Clear Sky Science · en
MHC1-TIP enables single-tube multimodal immunopeptidome profiling and uncovers intratumoral heterogeneity in antigen presentation
Peeking at Cancer’s ID Cards
Every cell carries tiny molecular “ID cards” on its surface that help the immune system decide whether that cell is healthy or dangerous. In cancer, these ID cards—short protein fragments called antigens—can reveal what makes a tumor cell abnormal and mark it for destruction by immune cells. This study introduces a new lab method, MHC1-TIP, that makes it much easier and cheaper to read these ID cards from very small amounts of patient tissue, opening doors for more precise cancer immunotherapies.
Why Cell Surface Signals Matter
Our immune system constantly inspects cells by checking antigens displayed on special molecules called MHC class I. Tumor cells often show unusual antigens that can, in principle, be recognized by T cells and targeted by therapies such as personalized vaccines. However, actually measuring which antigens are present on real patient samples has been technically demanding. Traditional methods need huge numbers of cells, many processing steps, and expensive antibodies, making them poorly suited to tiny biopsies or scarce clinical samples. At the same time, tumors are not uniform: different regions can express different proteins, raising the possibility that antigen presentation also varies from place to place inside a single tumor.
A One-Tube Shortcut to Cancer Antigens
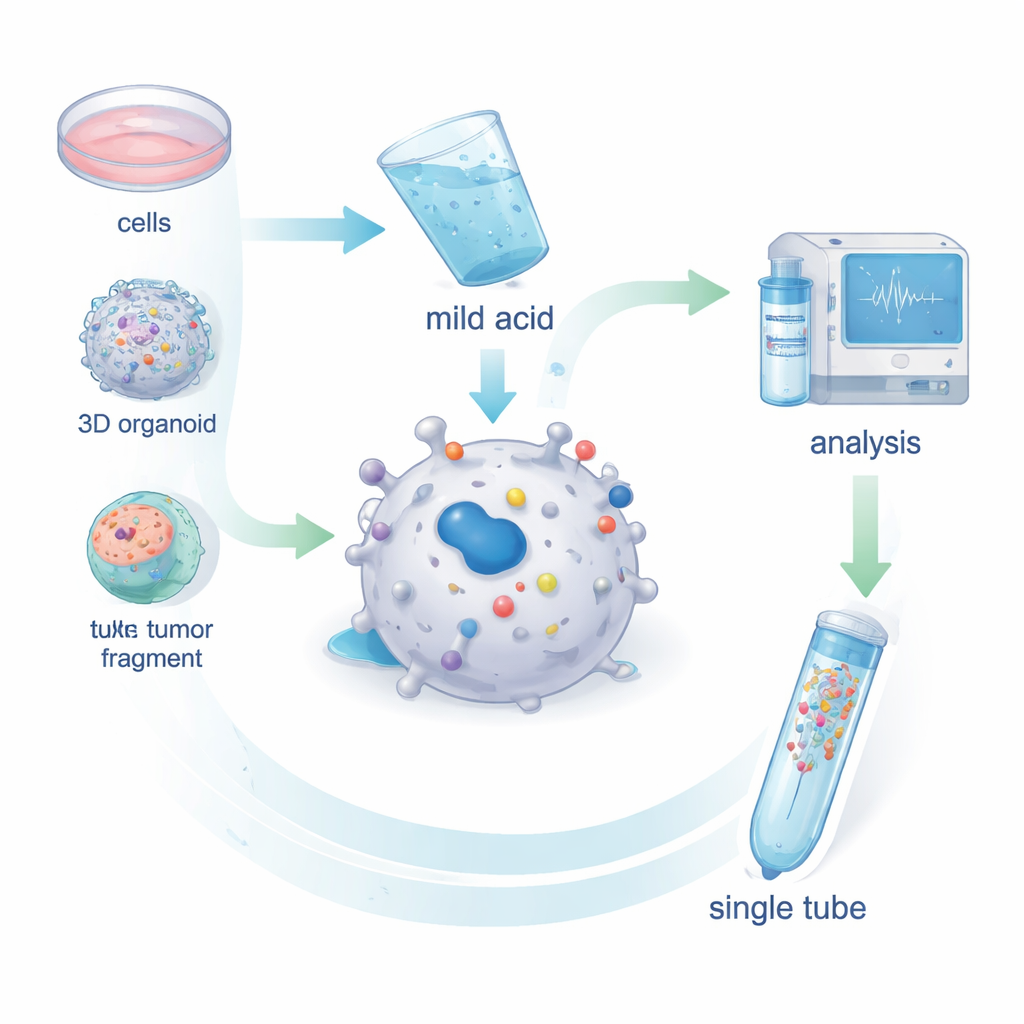
The researchers developed MHC1-TIP (MHC-I 1-Tube Immunopeptidomics) as a streamlined way to collect antigens from living cells. Instead of breaking cells open and fishing out MHC molecules with antibodies, they briefly wash intact cells—or tiny tumor fragments—with a mild acidic solution. This gentle treatment causes the antigens sitting in the MHC groove to fall off without killing the cells. The freed peptides then pass through a small filter that removes larger debris and are trapped on a tiny column inside a single tube. From there, they go straight into a mass spectrometer, which identifies the peptide sequences. Tests in melanoma cells showed that this mild acid step removes nearly all surface MHC–antigen complexes and that the recovered peptides look just like genuine MHC-bound antigens in terms of length and sequence patterns.
