Clear Sky Science · en
Masked autoencoding, generalizable pretraining, and integrated experts for enhanced glioma segmentation
Why smarter scans matter for brain tumors
Brain tumors called gliomas are among the deadliest cancers, yet doctors still spend a lot of time manually tracing tumor borders on MRI scans. This careful outlining guides surgery and radiation, but it can take 15–20 minutes per patient and must be repeated over time. The study introduces MAGPIE, an artificial intelligence system that learns from tens of thousands of brain scans without human labels, then needs only a handful of expertly labeled cases to reliably map out gliomas. For patients, this could mean faster, more consistent treatment planning even at hospitals that lack large, curated datasets.
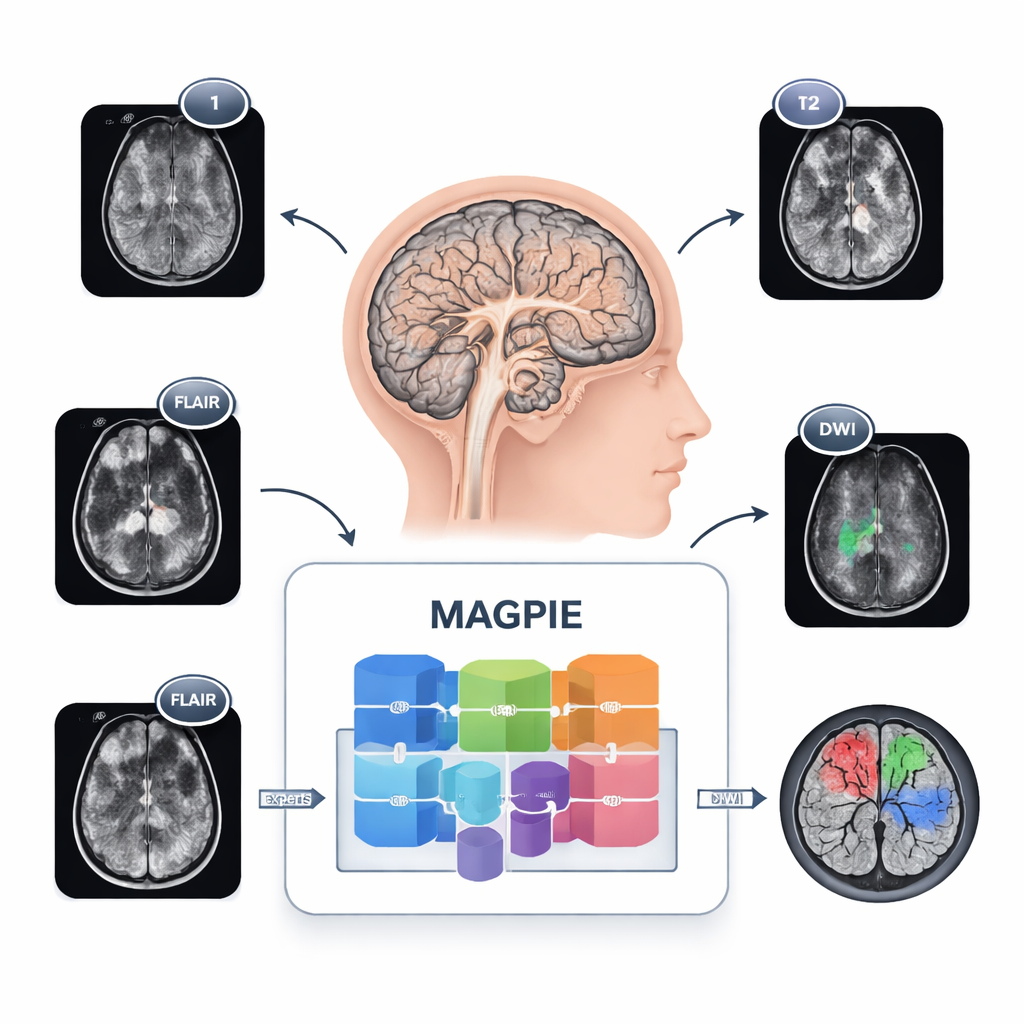
Seeing tumors in a new way
Gliomas are tricky to map because they do not form neat balls. Cancer cells spread along brain wiring, creating fuzzy margins and tiny satellite spots that are hard to see. Different hospitals also use different MRI settings and combinations of sequences, so a tool trained in one place can stumble in another. MAGPIE tackles all of this at once. It was first exposed to 43,505 unlabeled brain MRI scans drawn from many studies and scanner types. During this stage, it learned general patterns of healthy and diseased brain tissue by trying to rebuild missing pieces of images and by comparing different augmented views of the same brain, forcing it to focus on stable, meaningful features rather than brittle pixel details.
