Clear Sky Science · en
Characterising nanobody developability to improve therapeutic design using the Therapeutic Nanobody Profiler
Why tiny antibody cousins matter for future medicines
Many of today’s blockbuster drugs are antibodies—proteins that latch onto disease targets with great precision. A newer class of even smaller binders, called nanobodies, can slip into hard‑to‑reach pockets on viruses, tumors, and other molecules. But being able to bind a target is not enough: a candidate drug also has to be easy to make, stable in a vial, and safe in the body. This paper introduces the Therapeutic Nanobody Profiler, a computational tool designed to help scientists judge, early on and from sequence alone, which nanobodies are most likely to become practical medicines.
From promising idea to practical drug
Turning a protein into a real therapy involves a host of practical hurdles grouped under the term “developability.” Proteins must be produced in large quantities, stay soluble, avoid clumping together, and remain stable during shipping and storage. Over the past decade, researchers have learned how to predict many of these traits for full‑sized monoclonal antibodies, supported by abundant clinical data and specialized lab tests. Nanobodies, however, are structurally different: they consist of a single domain rather than a pair of chains, often have longer binding loops, and expose surface regions that are buried in conventional antibodies. As a result, methods tuned to regular antibodies can give misleading answers when applied to nanobodies.
A profiler built for nanobody quirks
To tackle this mismatch, the authors designed the Therapeutic Nanobody Profiler (TNP), inspired by an earlier tool for standard antibodies but re‑engineered around nanobody biology. They compiled nanobody sequences from many sources: clinical trials, natural immune repertoires, patents, scientific papers, and known crystal structures. Using deep‑learning structure predictors tailored to single‑domain antibodies, they generated 3D models for these sequences. From each model, they measured how long the binding loops are, how far one key loop reaches from the protein body, and how clusters of hydrophobic and charged residues are arranged on the surface—features that strongly influence solubility, aggregation, and non‑specific sticking.
Two structural flavors, both workable
