Clear Sky Science · en
Molecular QTL are enriched for structural variants in a cattle long-read cohort
Why cattle DNA can teach us about complex traits
Farmers, veterinarians, and geneticists all want to understand why some animals grow faster, resist disease, or produce more milk than others. Much of the answer lies in DNA, but our usual tools mostly look at tiny changes in single “letters” of the genome. This study shows that much larger DNA changes—structural variants—quietly shape how genes work in cattle, and that new long-read sequencing technologies are finally letting us see their full impact.
Looking at the genome with a sharper lens
Most genetic studies rely on short snippets of DNA sequence, which are cheap and accurate but struggle in repetitive or complex regions of the genome. The authors used a newer technique, long-read sequencing, on 120 bulls from a dairy-related cattle breed. These long reads span much larger stretches of DNA, making it easier to spot big insertions, deletions, and rearrangements known as structural variants. The team compared these long reads with existing short-read data from the same animals, and found that long reads uncovered more variants overall and dramatically improved coverage of difficult regions such as the X and Y chromosomes.
Uncovering thousands of hidden DNA rearrangements
With the long-read data, the researchers cataloged about 24 million small DNA changes and over 79,000 structural variants across the bulls. Many of these larger changes were linked to repetitive DNA elements that copy and paste themselves around the genome. About one in ten structural variants appeared in only one or two animals, revealing a rich reservoir of rare variation. Compared with an earlier cattle “pangenome” built from high-quality assemblies, the new dataset added tens of thousands of extra structural variants, especially insertions and complex duplications that are hard to detect with older methods. This suggests that long-read studies are still uncovering previously invisible layers of genetic diversity in livestock.
Connecting DNA changes to gene activity
To see how these DNA differences actually affect biology, the team turned to a tissue that matters for male fertility: the testis. For 117 of the bulls, they had deep RNA sequencing data that reveal which genes are turned on and how their messages are spliced. By statistically linking genetic variants near each gene to its activity, they identified over 27,000 “molecular QTLs”—genomic sites that change either how much of a gene is expressed or how its RNA is stitched together. Structural variants emerged as key players: they were more than twice as common among top expression signals and over five times as common among top splicing signals as expected by chance. In many cases, the most influential variant was a large insertion, deletion, or duplication sitting in a promoter, enhancer, exon, or splice site rather than a single-letter change.
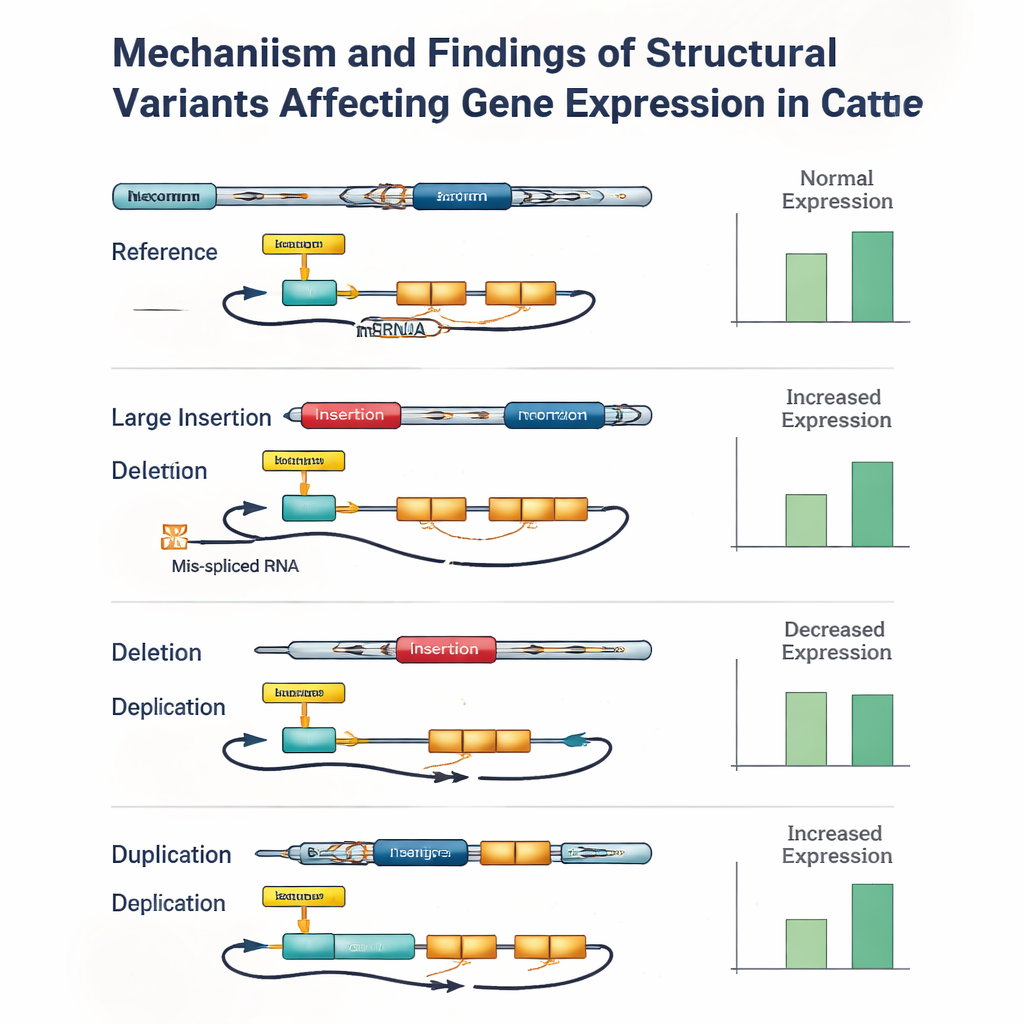
When genotyping errors hide important signals
However, the study also exposed the limits of current tools. Even with high-quality long reads, accurately assigning structural variant genotypes to each animal was challenging, especially for big insertions and long duplications. Small mistakes—sometimes involving only one or two bulls—could make a structural variant appear slightly less statistically significant than a nearby small variant that was in perfect genetic lockstep with it. When the authors manually checked some of the strongest signals, they repeatedly found cases where a structural variant within a gene or key regulatory region was the most plausible driver of the effect, but genotyping errors or missing data caused a linked small variant to top the ranking.
What this means for cattle breeding and beyond
For non-specialists, the takeaway is that “big” DNA changes matter a lot. This long-read survey in cattle shows that structural variants are strongly enriched among genetic sites that control how genes are switched on and spliced, particularly in reproductive tissue. Yet the study also warns that today’s analysis methods still miss or mislabel many of these variants, especially when sequencing depth is modest. As long-read sequencing becomes cheaper and more accurate, and as better software is developed, breeders and researchers will be able to trace economically important traits—like fertility, disease resistance, and milk production—back to specific structural variants. The same principles apply to human health and plant breeding: to fully understand complex traits, we must look beyond single-letter changes and embrace the larger rearrangements that reshape genomes.
Citation: Mapel, X.M., Leonard, A.S. & Pausch, H. Molecular QTL are enriched for structural variants in a cattle long-read cohort. Commun Biol 9, 290 (2026). https://doi.org/10.1038/s42003-026-09596-w
Keywords: structural variants, long-read sequencing, cattle genomics, gene expression, molecular QTL