Clear Sky Science · en
Quality assessment of RNA 3D structure models using deep learning and intermediate 2D maps
Why judging RNA shapes matters
Inside every cell, RNA molecules twist and fold into intricate three-dimensional shapes that help control which genes turn on, guide chemical reactions, and even fend off viruses. Today, powerful computer programs can guess many of these shapes, but scientists still face a basic problem: when the computer produces dozens or hundreds of candidate shapes for the same RNA, which one is actually close to reality? This paper introduces RNArank, an artificial-intelligence tool designed to tackle that question by grading RNA 3D models, much like a structural quality inspector, so researchers can focus on the most trustworthy predictions.
A new inspector for RNA models
RNArank is built to judge the quality of a proposed RNA shape without needing to know how that shape was produced. Whether the model comes from a deep-learning system, a physics-based simulation, or a human expert, RNArank looks only at the final 3D coordinates. It then asks, in effect, “Does this structure look like a realistic RNA?” This kind of independent quality assessment is essential because, unlike in the protein world where tools such as AlphaFold often deliver highly reliable answers out of the box, RNA prediction still benefits from mixing multiple methods and human insight—and from a smart way to rank the resulting models.
Teaching the AI what good RNA looks like
To train RNArank, the authors assembled about 200,000 RNA structures, covering a wide spectrum from clearly wrong to nearly perfect. These were built from known experimental structures using a wide variety of approaches, including modern deep-learning predictors, molecular dynamics simulations that mimic atomic motion, and deliberate distortions of accurate structures to create “decoys.” For each model, the team computed how closely it matched the real, experimentally determined RNA, using a refined accuracy score tailored to RNA called lDDTRNA. This score focuses on how well the distances between pairs of nucleotides are reproduced, capturing both overall fold and local details without being overly sensitive to molecule length.
How RNArank reads and scores an RNA
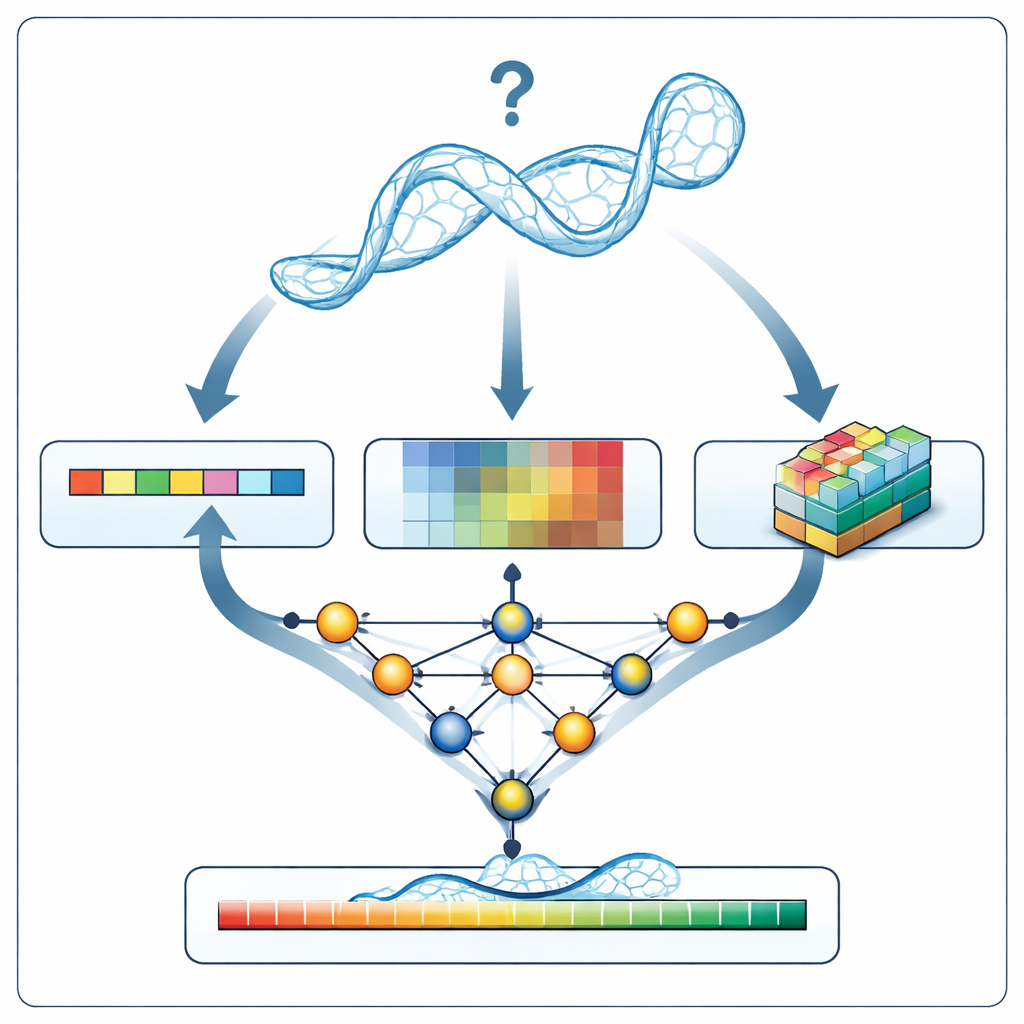
When RNArank examines a new RNA model, it first translates the structure into three kinds of information: a 1D description of the sequence and backbone geometry along the chain, 2D descriptions of how each pair of nucleotides relates to one another (their distances, estimated interaction energies, and possible atomic clashes), and 3D “voxel” snapshots, small grids that capture the local cloud of atoms around each nucleotide. A multi-part neural network weaves these clues together into a unified picture and then predicts two intermediate 2D maps: which nucleotides are likely to be in contact, and how far each modeled distance is likely to deviate from the unknown true structure. From these maps, RNArank reconstructs both a per-nucleotide confidence score and an overall score for the entire RNA model.