One of the most striking findings concerns the main binding loop, known as CDR3. When the team quantified how “compact” this loop is—comparing its length to how far it extends from the protein—they found a clear split into two structural styles. In one subtype, the loop is longer and folds back over the protein’s side, forming many stabilizing contacts with a set of hallmark residues. In the other, the loop sticks out more like in a conventional antibody fragment. Clinical‑stage nanobodies occupy both subtypes, and when the researchers compared dozens of practical lab measurements—such as aggregation, self‑association, and thermal stability—they saw no systematic penalty for either style. This means drug designers do not need to favor one overall loop shape over the other, as long as other properties look acceptable.
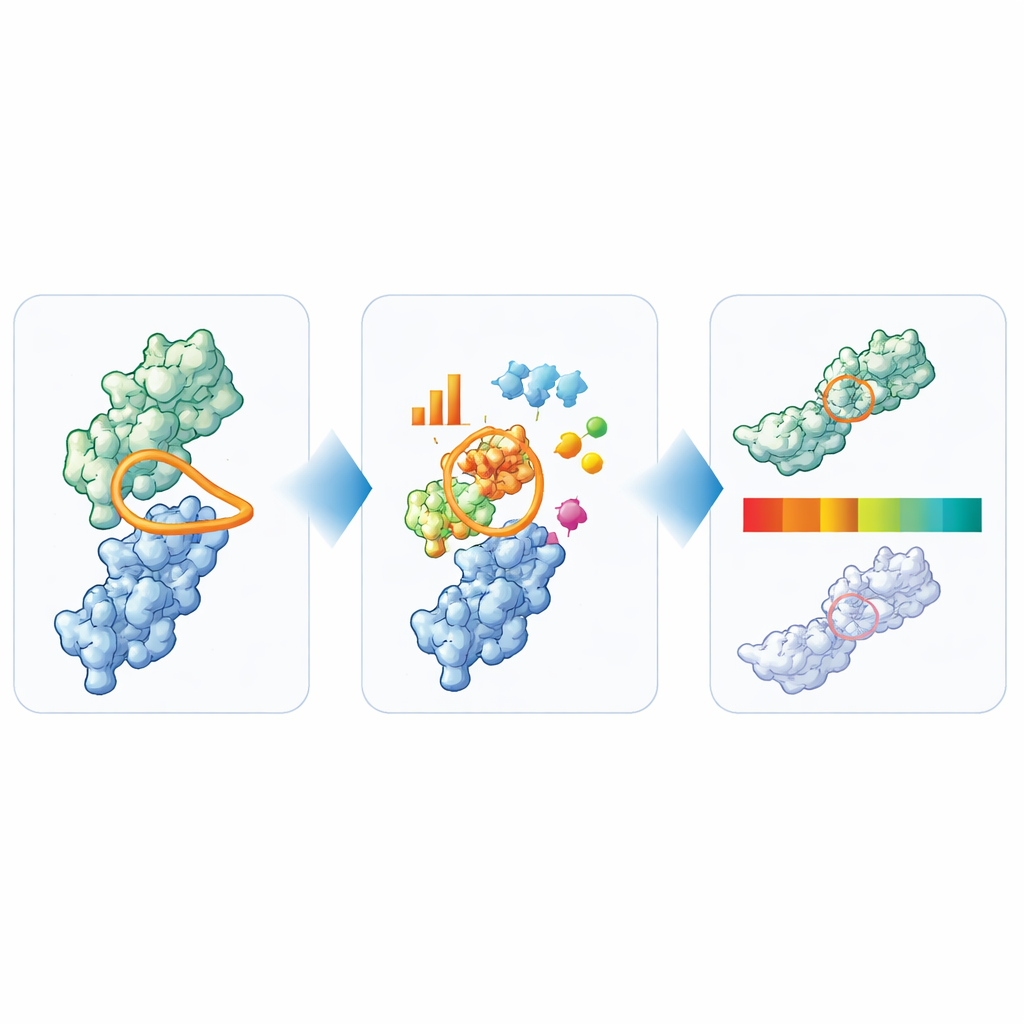
Turning structure into simple traffic lights
From their broad analysis, the authors distilled six key features that together capture the main developability concerns for nanobodies: total loop length, the specific length and compactness of CDR3, and the sizes of hydrophobic, positively charged, and negatively charged surface patches around the binding site. They then used the 36 nanobodies that have reached clinical testing to set practical boundaries for each measure. Values in the central, well‑traveled region are labeled “green,” borderline values “amber,” and clear outliers “red.” To test how informative these signals are, they applied TNP to 72 additional proprietary nanobodies and compared the flags to an extensive panel of lab assays. Nanobodies that TNP marked as clear outliers usually also showed multiple experimental warning signs, while those that looked clean computationally were more often well behaved in the lab.
What this means for tomorrow’s treatments
For non‑specialists, the core message is that the shape and surface pattern of a nanobody can now be converted into a straightforward developability profile before expensive lab work begins. The Therapeutic Nanobody Profiler does not replace experiments, and mismatches between predictions and assays still occur, especially because lab tests were done on nanobodies fused to larger antibody fragments. But by rapidly flagging candidates with unusually long or tightly folded loops or with problematic surface patches, TNP helps steer attention toward nanobodies that are more likely to become reliable drugs. As more nanobodies enter clinical trials and expand the reference set, this tool should grow even more accurate, accelerating the design of small, robust antibody‑like medicines for a wide range of diseases.
Citation: Gordon, G.L., Gervasio, J., Souders, C. et al. Characterising nanobody developability to improve therapeutic design using the Therapeutic Nanobody Profiler. Commun Biol 9, 344 (2026). https://doi.org/10.1038/s42003-026-09594-y
Keywords: nanobodies, biologic drug developability, computational profiling, antibody engineering, protein stability