Clear Sky Science · en
Large language models improve transferability of electronic health record-based predictions across countries and coding systems
Why smarter sharing of medical data matters
Hospitals and clinics around the world are sitting on a gold mine of information: electronic health records that capture people’s diagnoses, treatments, and outcomes over many years. In theory, this information could help doctors spot who is at high risk for serious illnesses early, long before symptoms become obvious. In practice, however, today’s computer models struggle to “travel” from one country or hospital system to another because each place records health data differently. This study introduces a new approach, called GRASP, that uses advances in artificial intelligence to bridge these gaps so that a model trained in one health system can work reliably in others.
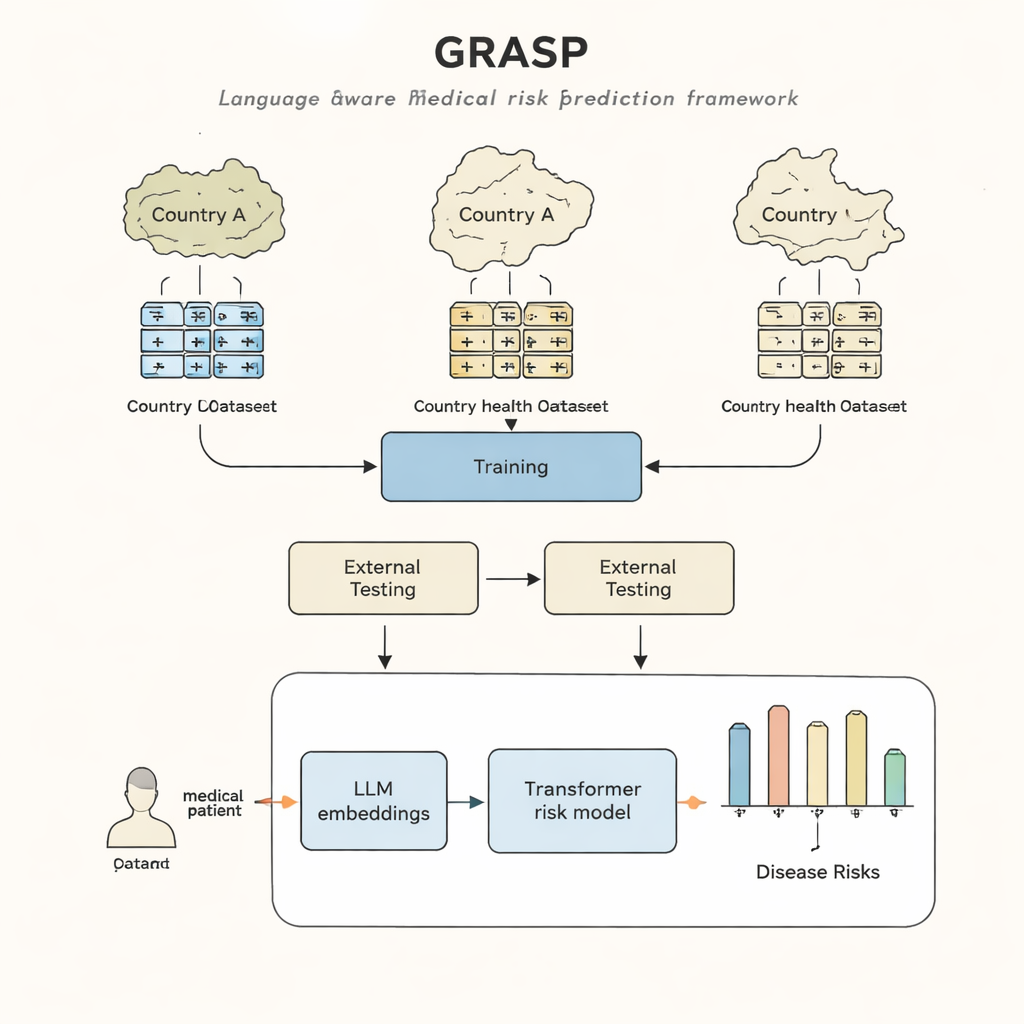
Different hospitals, different languages
Even when doctors treat the same disease, they often use different code systems and local habits to record it in the medical record. One hospital might store “high blood sugar” under one code, while another uses a different code for “hyperglycemia,” and a third uses yet another system entirely. Efforts to force everyone into a single common standard—such as big international coding schemes—are useful but slow, expensive, and still leave important differences behind. As a result, a computer model that predicts disease from records in one country can lose accuracy when applied elsewhere, limiting who can benefit from these tools.
Letting AI read the meaning, not just the code
The GRASP approach starts from a simple idea: instead of treating each medical code as a meaningless ID number, let a large language model read the human description behind it, such as “acute upper respiratory infection,” and turn that meaning into a numerical “embedding.” These embeddings place related concepts close together in a shared space, even if they come from different coding systems or countries. GRASP pre-computes such embeddings for millions of standard medical terms and stores them in a lookup table. A patient’s medical history is then represented as a series of these rich vectors, which are passed into a transformer network—a type of neural network well suited to handling collections of diverse inputs—to estimate that person’s risk of 21 major diseases plus overall risk of death.
