Clear Sky Science · en
Re-evaluating Gα protein–response element specificity in GPCR signaling
Why tiny cell switches matter to medicine
Many of today’s drugs work by flipping molecular “switches” on the surface of our cells called G‑protein‑coupled receptors, or GPCRs. These switches pass messages to the cell interior through helper proteins known as G proteins and eventually turn genes on or off. For decades, researchers have relied on simple light‑producing tests to see which G proteins a given drug or receptor is using. This study asks a basic but critical question: are those tests really telling us what we think they are—and can we still trust them when designing the next generation of precision medicines?
How scientists usually read GPCR signals
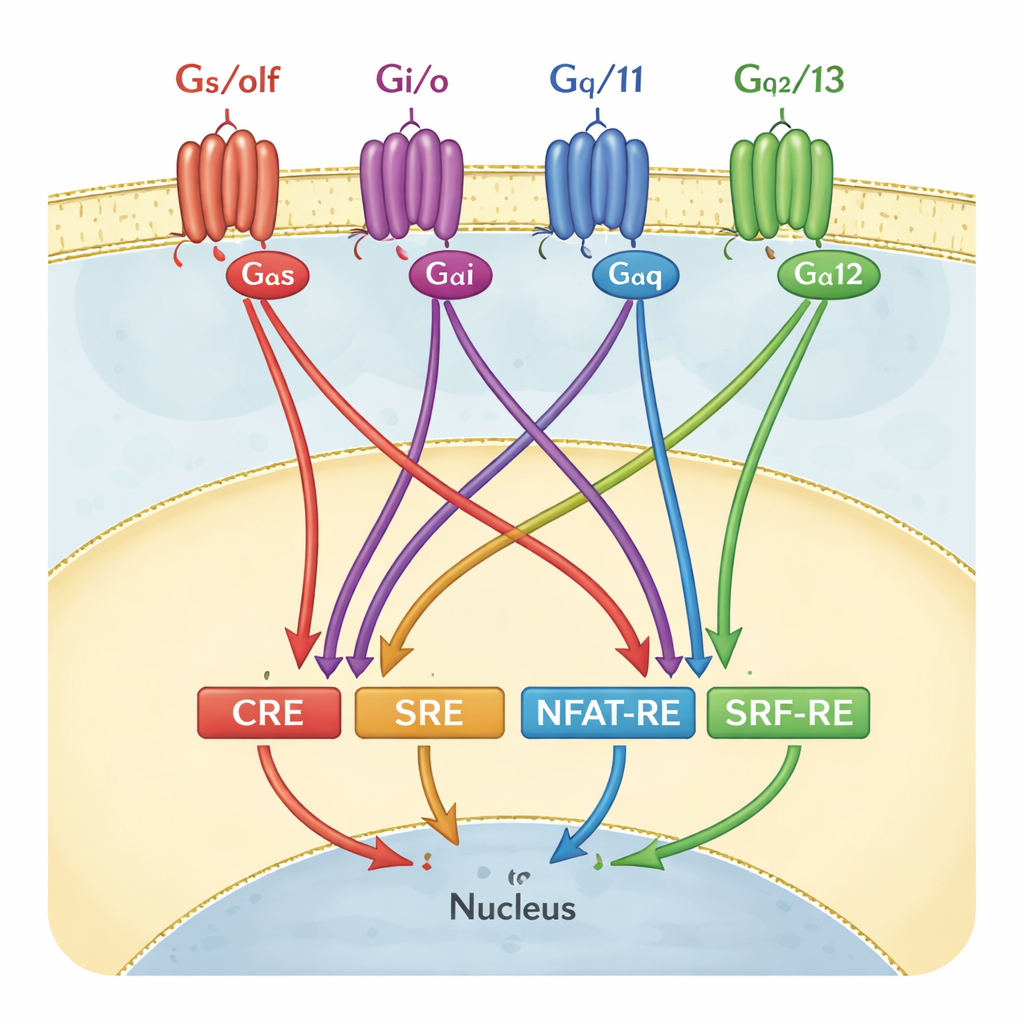
To track GPCR activity, many labs use reporter gene assays. In these tests, cells are engineered so that when a particular gene switch, or “response element,” is activated, the cells glow. Four such switches are especially popular: CRE, SRE, NFAT‑RE, and SRF‑RE. Each has long been treated as a stand‑in for one branch of G‑protein signaling—for example, CRE for the Gαs/olf family, or SRF‑RE for Gα12/13. In textbooks and screening pipelines, these pairings are often drawn almost like one‑to‑one wiring diagrams: turn on a specific G protein, and only one reporter lights up.
Putting the old wiring diagram to the test
The authors systematically challenged this picture using human HEK293 cells in which specific G‑protein families—or even all G proteins—were removed with genome editing. They then reintroduced individual G‑protein types one by one. Into these cells they expressed nine different GPCRs, chosen to represent receptors that either signal very selectively or through many G‑protein partners. By measuring how strongly each of the four reporters glowed across many drug concentrations, they could see which G proteins were truly required for each response element, and which ones could help out indirectly.
