Clear Sky Science · en
Multimodal fusion of pathology and radiology foundation models for WHO 2021 glioma subtyping
Bringing Two Views of Brain Tumors Together
When a person is diagnosed with a brain tumor, doctors need to know not just that a tumor is present, but what exact kind it is. Different tumor types respond very differently to surgery, radiation, and drugs. Today, this detailed "subtyping" usually requires genetic tests that can be slow, costly, and not available everywhere. This study explores whether a smart computer system that looks at both brain scans and microscope images of tumor tissue can reliably infer these subtypes, potentially speeding up and broadening access to precision treatment.
Why Tumor Type Matters
Adult diffuse gliomas are among the deadliest brain cancers, yet they often look similar on standard scans and under the microscope. Modern guidelines group them into three genetic subtypes that differ greatly in how aggressive they are and how long patients tend to live. The current gold standard for telling these subtypes apart depends on molecular tests of tumor DNA. These tests require extra tissue, specialized labs, and waiting days to weeks for results. The authors ask whether routinely collected magnetic resonance imaging (MRI) and digital pathology slides could be combined to extract enough information to stand in for some of this genetic workup.
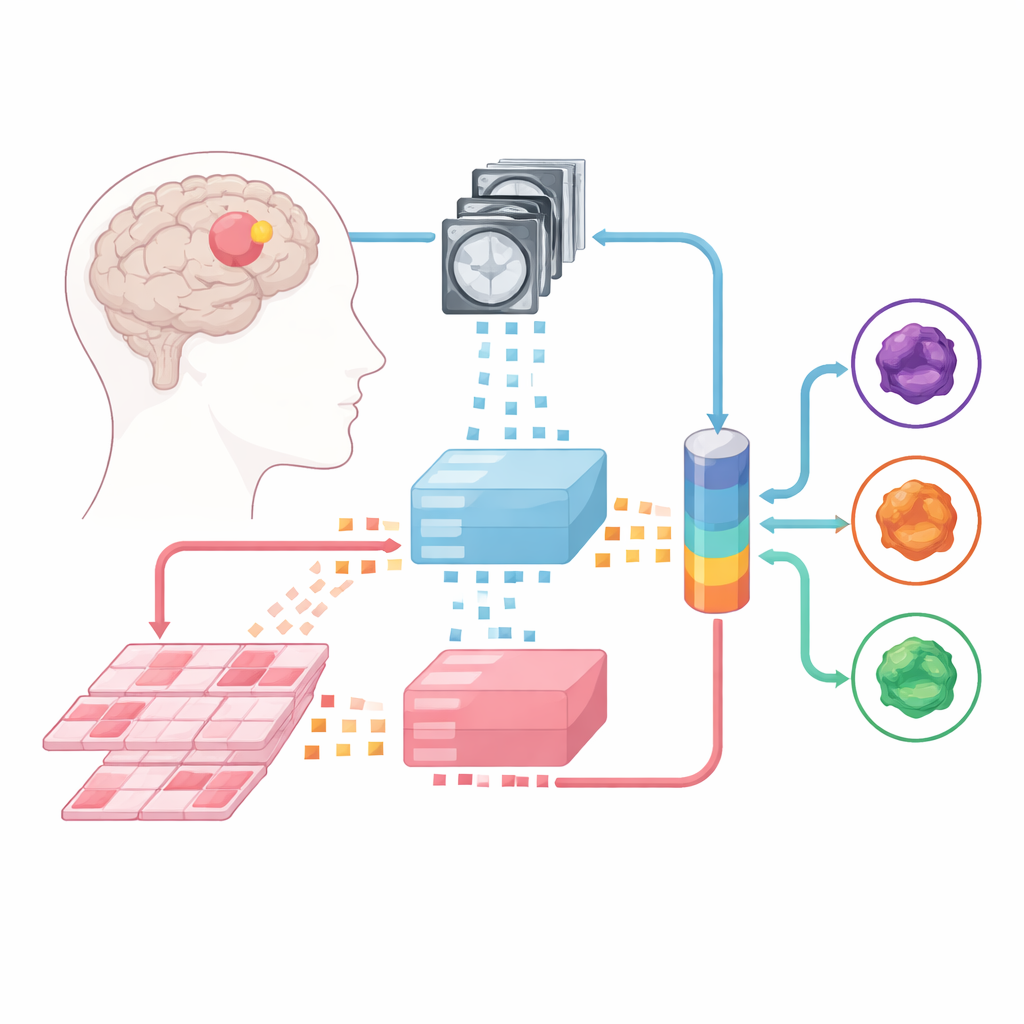
Teaching Machines to Read Scans and Slides
The team built upon large “foundation models” – powerful image analyzers trained beforehand on enormous collections of medical images. One such model digests multiparametric MRIs, and another handles high-resolution pathology slides made from tumor tissue. Each incoming case is split into small image patches, which the foundation models convert into numerical fingerprints. On top of these fixed "experts," the researchers trained three types of fusion models that learn to combine information from both MRI and pathology: a late-fusion design, an early-fusion design, and a more flexible mixture-of-experts architecture that can dynamically decide how much to rely on each source.