Letting multiple experts share the work
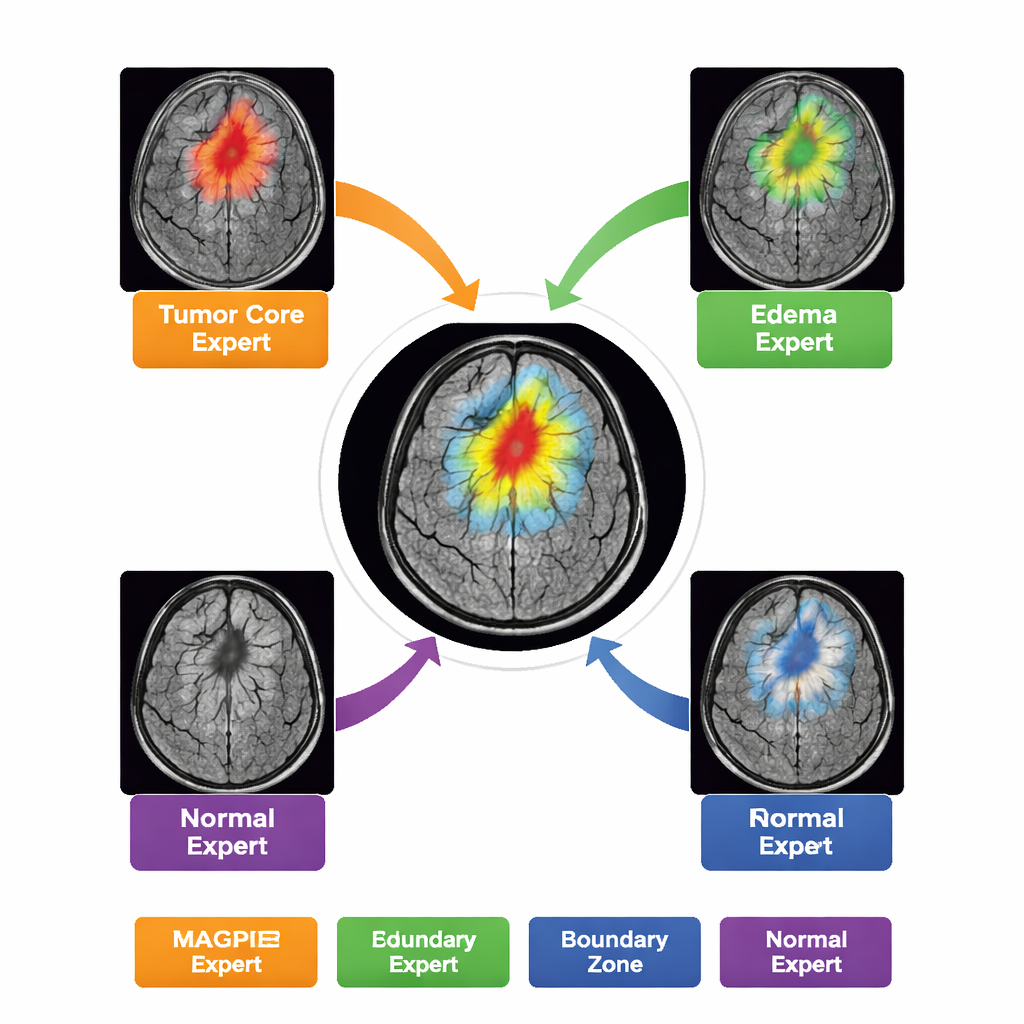
Rather than acting as a single, monolithic model, MAGPIE contains a “mixture of experts” inside. When it analyzes a new scan, it only activates a small subset of eight specialized sub-networks for each region of the image. Over training, these experts naturally divide up the task: some become sensitive to the bright, actively growing rim of tumor; others lock onto the dead core; others learn the hazy ring of swelling around the tumor; and some focus mainly on normal brain background and boundaries. The authors show this by measuring how strongly each expert’s activity overlaps with the different tumor zones drawn by radiologists. This division of labor improves accuracy while keeping the computation manageable—only about half of the model’s parameters are active for any given patch.
Handling messy, real-world scans
Clinical MRI protocols are far from uniform. Some patients have four sequences, others fewer; machines from different manufacturers produce subtly different images. MAGPIE’s design treats each MRI sequence like a separate “token” and learns how much weight to give each one on the fly, instead of expecting a fixed set of inputs in a fixed order. This channel-agnostic approach means the system can adapt if, for example, a contrast-enhanced sequence is missing but FLAIR is present. The model also uses advanced attention mechanisms that let it both “see far,” capturing long-distance spread along white matter tracts, and “see precisely,” spotting very small lesions only a few millimeters across.
Doing more with far fewer labels
After pretraining, the researchers fine-tuned MAGPIE on just 20 fully labeled glioma cases and compared it with standard models trained from scratch under the same conditions. On a major brain tumor benchmark (BraTS21), MAGPIE achieved a Dice score—a common overlap measure in medical imaging—of about 61%, beating the best scratch-trained version by roughly 2.6 percentage points and outperforming a strong earlier self-supervised method without showing any harmful “negative transfer.” On challenging out-of-distribution data—scans from different diseases, scanner types, and image settings—it also held up better, reaching over 70% Dice on one white-matter lesion dataset without any extra tuning. Crucially, this level of performance normally requires on the order of 400 labeled cases; MAGPIE reaches it with only about 5% of that effort.
What this could mean for patients and clinics
For non-experts, the central message is that MAGPIE turns a mountain of unlabeled MRI scans into a powerful assistant that needs very little expert training to become clinically useful. It can outline complex brain tumors with realistic boundaries, detect small satellite spots that other systems miss, and keep working reliably when scans come from unfamiliar machines or lack certain sequences. That combination could cut radiologists’ annotation time by around 95%, lower the barrier for smaller hospitals to deploy advanced imaging AI, and support more precise surgery and radiation planning. While further validation on rare tumor types and low-grade cases is still needed, this study shows how carefully designed self-supervised learning can bring robust, data-efficient brain tumor segmentation closer to everyday clinical reality.
Citation: Xie, M., Xiao, Q., Wu, H. et al. Masked autoencoding, generalizable pretraining, and integrated experts for enhanced glioma segmentation. npj Digit. Med. 9, 163 (2026). https://doi.org/10.1038/s41746-026-02347-5
Keywords: glioma segmentation, brain MRI, self-supervised learning, mixture of experts, medical imaging AI