Overlapping pathways instead of clean channels
The results overturn the neat one‑receptor‑one‑reporter model. While CRE still depended mainly on Gαs/olf proteins—the classic cAMP‑raising pathway—other G‑protein families could noticeably influence CRE activity, but only when Gαs/olf was present. For the other three reporters, SRE, NFAT‑RE, and SRF‑RE, the Gαq/11 family emerged as the major driver, even though SRE and SRF‑RE were traditionally linked to other G‑protein branches. SRE and SRF‑RE behaved especially similarly, suggesting that they tap into shared downstream machinery rather than distinct, isolated routes. In many cases, additional G proteins such as Gα12/13 and Gαi/o provided extra push or background activity, highlighting a web of crosstalk rather than separate channels.
Why the cell’s network blurs our readouts
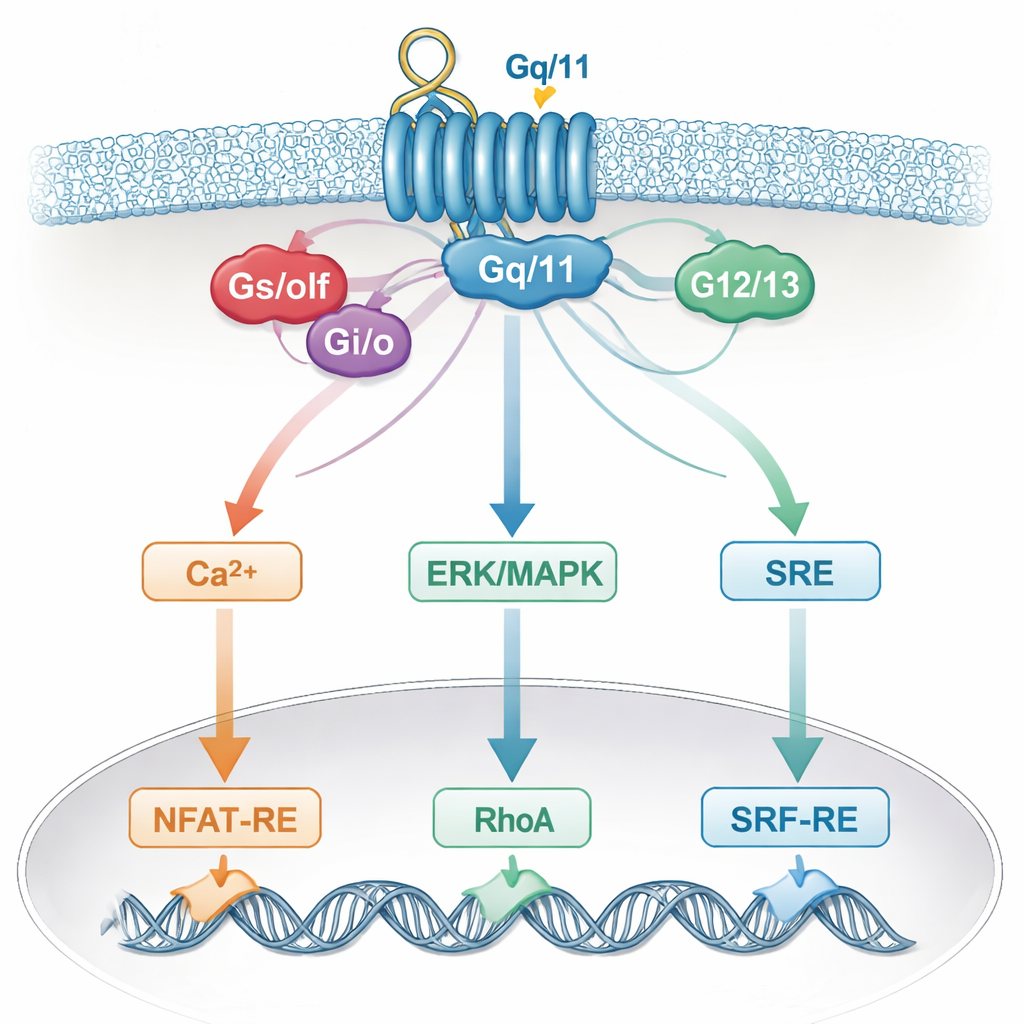
This blurred specificity likely reflects how real cells integrate signals. A single receptor can activate multiple G proteins, which then converge on common messengers like calcium, ERK/MAPK, RhoA, or cAMP, and on shared helpers such as Gβγ subunits. These shared pathways, in turn, feed into the same gene switches in the nucleus. As a result, a reporter that was thought to respond to one G‑protein family actually “listens” to several. Cell‑type‑specific factors and the exact set of receptors present add even more complexity, meaning that conclusions drawn from one cell system may not hold in another.
What this means for drug discovery
For non‑specialists, the key takeaway is that many widely used glow‑based GPCR tests are less specific than their labels suggest. They can still tell us that a receptor is active and roughly which broad pathways are engaged, but they are not reliable proof that a particular G‑protein family—and only that family—is involved. The authors argue that researchers and drug developers should treat these reporters as coarse indicators and pair them with more direct methods that watch proteins interact in real time. Doing so will give a truer picture of how potential medicines steer cellular signaling, and help avoid misleading shortcuts in the search for safer, more targeted therapies.
Citation: Saito, A., Kise, R., Yamaguchi, S. et al. Re-evaluating Gα protein–response element specificity in GPCR signaling. Commun Biol 9, 288 (2026). https://doi.org/10.1038/s42003-026-09569-z
Keywords: GPCR signaling, G proteins, reporter assays, cell signaling networks, drug discovery