More Data from Less Sample
MHC1-TIP was designed to work with very few cells and still deliver many identifiable antigens. By pairing the new workflow with a modern mass spectrometry mode called data-independent acquisition, the team detected hundreds of antigens from as few as 100,000 cells and thousands from a few million cells—similar depth to the field’s gold-standard antibody method, but with less material and lower cost. The method also worked on patient-derived organoids, which are 3D mini-tumors grown in the lab, and on minute ex vivo tumor fragments smaller than a cubic millimeter. Importantly, because the acid wash leaves cells largely intact, the leftover material can be used for full-scale protein profiling from the very same sample, allowing direct comparison between how much of a given protein is present and how much of it is actually presented as an antigen.
Hidden Differences Inside a Single Tumor
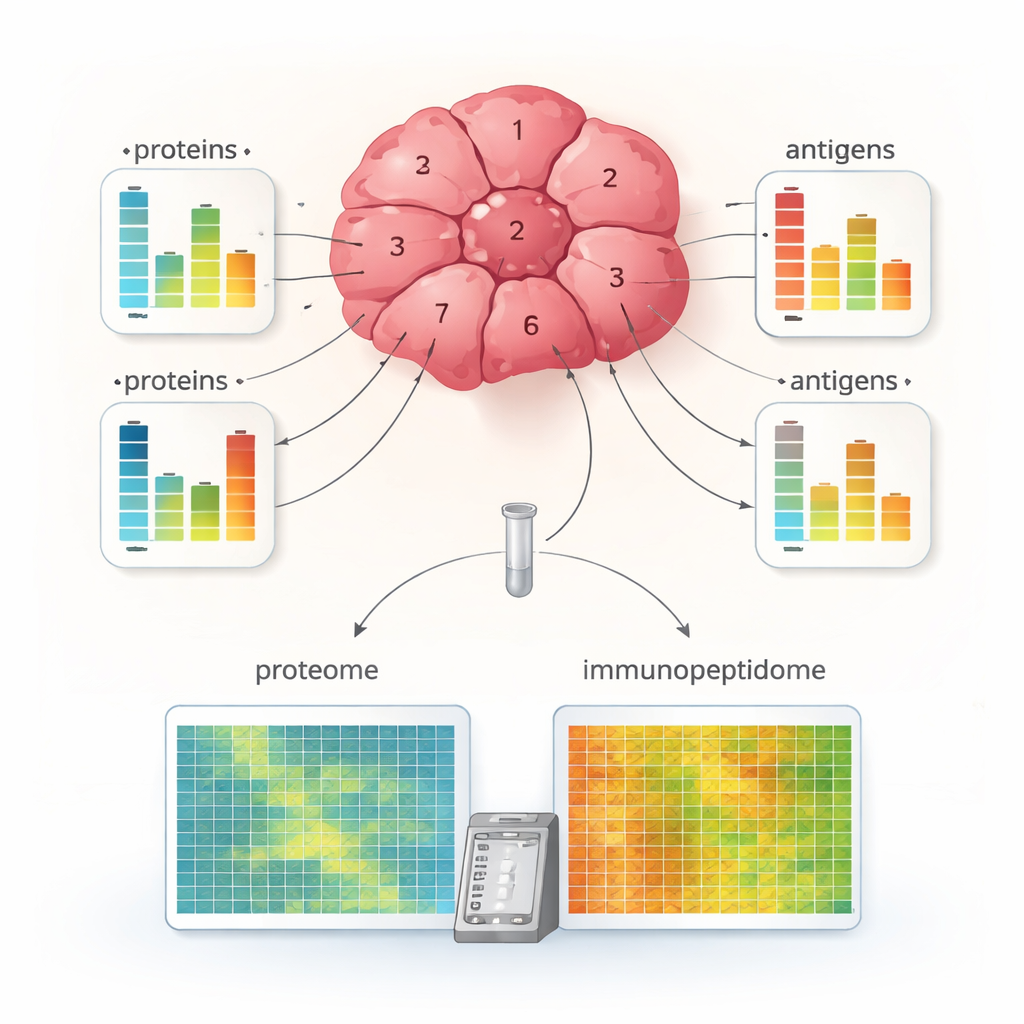
Applying MHC1-TIP to multiple small fragments taken from different regions of a renal cell carcinoma tumor revealed striking internal diversity. Some fragments showed rich antigen presentation, while others displayed very few antigens, even though their overall protein content could be similar. For many proteins, changes in antigen levels did not match changes in their protein abundance, suggesting that antigen processing and loading onto MHC molecules are regulated independently of how much of the parent protein is present. The team also combined antigen data with markers of immune cells and MHC machinery, identifying tumor regions that were “immune-hot,” with high antigen display and signs of active T cells, and “immune-cold,” with fewer antigens and less apparent immune attack. This level of detail has rarely been possible from such tiny tissue pieces.
What This Means for Future Cancer Care
To a non-specialist, the main message is that simply measuring which proteins a tumor makes is not enough to know what the immune system can actually see. The new MHC1-TIP method offers a practical way to read the true antigen display from very small, clinically realistic samples, while at the same time measuring the broader protein landscape. The discovery that antigen presentation can vary greatly across different parts of the same tumor, and does not always follow protein levels, warns against choosing vaccine or T-cell targets based only on gene or protein expression. In the future, approaches like MHC1-TIP could help design more effective, personalized immunotherapies by focusing on antigens that are genuinely shown on the surface and more consistently presented throughout the tumor.
Citation: Bathini, M., Bocaniciu, D., Johnson, F.D. et al. MHC1-TIP enables single-tube multimodal immunopeptidome profiling and uncovers intratumoral heterogeneity in antigen presentation. Commun Biol 9, 296 (2026). https://doi.org/10.1038/s42003-026-09570-6
Keywords: antigen presentation, immunopeptidomics, cancer immunotherapy, tumor heterogeneity, mass spectrometry