Clear Sky Science · en
Targeted blocking of gene splicing can dysregulate intron-embedded primary microRNAs
Hidden Messages Inside Our Genes
Many new gene therapies work by nudging our cells’ RNA messages, but those RNAs often hide extra “notes in the margins” that scientists are only beginning to understand. This paper shows that drugs designed to correct faulty splicing of genes can accidentally crank up tiny regulatory RNAs buried inside introns—the non‑coding stretches between gene segments—with surprising effects on brain wiring and development.
Gene Therapies That Rewrite RNA
Antisense oligonucleotides, or ASOs, are short synthetic pieces of genetic material that stick to RNA and change how it is read or processed. Several ASOs are already being tested or used for rare genetic diseases, especially those caused by faults in how RNA is spliced—the step that stitches together coding segments of a gene. Most safety tests focus on whether an ASO restores the right protein and avoids obvious side effects. But many genes also carry non‑coding RNAs, including microRNAs, inside their introns. These tiny molecules can strongly tune gene activity, and up to half of known microRNAs in vertebrates come from introns. The new study asks: what happens to these embedded microRNAs when we force changes in splicing with ASOs?
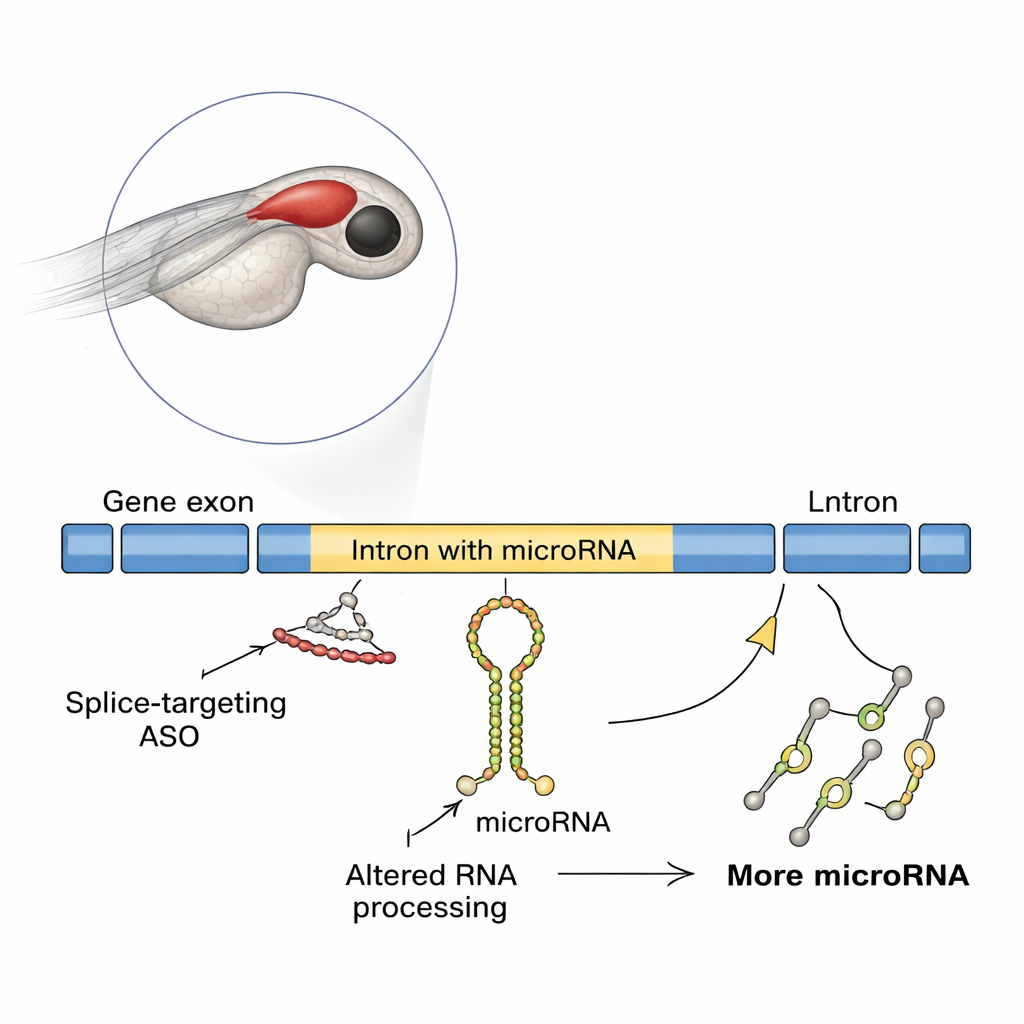
A Surprising Effect in Developing Fish Brains
The researchers used zebrafish, a popular vertebrate model whose transparent embryos make brain development easy to track. They focused on a gene called slit3, which helps guide nerve fibers and contains a microRNA called mir-218a-1 inside one of its introns. When the team blocked the start of slit3 translation, development of hormone‑secreting pituitary axons looked normal. But when they used a different ASO that blocked splicing at a specific slit3 splice site, the axons that should project to the posterior pituitary were drastically reduced, even though the number of oxytocin‑producing neurons stayed the same. Molecular tests showed that this splice‑blocking ASO caused intron retention in the gene’s RNA, unexpectedly increasing the levels of slit3 transcript, its long intron, and the intron‑embedded primary mir-218a-1 transcript.