Testing across countries and record systems
The researchers trained GRASP using data from nearly 400,000 participants in the UK Biobank, then tested it without retraining in two very different settings: the FinnGen project in Finland and a large hospital network in New York City. GRASP matched or outperformed strong alternatives, including a popular method called XGBoost and a similar transformer that did not use language-based embeddings. In Finland, GRASP did especially well, showing clear gains for conditions such as asthma, chronic kidney disease, and heart failure. Strikingly, even when the American hospital data were left in a different coding scheme instead of being converted to a shared standard, GRASP still delivered better predictions than demographics alone, because it could align codes purely by understanding the wording of their descriptions.
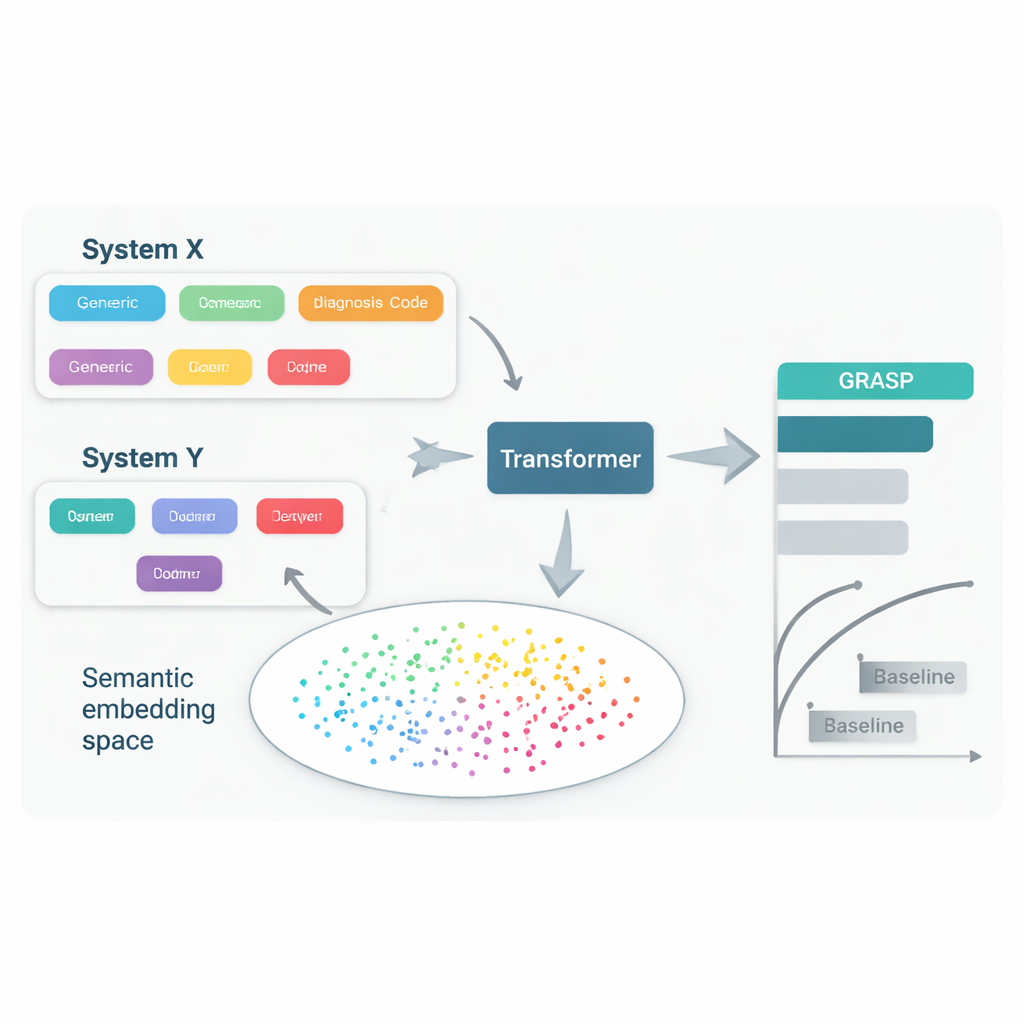
Getting more from less data
Another advantage of GRASP is efficiency. Because the language model has already learned that many medical concepts are related, the prediction network does not need to rediscover these links from scratch. When the authors trained GRASP on much smaller subsets of the UK data—down to just 10,000 people—it still outperformed competing models trained on the same limited samples, both in the UK and when transferred abroad. GRASP’s risk scores were also more closely aligned with people’s inherited genetic risk for several diseases, suggesting that it is capturing deeper aspects of disease susceptibility rather than just memorizing patterns in one dataset.
What this means for future care
For non-specialists, the key message is that GRASP shows how modern language-based AI can help different health systems “speak the same language” without forcing them into a single rigid coding scheme. By reading the meaning of medical terms, GRASP can make disease risk predictions that generalize better across countries and record formats, and it can do so with fewer patient examples. While the method still needs careful testing, recalibration, and checks for fairness before use in everyday care, it points toward a future in which powerful risk tools developed in one place can be safely and efficiently shared with hospitals and clinics around the world.
Citation: Kirchler, M., Ferro, M., Lorenzini, V. et al. Large language models improve transferability of electronic health record-based predictions across countries and coding systems. npj Digit. Med. 9, 177 (2026). https://doi.org/10.1038/s41746-026-02363-5
Keywords: electronic health records, disease risk prediction, large language models, medical data sharing, healthcare AI