Clear Sky Science · en
Decoding the ERS–CAF immunoregulatory axis via multimodal AI and its pan-cancer prognostic and therapeutic predictive value
Peering into Tumors Without a Scalpel
Cancer doctors increasingly recognize that what surrounds a tumor can matter as much as the tumor itself. But repeatedly sampling this hidden neighborhood with biopsies is invasive and often impractical. This study shows how artificial intelligence (AI) can read routine medical scans and microscope images to infer hard‑to‑measure immune and scar‑like processes inside tumors, potentially turning everyday imaging into a kind of "digital biopsy" that works across different cancers.
The Hidden Support Cells That Shape Cancer
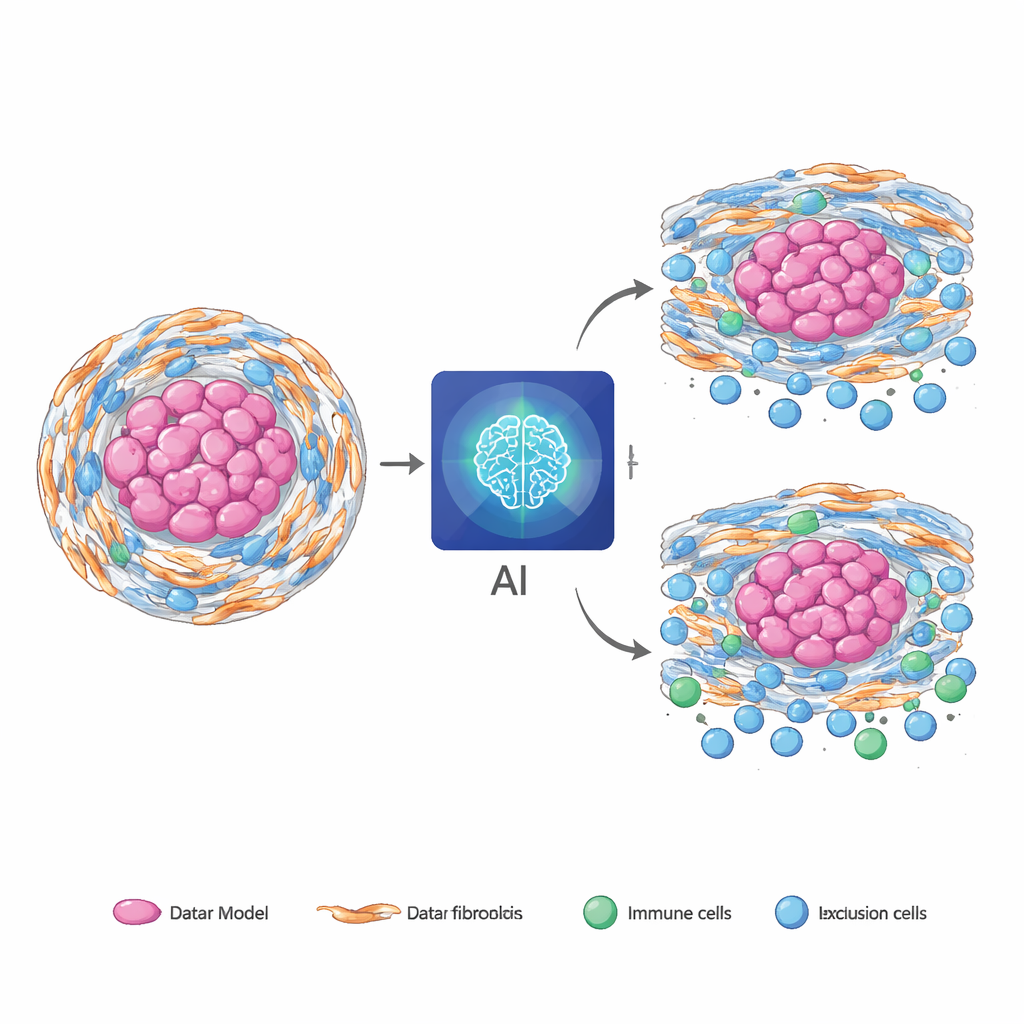
Many solid tumors are wrapped in a tough, fibrous shell made by specialized support cells called fibroblasts. When these cells are under stress inside the cell’s protein factory (the endoplasmic reticulum), they adopt an aggressive, cancer‑helping state. In chordoma, a rare bone cancer, these stressed fibroblasts build a dense matrix and help keep immune cells out, making treatments less effective. Similar fibrotic, immune‑poor environments show up in other cancers such as pancreatic and colorectal tumors, suggesting that this biology is not unique to one disease. The challenge is that current ways to measure these stressed fibroblasts and their immune‑blocking behavior rely on tissue samples and complex molecular tests, which are difficult to repeat and may miss important regions of the tumor.