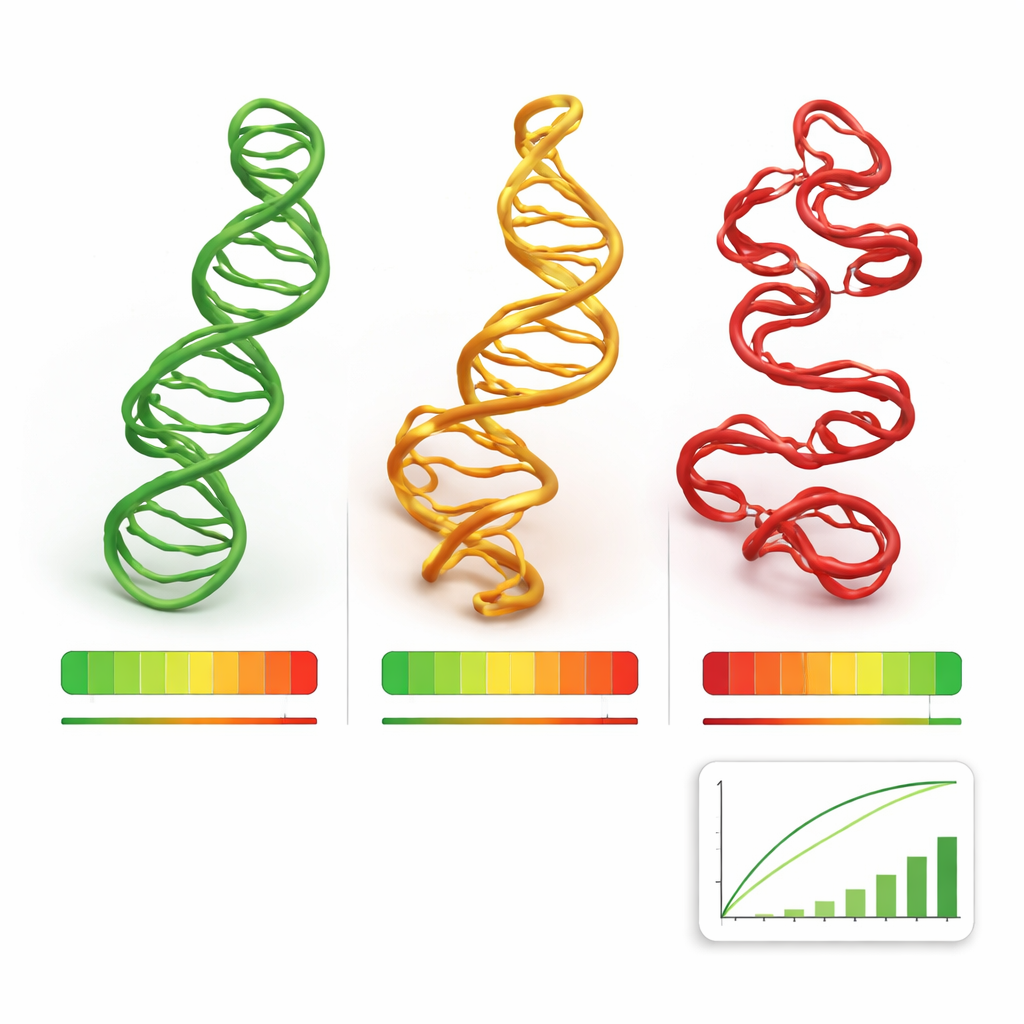
Putting the method to the test
The team benchmarked RNArank on three demanding datasets: a set of 24 newly solved RNAs from the Protein Data Bank, and the RNA targets from two international blind prediction contests, CASP15 and CASP16, where many groups submit models without knowing the answers in advance. Across thousands of candidate structures, RNArank’s scores tracked the true model quality more closely than several established energy-based scoring methods and other deep-learning approaches. It was especially strong at picking out the best or near-best models from a pool, and at identifying which parts of a structure were likely to be unreliable. The authors also showed that RNArank maintained its performance even on RNAs that were clearly different in sequence from those it had seen during training, a sign of genuine generalization rather than memorization.
Limits today and prospects tomorrow
RNArank is not perfect: it still struggles with especially flexible RNAs that adopt many shapes, and with RNAs that change their form when clamped by proteins inside large molecular machines. However, it is fast enough to handle many models for RNAs of a few hundred nucleotides in just seconds, and it already helps automated servers choose higher-quality predictions in community-wide tests. By providing a method-agnostic, structure-only judge of RNA models, RNArank gives biologists a sharper filter for turning raw computer output into reliable structural hypotheses, bringing the field a step closer to routine, trustworthy prediction of RNA shapes and, in turn, to deeper insights into how these versatile molecules work.
Citation: Liu, X., Wang, W., Du, Z. et al. Quality assessment of RNA 3D structure models using deep learning and intermediate 2D maps. Commun Biol 9, 293 (2026). https://doi.org/10.1038/s42003-026-09582-2
Keywords: RNA 3D structure, deep learning, model quality assessment, structural bioinformatics, RNArank