Clear Sky Science · en
Integrated analysis of GWAS and molQTLs reveals cell-specific genetic variants in the porcine immune system
Why pig immunity matters to all of us
Pigs help feed much of the world and also serve as important stand-ins for humans in medical research. Yet infectious diseases in pigs cost farmers billions of dollars each year and drive heavy use of antibiotics. This study asks a simple but powerful question: how do tiny DNA differences shape the way pig immune cells work, and can we trace those effects down to the behavior of individual cell types in the blood? Understanding that chain of cause and effect could guide breeding for healthier herds and offer clues about our own immune systems.
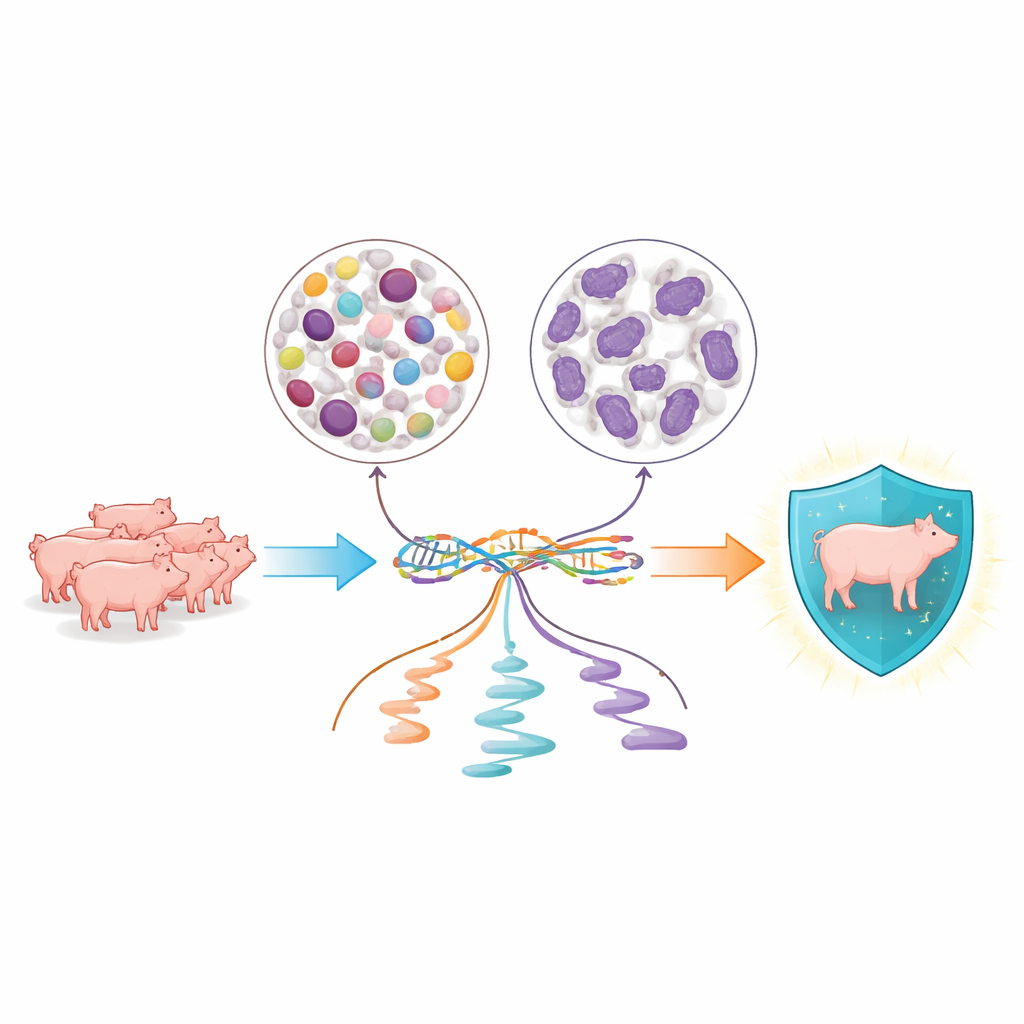
Looking closely at two key blood cell families
The researchers focused on two major groups of white blood cells that constantly patrol pig blood: peripheral blood mononuclear cells, which include lymphocytes and monocytes involved in slower, targeted defenses, and neutrophils, which mount rapid, first-line attacks on microbes. From 134 young Yorkshire pigs, they purified each cell type separately and captured both whole-genome DNA data and detailed snapshots of RNA, the molecules that carry genetic instructions inside cells. This allowed them to see not only which genes were turned on, but also how the cell cut and finished those messages before using them.
Connecting DNA variants to molecular switches
Instead of looking only at which DNA variants associate with disease risk, the team mapped how variants influence three kinds of molecular features inside each cell type. Some variants altered how much RNA a gene produced. Others changed how the RNA was spliced, trimming or rearranging pieces. A third class shifted how the RNA was finished at its tail end, a process called polyadenylation that can affect message stability. These variant–effect pairs are called molecular quantitative trait loci, or molQTLs. The maps revealed thousands of such sites in each cell type, and strikingly, more than four out of five were specific to either the mixed mononuclear cells or the neutrophils, underscoring how differently the same genome can be read in different immune cells.