Teaching AI to See Invisible Biology
The researchers asked whether standard pre‑surgery MRI scans and routine H&E pathology slides already contain visual clues about this stressed‑fibroblast immune barrier. They created three numerical “reference scores” from tumor RNA sequencing: one capturing how active the stress program in fibroblasts is, one summarizing how strongly these cells appear to be signaling to immune cells, and one describing how diverse the surrounding immune and support cell populations are. Instead of predicting thousands of genes, their AI was trained to predict just these three biologically meaningful scores from images alone. To do this, the team combined two branches: one that analyzes MRI texture and shape features, and another that scans thousands of small regions in the digital slide and uses a language‑guided attention mechanism to focus on areas that match expert descriptions of fibrotic, immune‑poor tissue.
Blending Scans and Slides for Stronger Signals
In 126 chordoma patients with matched MRI, pathology slides, RNA data, and follow‑up, the fused AI model outperformed models that used only MRI or only slides. Its predictions of the three molecular scores agreed closely with the RNA‑based measurements and remained well‑calibrated across different hospitals and scanners. When pathologists independently marked fibrotic and immune‑excluded regions, the AI’s “hotspots” tended to light up in those same areas, suggesting it was tracking genuine biology rather than just tumor size. The model also captured prognosis: higher predicted stress‑fibroblast and signaling scores were linked to worse survival, while greater predicted microenvironment diversity offered partial protection. Adding these AI‑derived scores to routine clinical factors improved the ability to separate high‑ and low‑risk patients over time.
From Rare Tumors to Common Cancers
A key test was whether a model trained entirely in chordoma could be used “as is” in other, more common cancers. Applied without retraining to pancreatic, stomach, and colorectal tumors from large public datasets, the slide‑only version of the model still showed meaningful alignment between its image‑based predictions and freshly computed RNA‑based scores. In some of these cancers, the AI scores improved prediction of patient survival beyond standard clinical information and helped distinguish which patients were more likely to benefit from chemotherapy. To make the approach easier to deploy where digital pathology is limited, the team distilled the full multimodal model into an MRI‑only version that retained most of the predictive power while running faster and using less computing power.
What This Could Mean for Patients
Taken together, the results support the idea that routine medical images quietly encode information about stressed support cells, immune exclusion, and microenvironment diversity—features that normally require expensive molecular tests. While the current work is retrospective and needs prospective validation, it points toward a future in which a standard scan and slide can non‑invasively flag tumors with a hostile, fibrotic immune barrier, guide which patients might benefit from additional testing or tailored therapies, and do so across multiple cancer types without extra burden on patients.
Citation: Zheng, BW., Xia, C., Tang, M. et al. Decoding the ERS–CAF immunoregulatory axis via multimodal AI and its pan-cancer prognostic and therapeutic predictive value. npj Digit. Med. 9, 199 (2026). https://doi.org/10.1038/s41746-026-02388-w
Keywords: tumor microenvironment, cancer imaging, artificial intelligence, fibroblasts, immunotherapy