Clear Sky Science · en
Structural insights into WRN helicase reveal conformational states and opportunities for MSI-H cancer drug discovery
Why this matters for cancer treatment
Certain cancers carry a built-in weakness: they struggle to repair mistakes in their DNA. A protein called WRN helicase acts like a molecular repair tool that keeps these fragile tumors alive. This study reveals, in atomic detail, how WRN moves along DNA and how experimental drugs can jam its motion—offering a roadmap for new treatments that selectively kill these vulnerable cancer cells while sparing healthy tissue.
The DNA handyman under the microscope
WRN helicase is part of the cell’s maintenance crew, helping unwind DNA so damage can be spotted and fixed. People born without working WRN develop Werner syndrome, a rare disorder marked by premature aging, showing how crucial this protein is for genome upkeep. Tumors with “microsatellite instability–high” (MSI-H)—a common defect in colorectal and some other cancers—turn out to be especially dependent on WRN. When WRN is shut down in these cells, their already shaky DNA quickly falls apart, and the cancer cells die. That makes WRN an appealing drug target, but until now scientists lacked a clear view of how the protein changes shape as it grabs DNA, burns chemical fuel, and moves along the genetic material.
Watching WRN flex and breathe
The authors used X‑ray crystallography to capture several high‑resolution “snapshots” of the human WRN helicase core. They solved structures of WRN alone and of WRN bound to single‑stranded DNA and a non‑breakable ATP‑like fuel molecule. These views revealed that the protein is built from two main lobes connected by a flexible hinge that behaves like a breathing joint. In its resting, fuel‑free state, WRN adopts a compact “closed” form in which the lobes sit close together. When ATP‑like fuel and DNA are present, the lobes swing apart into a more “open” configuration that can cradle DNA in a positively charged groove. A small aromatic loop within WRN reshapes into a short helix and wedges between DNA bases, acting like a ratchet that helps the protein step forward without sliding backward.
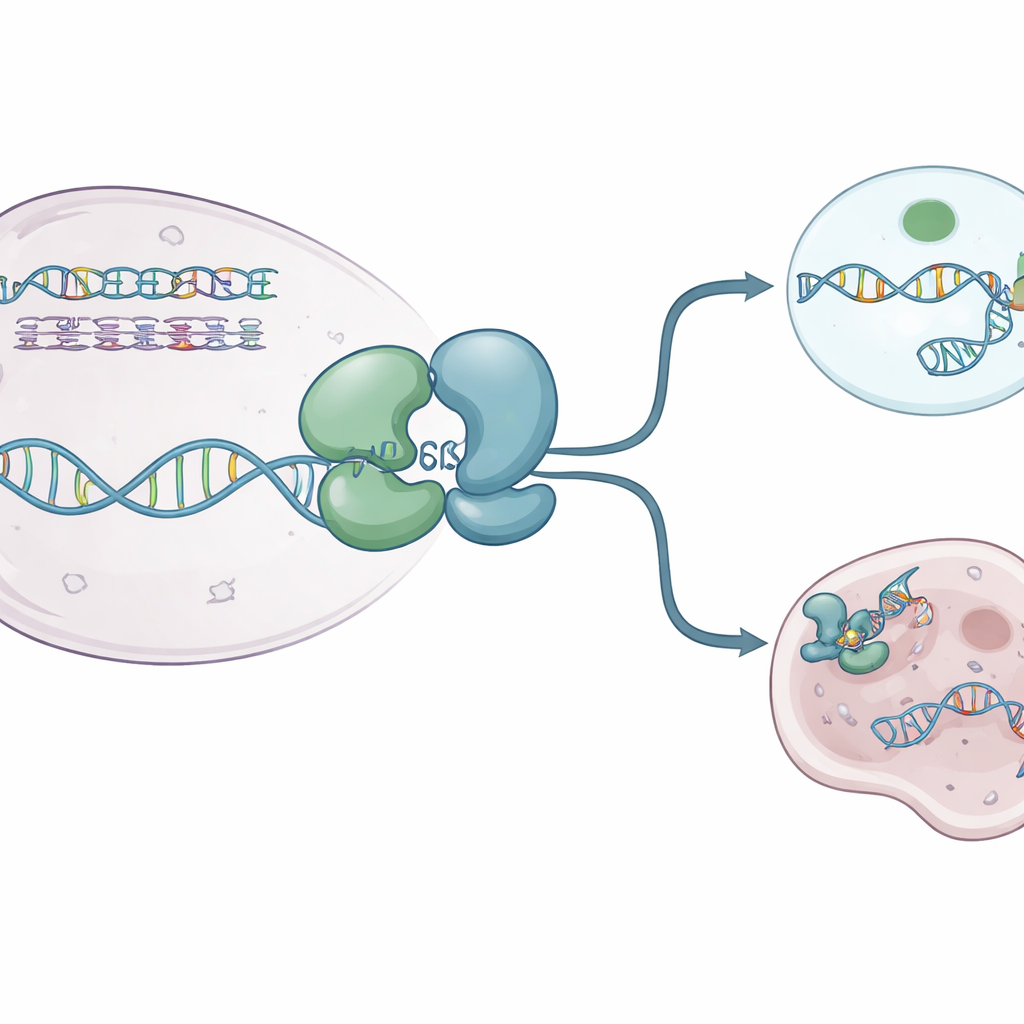
How current drugs switch WRN off
Several WRN‑blocking compounds have recently entered clinical testing. They do not clog the protein’s active groove directly. Instead, they act at a distance, gripping the hinge region and locking WRN into shapes that cannot properly engage DNA. Some molecules, such as HRO761 and a related clinical candidate from GSK, flip one lobe roughly 180 degrees relative to the other, creating a dramatic “twisted” form that sits off DNA. Others, including VVD‑133214 and a closely related compound studied here, pin WRN in a tightly “closed” arrangement that fails to expose key DNA‑gripping surfaces. Biophysical experiments confirmed that when these drugs are bound, WRN can no longer form a stable complex with single‑stranded DNA, effectively cutting the link between ATP use and DNA unwinding.