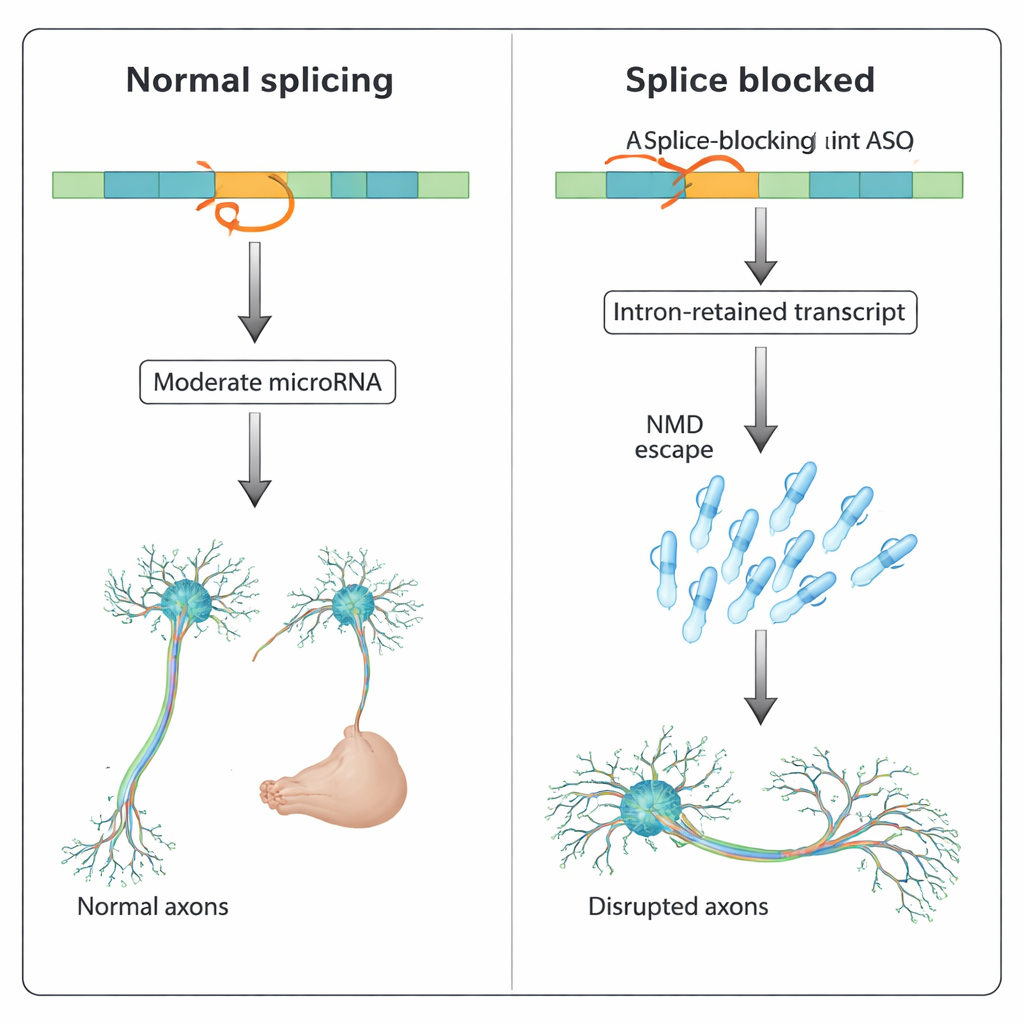
MicroRNA, Not Protein Loss, Drives the Defects
To untangle what caused the axonal problem, the authors compared several scenarios. Blocking mature mir-218a-1 alone did not change slit3 levels, and a separate ASO aimed at a different splice site near the microRNA did not produce axonal defects or boost mir-218a-1. Crucially, when the splice‑blocking slit3 ASO was combined with an ASO that neutralizes mir-218a-1, the axonal wiring largely recovered. Likewise, in zebrafish genetically lacking mir-218a-1, the same splice‑blocking slit3 ASO no longer caused axonal defects, whereas injecting a synthetic mir-218a mimic could mimic the phenotype in normal fish. These experiments point to elevated intronic microRNA—rather than a truncated Slit3 protein—as the main driver of the developmental disturbance.
A Broader Pattern Across Multiple Genes
The team then asked whether this effect is unique to slit3. They searched the zebrafish genome and found dozens of protein‑coding genes that host intronic microRNAs, many involved in development and signaling. Two such genes, pank2 and dnm2a, had previously shown severe developmental problems when their splicing was blocked by ASOs, including brain fluid buildup, heart swelling, and body deformities—problems that genetic knockouts of the same genes did not reproduce. In this study, co‑injecting ASOs that specifically blocked the intronic microRNAs mir-103 (in pank2) or mir-199-5p (in dnm2a) partially rescued these ASO‑induced phenotypes. In some cases, microRNA blockade also restored host gene expression, while in others it did not, suggesting that elevated microRNA activity itself can be harmful even when overall levels change only modestly.
What This Means for Future Treatments
The findings reveal that splice‑targeting ASOs can do more than cause exon skipping or intron retention: they can stabilize intron‑retained transcripts that evade the cell’s quality‑control system and boost intronic primary microRNAs, which then mis‑regulate networks of target genes. In developing neurons, such runaway microRNA activity can distort axon growth and connectivity. For drug developers, this means that unusual or “off‑target” phenotypes in animal tests may not always stem from generic toxicity pathways like p53 activation, but from unintended effects on concealed intronic RNAs. Carefully comparing translation‑blocking and splice‑blocking ASOs, and monitoring intronic microRNA expression, could help flag these risks early and refine ASO designs before they reach patients.
Citation: Ali, M.H., Ramesh, A.R., Nedunchezhian, N. et al. Targeted blocking of gene splicing can dysregulate intron-embedded primary microRNAs. Commun Biol 9, 252 (2026). https://doi.org/10.1038/s42003-026-09529-7
Keywords: antisense oligonucleotides, microRNA, RNA splicing, zebrafish development, gene therapy safety