Finding networks that link genes to real-world immune traits
To see how these molecular switches relate to whole-animal immunity, the scientists combined the molQTL maps with standard blood tests and measurements of immune signaling proteins. They built co-expression networks—groups of genes that tend to rise and fall together—and asked which ones track with traits such as the proportion of neutrophils or lymphocytes, white blood cell count, or levels of interferons and tumor necrosis factor. Modules tied to lymphocyte traits were enriched for adaptive immune pathways, while those linked to neutrophil traits highlighted rapid innate defenses and inflammatory signaling. Many key genes in these modules were directly regulated by cell type-specific molQTLs, suggesting clear routes from DNA variant to altered cell composition or function.
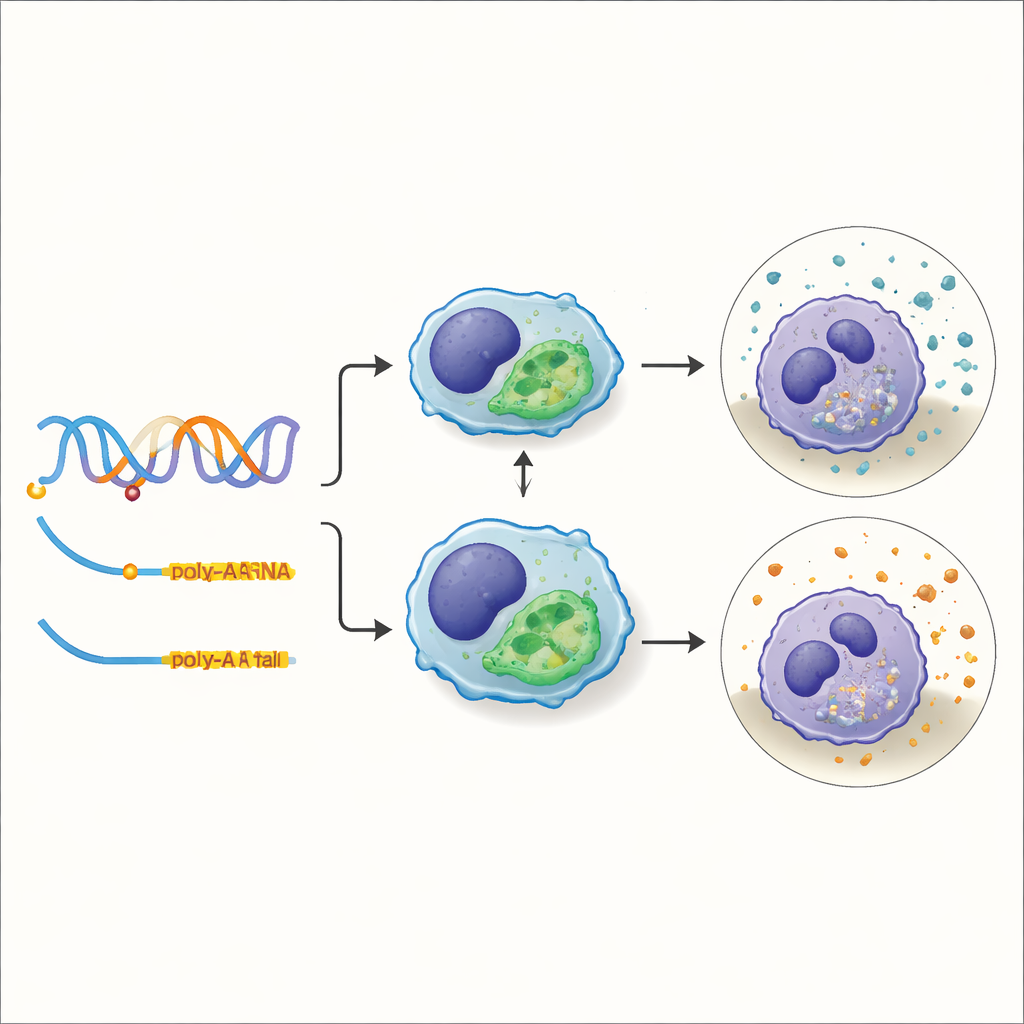
Zooming in on a variant that tunes cellular “cleanup” ability
One of the most practical traits the team examined was phagocytic capacity—the ability of immune cells to engulf and clear debris or pathogens. By overlaying their molQTLs with the results of previous genome-wide association studies for phagocytosis, they pinpointed 588 regions where the same DNA change likely drives both a molecular shift and a change in cellular cleanup performance. A standout example was a variant within a gene called TXNDC15. This change did not simply turn the gene on or off; instead, it altered which polyadenylation site the cell chose for the gene’s RNA. Pigs carrying one version of the variant favored a shorter tail on the RNA, which proved more stable and accumulated at higher levels, and these animals showed stronger phagocytic activity in their immune cells.
What this means for healthier pigs and beyond
In everyday terms, this work shows how small DNA differences can rewire the “instructions” inside specific immune cell types and, through that, influence how many such cells exist and how well they perform their jobs. By carefully separating cell types and tracking not only gene activity but also how messages are cut and finished, the study uncovers hidden control points that bulk tissue studies miss. The resulting genetic map provides a foundation for breeding pigs that are naturally more disease-resistant, potentially reducing the need for antibiotics. At the same time, because pig immunity closely resembles our own, these insights may also help scientists understand how similar mechanisms operate in human health and disease.
Citation: Yang, J., Chen, S., Tang, Y. et al. Integrated analysis of GWAS and molQTLs reveals cell-specific genetic variants in the porcine immune system. Commun Biol 9, 408 (2026). https://doi.org/10.1038/s42003-026-09605-y
Keywords: pig immunity, genetic variants, immune cells, phagocytosis, molecular QTLs