Clear Sky Science · en
Strategic advances for cryo-EM structural studies of small (<100 kDa) GPCRs
Why tiny cell switches matter for medicine
Much of modern medicine works by quietly nudging tiny switches that sit in our cell membranes, called G protein‑coupled receptors (GPCRs). These switches sense hormones, smells, and drugs, and about a third of all approved medicines act on them. To design safer, smarter drugs, scientists need detailed 3D pictures of these receptors, especially in their “off” or inactive forms, which many drugs specifically target. This article explains how researchers are learning to see some of the smallest GPCRs with cryo‑electron microscopy (cryo‑EM), a powerful imaging method that can look at molecules frozen in a near‑natural state.
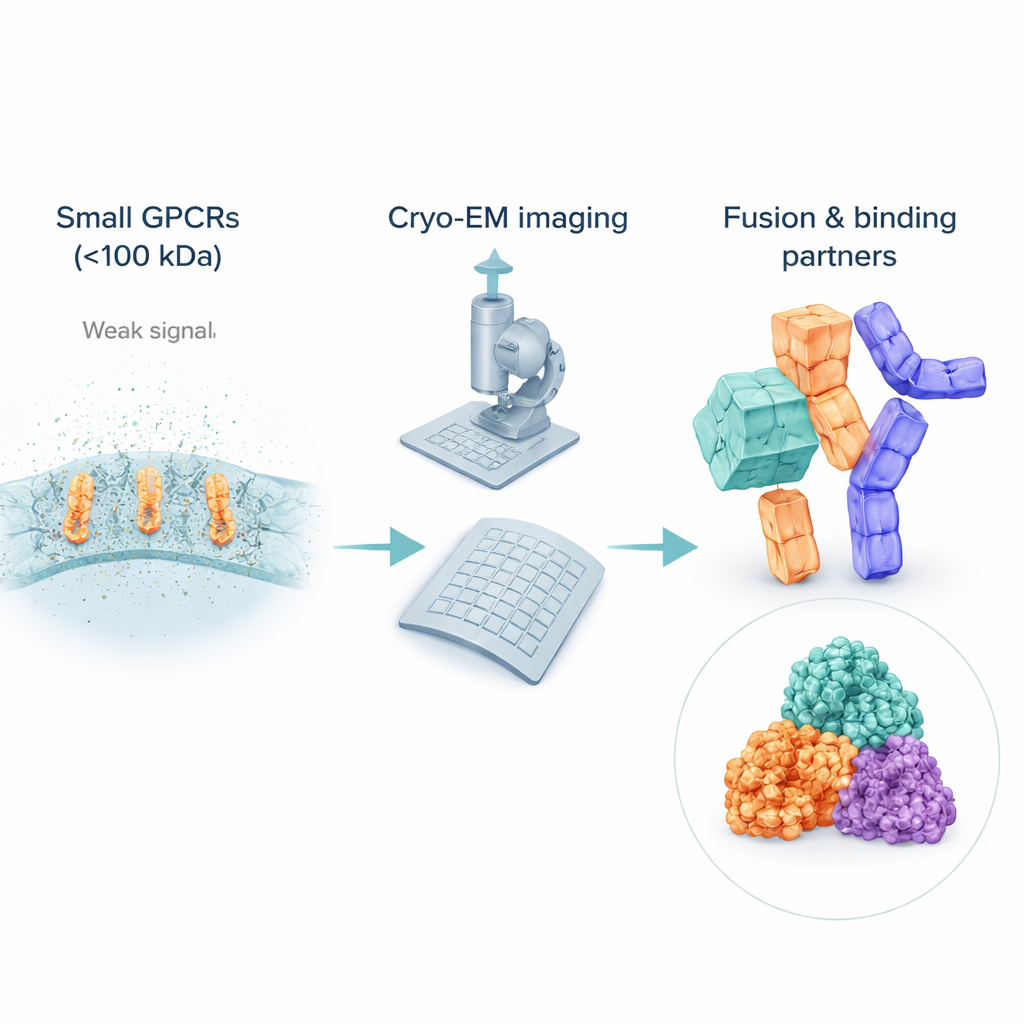
The challenge of seeing very small receptors
Although cryo‑EM has revolutionized structural biology, it still struggles with small, flexible molecules. Many inactive GPCRs weigh far less than 100 kilodaltons, making them difficult to spot and align in noisy images. To keep them soluble, scientists wrap these receptors in detergent bubbles or lipid mimics, which can actually look “bigger” to the electron beam than the receptor itself, drowning out the signal from the protein of interest. Unlike active receptors that are bound to bulky partners such as G proteins, inactive GPCRs often lack prominent external features, so computers have a hard time averaging many images into a sharp 3D view. As a result, most of the inactive GPCR structures still come from older X‑ray methods, leaving a gap in cryo‑EM coverage right where many drugs act.