How cancer cells learn to escape
To see how tumors might evade these drugs, the team grew MSI‑H colorectal cancer cells in the presence of WRN inhibitors for weeks to months. Resistant populations emerged quickly. Genetic analysis showed that, in each case, the cells had acquired a single, precise change in the WRN gene near the drug‑binding hinge. One mutation weakened the effect of the “twisted‑state” inhibitor HRO761, while a different mutation reduced sensitivity to VVD‑133214, which favors the closed state. In addition, many resistant cells ramped up WRN production, giving themselves more copies of the target and partially diluting drug impact. These findings echo resistance patterns seen with other targeted cancer drugs, where small structural tweaks in the target protein or overexpression can blunt treatment.
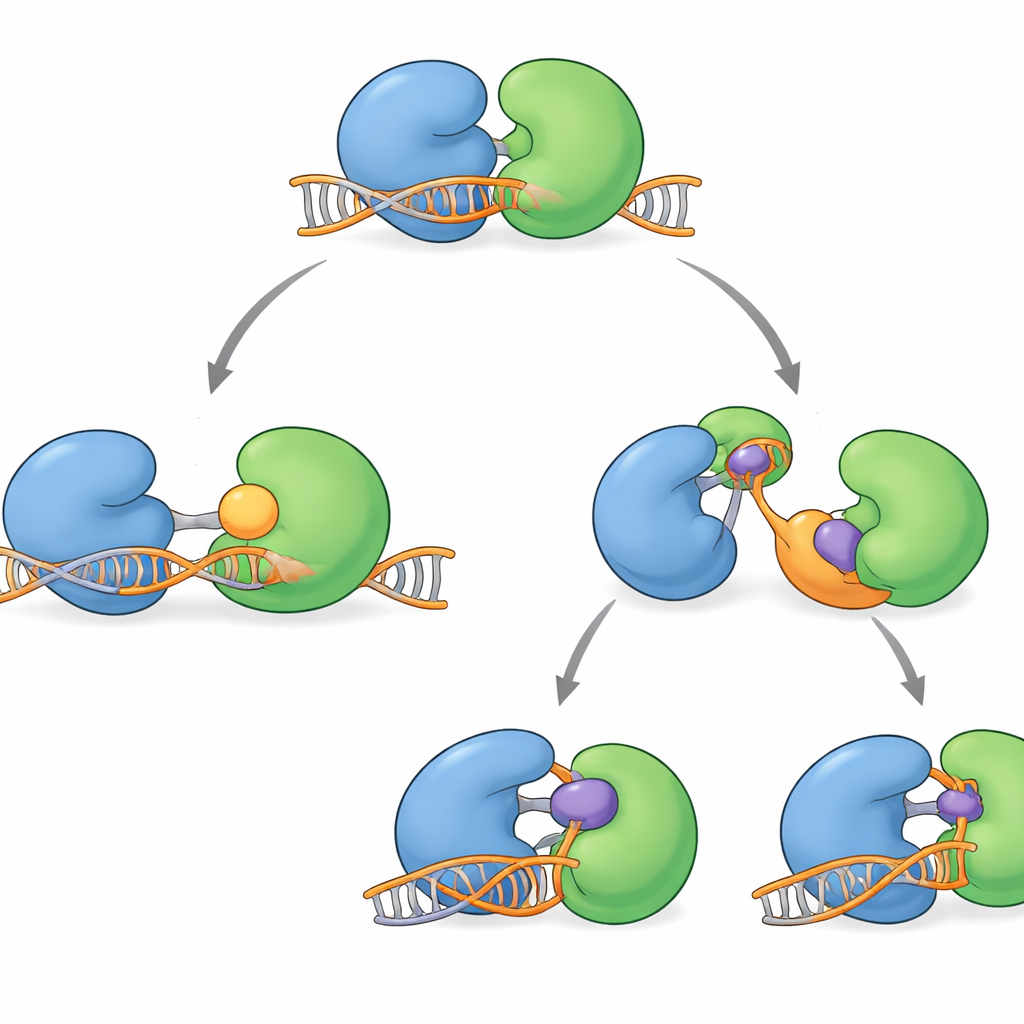
Looking ahead to smarter WRN blockers
Taken together, the new structures map out a full working cycle for WRN as it latches onto DNA, steps along, and then resets. They also show that today’s clinical compounds mainly stabilize “off‑DNA” forms of the protein. For a general reader, the key point is that we now understand where WRN is vulnerable and how tumors may adapt. This suggests a next wave of inhibitors that bind WRN when it is locked onto DNA, potentially trapping it in a toxic state—much like successful drugs that trap other DNA repair enzymes. Such “on‑DNA” WRN blockers, used alone or alongside existing agents, could offer more durable ways to exploit the hidden weakness of MSI‑H cancers.
Citation: Fletcher, C.T., Mornement, A.A., Barrett, C. et al. Structural insights into WRN helicase reveal conformational states and opportunities for MSI-H cancer drug discovery. Commun Biol 9, 334 (2026). https://doi.org/10.1038/s42003-026-09584-0
Keywords: WRN helicase, microsatellite instability, DNA repair, allosteric inhibitors, drug resistance