Clear Sky Science · en
The Neijiang pig T2T genome reveals domestication history and germplasm traits of Southwest Chinese local breeds
Why a pig genome matters to people
Pigs are not only a major source of meat; they are also key models for understanding genetics, evolution, and even human disease. This study delivers the first nearly gap-free, "telomere-to-telomere" (T2T) genome for the Neijiang pig, a traditional breed from Southwest China. By reading this genome end to end, researchers can trace how these animals were shaped by history, environment, and breeding—and use that knowledge to protect rare local pigs while improving future herds.

A detailed map of a traditional Chinese pig
The Neijiang pig has been raised in Sichuan Province for about 1,800 years and is prized for its high fertility, toughness, and distinctive head shape. Yet like many traditional breeds, it has been overshadowed by modern commercial pigs, putting its genetic diversity at risk. Earlier reference genomes for pigs were based on European breeds and differ by around 7% from Asian pigs, limiting their use for studying China’s local animals. In this work, scientists combined cutting-edge DNA sequencing methods to build a highly accurate reference specifically for the Neijiang pig, called NJP-T2T.
Building a genome from end to end
To assemble NJP-T2T, the team used long but precise reads from PacBio HiFi sequencing, ultra-long reads from Oxford Nanopore, and 3D genome contact maps from Hi-C technology. They first created long stretches of continuous DNA sequence, then used chromosome contact data to arrange them into 19 chromosomes (18 autosomes plus the X chromosome). Nanopore reads were then used to close almost all remaining gaps, leaving just a tiny 500-base gap on one chromosome. The final genome, about 2.55 billion DNA letters long, has excellent accuracy and completeness, outperforming the standard pig reference when data from both Chinese and foreign pig breeds are aligned to it.
Hidden regions and special gene families
One strength of a T2T genome is that it reveals regions that were previously missing or badly assembled, including the highly repetitive stretches found at chromosome centers (centromeres) and ends (telomeres). In Neijiang pigs, the researchers mapped centromeres across all chromosomes by tracking patterns such as low gene density, dense repeats, and characteristic satellite DNA sequences, and confirmed telomere structures at every chromosome tip. They also identified more than a thousand genes in regions that older genomes had left unresolved. Comparing Neijiang with other pig breeds, the study found 75 gene families and over 300 genes unique to this breed, many involved in sensing the environment, immunity, metabolism, and aging—traits that likely support its hardiness and adaptability.
From population history to fertility and head shape
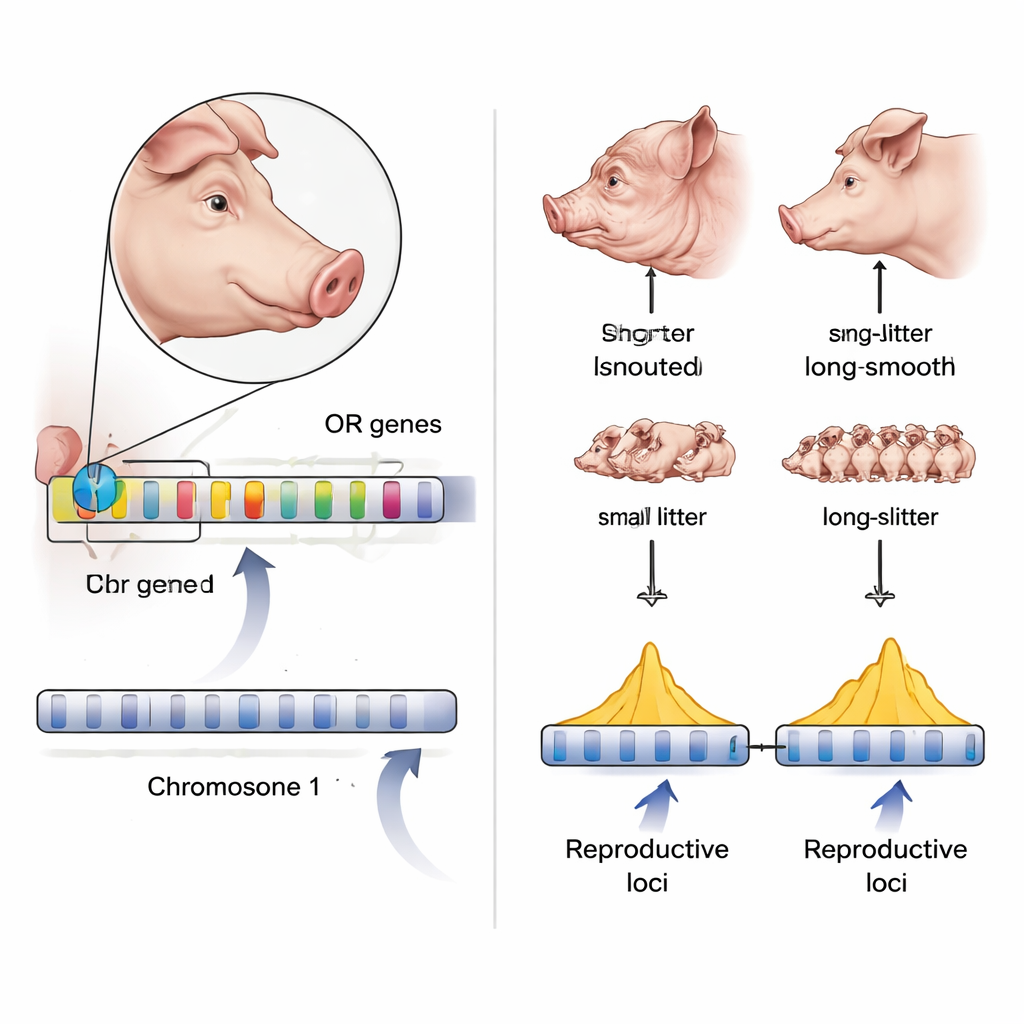
Armed with this detailed genome, the team examined a conservation herd of Neijiang pigs. They found moderate genetic diversity and generally low inbreeding, although some individuals showed signs of past inbreeding, underscoring the need for careful management. Using genome-wide association studies, they linked specific DNA sites to litter size and number of piglets born alive, pinpointing genes involved in early embryo development as promising targets for selective breeding. They also probed one of the breed’s most striking features: contrasting head shapes, ranging from a short, wrinkled “lion head” type to a longer, smoother form. By comparing pigs with extreme head types, they detected strong selection signals near clusters of olfactory receptor genes—genes that help pigs smell. Several of these smell genes are more active in pigs with longer snouts, suggesting that changes in local feeding practices and reliance on smell may have helped drive head-shape evolution.
What this means for pigs and people
For non-specialists, the bottom line is that NJP-T2T is akin to an ultra-high-resolution map of a rare regional pig. It shows how centuries of farming and environment have sculpted the Neijiang’s body and behavior and reveals genetic levers that influence fertility, resilience, and even skull shape. This map will help conserve a culturally important Chinese breed, guide more precise and sustainable pig breeding, and add to a growing library of complete animal genomes that deepen our understanding of domestication and adaptation.
Citation: Chen, D., Cui, S., Zhao, Z. et al. The Neijiang pig T2T genome reveals domestication history and germplasm traits of Southwest Chinese local breeds. Commun Biol 9, 278 (2026). https://doi.org/10.1038/s42003-026-09557-3
Keywords: Neijiang pig, genome assembly, domestication, reproductive traits, head morphology