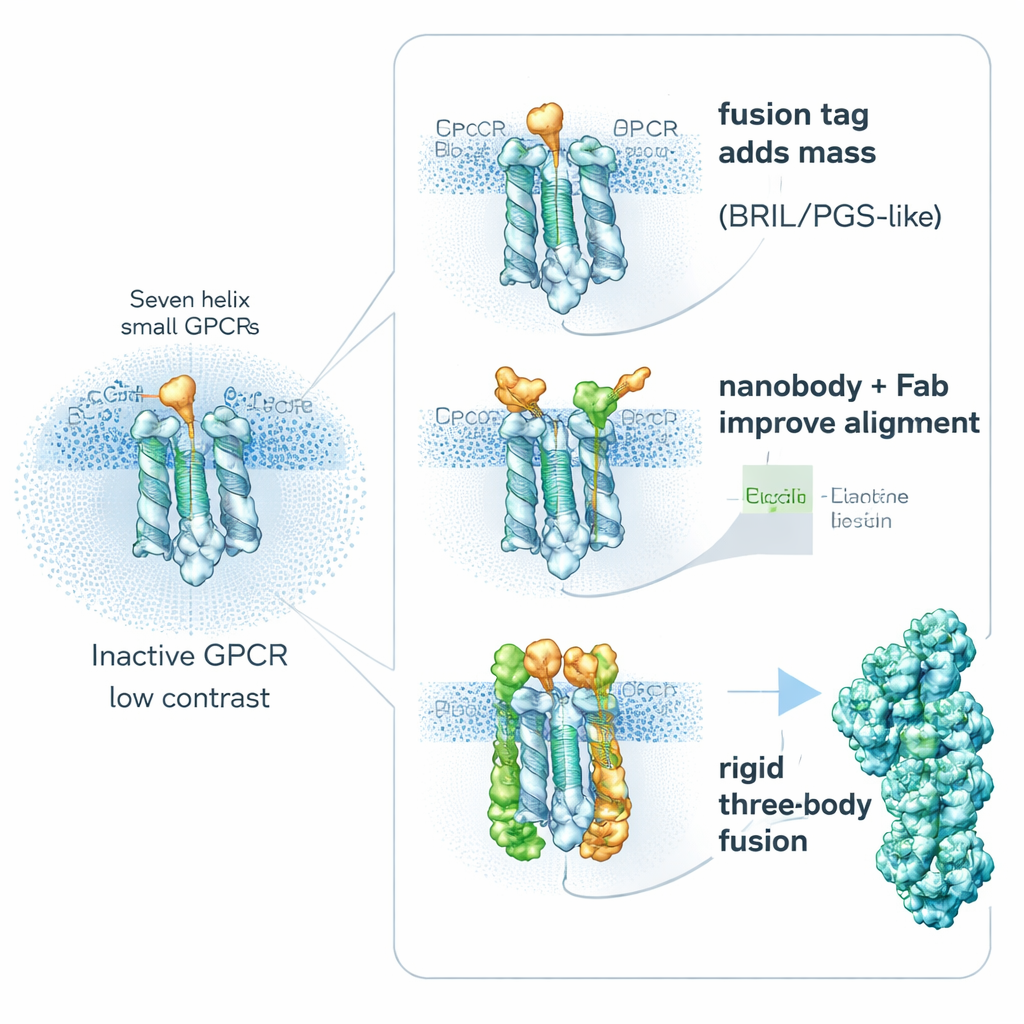
Bulking up receptors with built‑in helpers
One key strategy is surprisingly simple: make the receptor “heavier” on purpose. Researchers genetically fuse small, stable helper proteins onto flexible parts of GPCRs to increase their size and rigidity, without blocking the important drug‑binding regions. Popular helpers include BRIL (a four‑helix bundle), PGS (a thermostable enzyme fragment), and a larger clamp‑like protein called calcineurin. These fusion partners are often inserted into a mobile inner loop that connects two helices. By stiffening this region, they reduce the blurring that comes from molecular motion and provide recognizable shapes that help computers align particles. Studies show that the exact position and orientation of the fusion piece can make or break a project: in one receptor, a differently placed PGS tag improved the resolution from a fuzzy 6 angstroms to a much clearer 3.7 angstroms.
Borrowing antibodies and designer binders as handles
A second route avoids building large fusions into the receptor and instead attaches high‑affinity “handles” from the outside. Tiny antibody fragments called nanobodies and related binders can be raised to recognize inactive GPCRs and latch onto flexible inner loops. A standout example is Nanobody‑6, which was first created as a sensor for one opioid receptor and then re‑used, with modest tweaks, to stabilize several other GPCRs in their inactive forms. Because it binds off‑center, it gives each particle a clear orientation cue, making image averaging more reliable. Researchers often extend this idea by adding a second layer—an ordinary antibody fragment that recognizes the nanobody itself—creating a bulkier, more rigid complex that shows up crisply in cryo‑EM. Other engineered binders, such as DARPins, can be used as modular spacers or as connectors to larger protein cages, further boosting signal from very small targets.
Smarter sample design and AI‑boosted image processing
Fusing helpers or adding binders is only part of the story. The review emphasizes that good structures begin with carefully engineered receptor constructs: trimming floppy tail regions, introducing stabilizing mutations, and using modern structure prediction tools to identify parts likely to cause trouble. On the imaging side, older tricks like phase plates, which increase contrast for faint particles, are being complemented or even replaced by artificial‑intelligence tools. Deep‑learning programs can pick out tiny particles hidden in noisy micrographs, and new algorithms can sort images into different shapes when receptors sample many conformations. Together with fiducial helpers, these advances push cryo‑EM towards reliably resolving membrane proteins that were once considered too small or too dynamic to study.
What this means for future drugs
The article concludes that there is no single “magic” helper that works for all receptors, but a growing toolbox of fusion partners, antibody‑like binders, and AI‑driven methods is steadily opening the inactive landscape of GPCRs to cryo‑EM. For non‑experts, the key takeaway is that by making small receptors appear bigger and more orderly to the microscope, scientists can finally obtain detailed snapshots of the exact shapes that many drugs prefer to bind. These structural blueprints should accelerate the design of medicines that more precisely turn these cellular switches off—or fine‑tune their activity—with fewer side effects.
Citation: Singh, S.K., Agrawal, M., Pattanayak, A. et al. Strategic advances for cryo-EM structural studies of small (<100 kDa) GPCRs. Commun Biol 9, 237 (2026). https://doi.org/10.1038/s42003-026-09516-y
Keywords: G protein-coupled receptors, cryo-electron microscopy, drug discovery, structural biology, nanobody scaffolds