Clear Sky Science · en
Epigenome-wide association study of circulating interleukin-6 connects DNA methylation to immunometabolic and inflammatory health
Why this matters for everyday health
Chronic, low-level inflammation quietly shapes our risk of diseases such as type 2 diabetes, heart disease, and inflammatory bowel disease. A key player in this process is a signalling protein in the blood called interleukin‑6 (IL‑6). This study asks a fundamental question: how does IL‑6 interact with our epigenome—the chemical tags on DNA that help control gene activity—and what does that mean for our long‑term health?
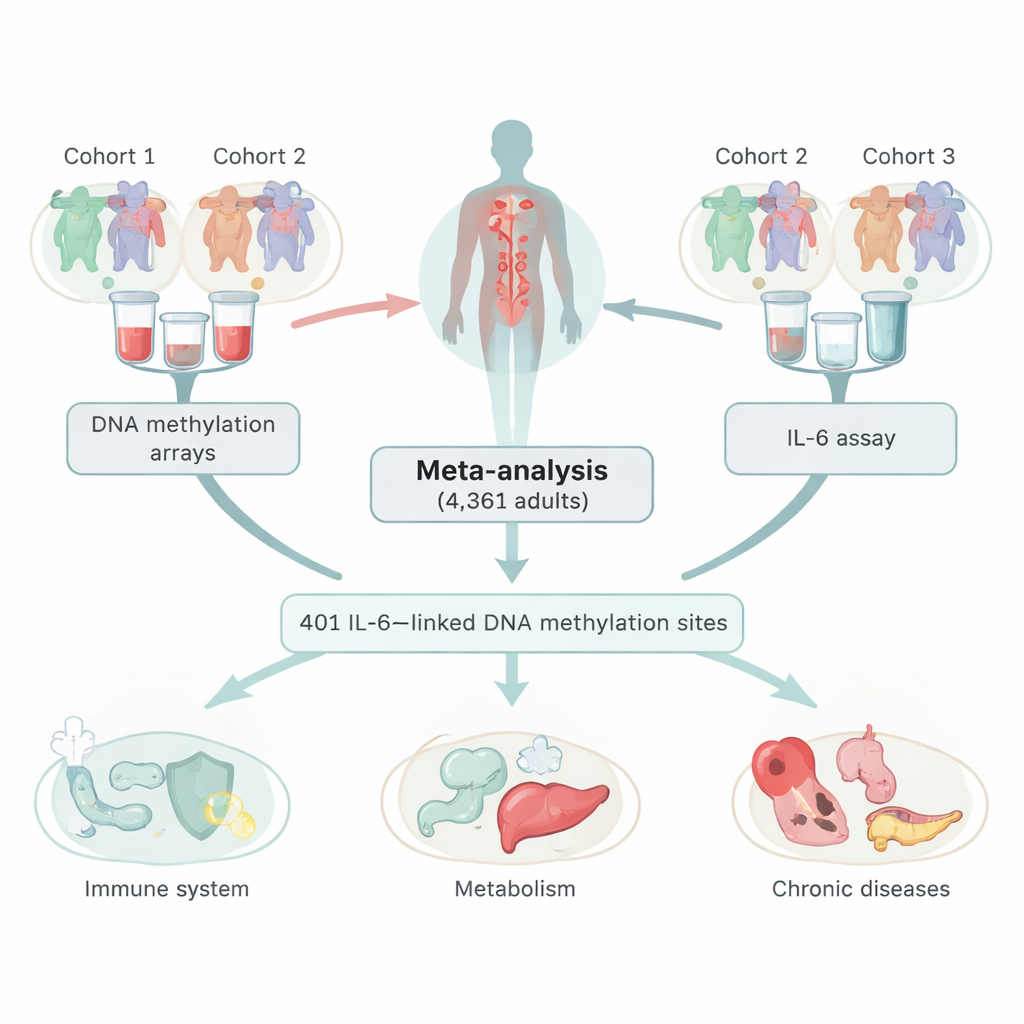
Looking at chemical markers on DNA
The researchers combined data from three large European studies, covering 4,361 adults who gave blood samples. In these samples, they measured both IL‑6 levels and DNA methylation, a common chemical mark on DNA that can turn genes up or down without changing the genetic code itself. By scanning over 400,000 sites across the genome, they identified 401 locations where methylation levels tracked with the amount of IL‑6 circulating in the blood. Most of these sites showed an inverse pattern: higher IL‑6 went hand‑in‑hand with lower methylation at those spots. The team carefully checked that these links were not simply side effects of smoking, differences in blood cell types, or another inflammation marker called C‑reactive protein (CRP).
Connecting inflammation to metabolism and disease
Next, the authors asked whether these IL‑6‑linked DNA sites overlapped with signals seen in other large epigenetic studies. They found strong enrichment for traits with an inflammatory or metabolic component, including body mass index, blood fats, blood pressure, blood sugar, type 2 diabetes, chronic kidney disease, and psychiatric and stress‑related conditions. Many of the same methylation spots had previously been tied to CRP, but the patterns suggested that IL‑6 and CRP each carry partly distinct epigenetic signatures. In other words, the chemical marks associated with IL‑6 are not just a carbon copy of those associated with CRP, and may provide extra information about a person’s inflammatory and metabolic state.
