proteins and organelles. Under normal conditions, autophagy is a healthy housekeeping process, but when driven too hard for too long it can push injured kidneys toward maladaptive repair and scarring. In stressed kidney cells, P4HA2-AS1 boosted the flow of material through autophagy, leading to an accumulation of recycling vesicles. When the RNA was blocked, this overactive recycling dropped back toward normal, both in cells and in kidneys from obstructed mice. Mice lacking ULK1, a central starter enzyme for autophagy, were also protected from scarring, and in these animals P4HA2-AS1 overexpression could no longer worsen the damage, firmly linking the RNA’s effects to this pathway.
A Three‑Part Molecular Chain
Digging deeper, the team used biochemical fishing techniques to identify proteins that physically bind to P4HA2-AS1. One stood out: TRIM32, an enzyme that tags other proteins with small molecular “flags” called ubiquitin. The RNA binds to TRIM32 and shields it from being broken down, raising its levels inside kidney cells. Stabilized TRIM32, in turn, attaches a specific type of ubiquitin chain to ULK1. Rather than sending ULK1 for destruction, this chain acts like a power‑up, enhancing ULK1’s ability to trigger autophagy. When the scientists either removed TRIM32 in mice or silenced it in kidney cells, scarring and excessive autophagy were both reduced. Re‑introducing TRIM32 into cells where P4HA2-AS1 had been blocked partially restored the scarring response, showing that this enzyme is a key middle link in the chain. 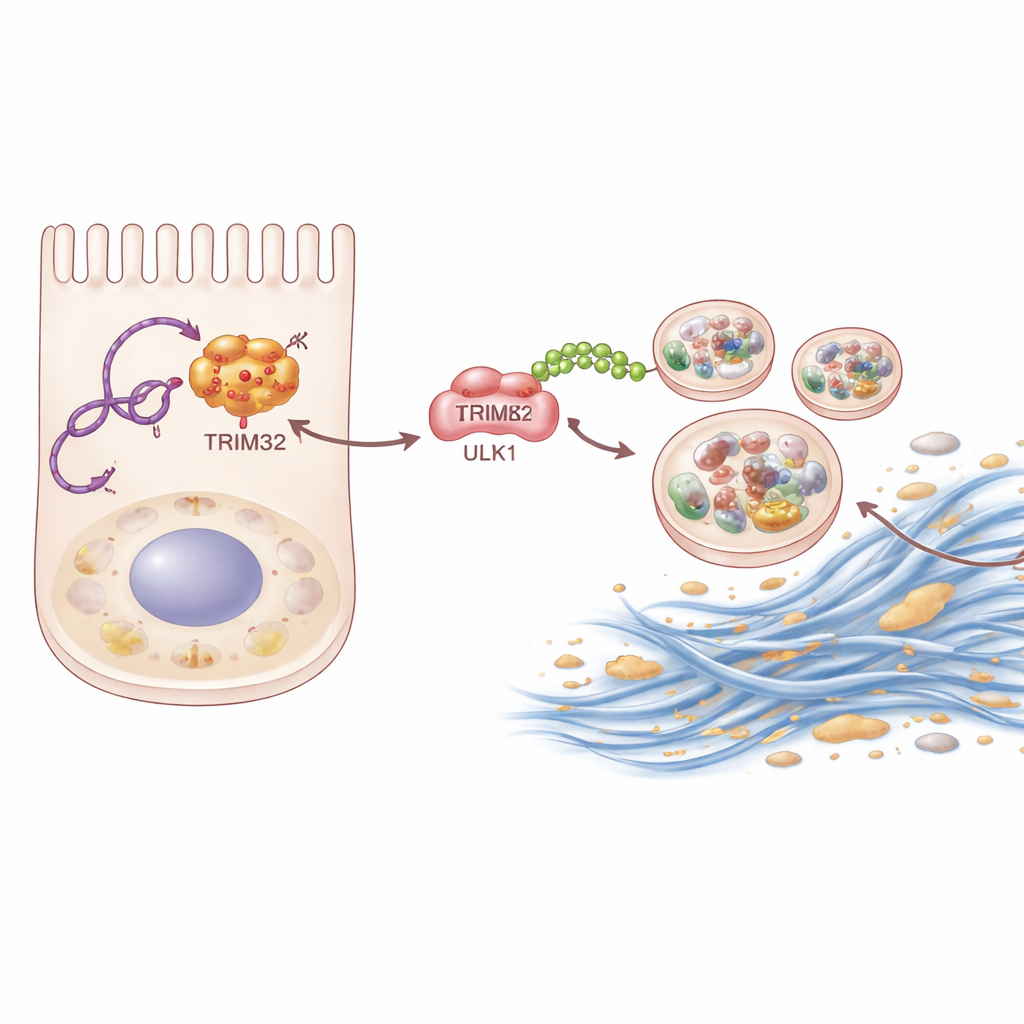
New Paths Toward Softer Kidneys
Taken together, the work reveals a previously unknown molecular route that connects long non‑coding RNA, protein tagging, and cellular recycling to the buildup of kidney scar tissue. In injured kidneys, P4HA2-AS1 rises, stabilizes TRIM32, supercharges ULK1, and drives autophagy beyond its helpful range, ultimately encouraging fibrotic protein buildup and loss of function. For non‑specialists, the take‑home message is that kidney scarring is not an inevitable black box: it is driven by identifiable switches that might be druggable. Targeting this P4HA2-AS1–TRIM32–ULK1 axis—by lowering the rogue RNA, taming TRIM32, or fine‑tuning ULK1 activity—could one day offer new treatments to slow or prevent the progression of chronic kidney disease.
Citation: Pan, Z., Xiao, F., Hu, W. et al. LncRNA P4HA2-AS1 drives renal interstitial fibrosis via trim32-mediated k63 ubiquitination of ULK1 and autophagic dysregulation. Commun Biol 9, 339 (2026). https://doi.org/10.1038/s42003-026-09618-7
Keywords: chronic kidney disease, renal fibrosis, autophagy, long noncoding RNA, kidney tubules