Where in the genome the action happens
To understand what these methylation sites might be doing, the team mapped them onto known regulatory regions of the genome. They found that IL‑6‑associated sites were concentrated in active control elements such as enhancers—stretches of DNA that help switch genes on—rather than in silent regions. These sites also sat near binding sequences for transcription factors that directly regulate IL‑6 and stress responses, including NF‑κB, Atf4, CHOP, and Nrf2. By combining methylation data with large gene‑expression datasets, the researchers linked over 400 genes to the IL‑6‑related sites. Many of these genes sit at the heart of immune and metabolic control, including SOCS3, IL6R, AIM2, IFI16, MTOR, and RORC, and participate in pathways that drive activation and energy use in a specialised type of T cell (Th17 cells) implicated in chronic inflammatory diseases.
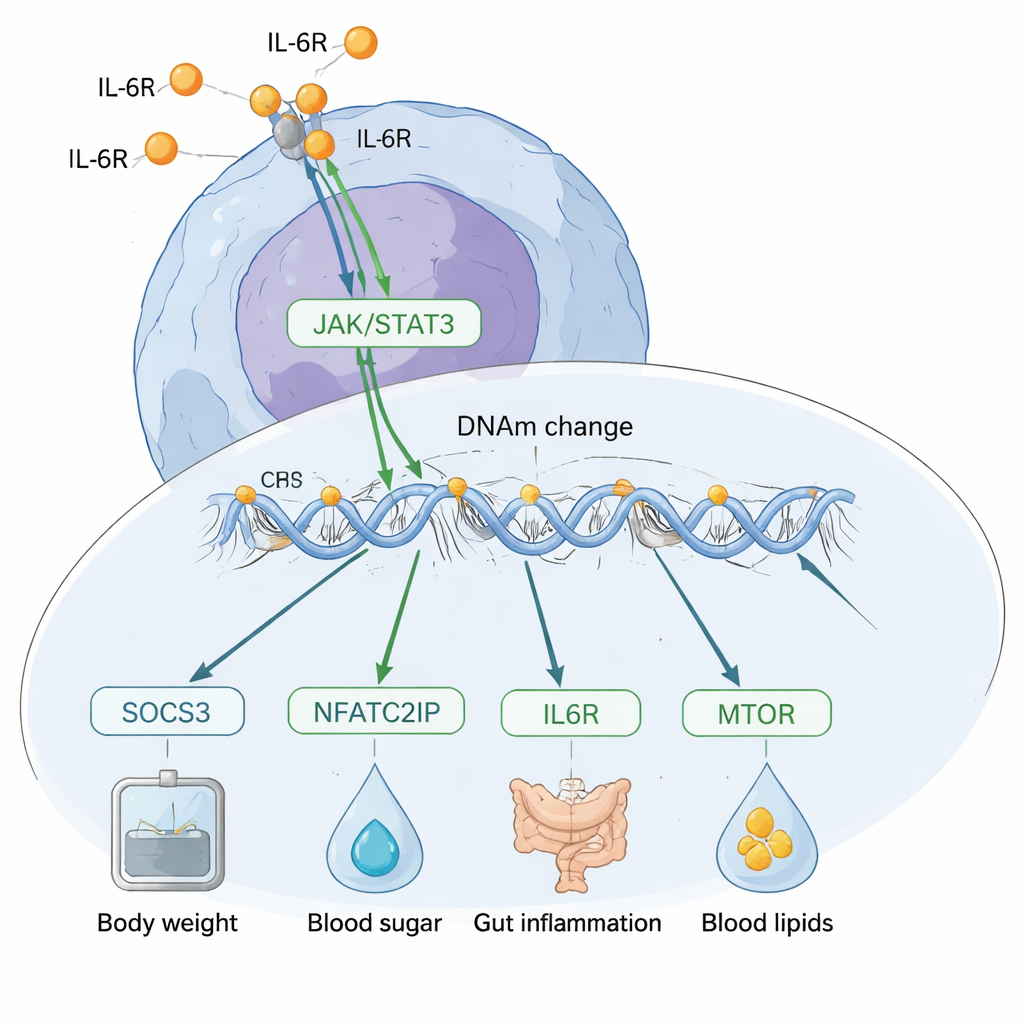
Which way does the cause and effect run?
A major challenge is to work out whether DNA methylation is causing changes in IL‑6, or whether IL‑6 is reshaping methylation. The team used several genetic tricks to probe this direction. Across the genome, their “triangulation” analyses supported a model in which IL‑6 mostly drives changes in DNA methylation, rather than the other way around. They then zoomed in on specific sites and found that methylation changes near the SOCS3 gene appear to mediate IL‑6’s impact on risk factors such as body weight, cholesterol, CRP levels, and inflammatory bowel disease. In contrast, one standout site near a gene regulator called NFATC2IP showed signs of influencing IL‑6 production itself, and also appeared to affect multiple conditions including body mass index, type 2 diabetes, and gut inflammation.
What this means for future prevention and treatment
For a non‑specialist reader, the central message is that IL‑6 and epigenetic marks on DNA are tightly intertwined in ways that touch many common diseases. Most of the time, IL‑6 seems to leave a chemical "footprint" on the genome, particularly in immune cells, marking and possibly stabilising pro‑inflammatory states. At a few key sites, however, methylation may help set how much IL‑6 is produced and how strongly immune pathways respond. These findings point to DNA methylation patterns as promising blood‑based indicators of inflammatory and metabolic health, and suggest that carefully targeted epigenetic changes—especially around genes like SOCS3 and NFATC2IP—could one day complement IL‑6‑blocking drugs in preventing or treating chronic disease.
Citation: Sinke, L., van Dongen, J., Delerue, T. et al. Epigenome-wide association study of circulating interleukin-6 connects DNA methylation to immunometabolic and inflammatory health. Commun Biol 9, 242 (2026). https://doi.org/10.1038/s42003-026-09520-2
Keywords: interleukin-6, DNA methylation, inflammation, metabolic disease, epigenetics