Mixing Modalities Without Matching Patients
A practical obstacle for such multimodal methods is that hospitals rarely have large datasets where every patient has both MRI and pathology images neatly paired. Instead of relying on perfectly matched data, the authors assembled separate collections: hundreds of MRI cases from several centers and hundreds of pathology cases from another resource, plus a smaller set of 171 patients from a public cancer project who had both. During training, they often paired an MRI from one person with a pathology slide from another, as long as the tumors belonged to the same subtype. Surprisingly, models trained on such "unpaired" data worked just as well as those trained on true patient pairs, and clearly better than simply averaging two separate single-modality models.
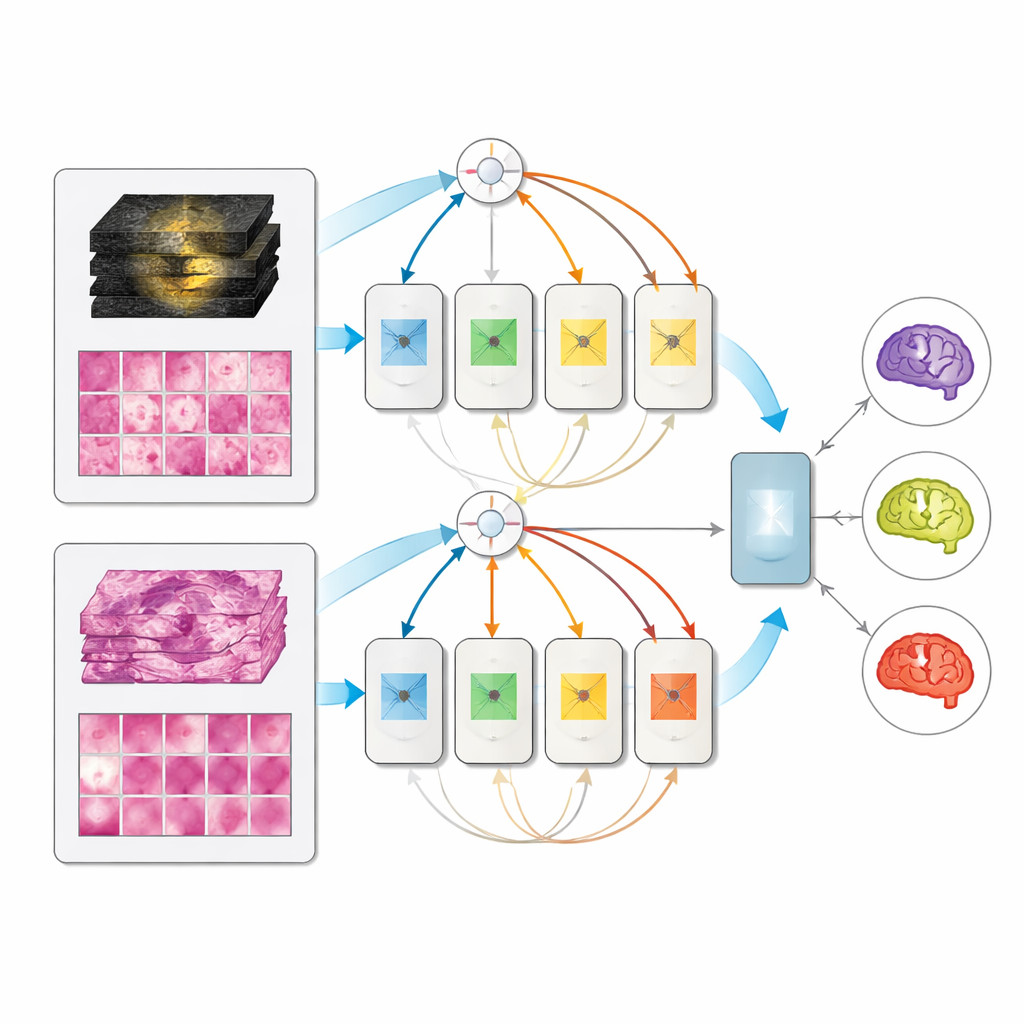
A Single Model That Adapts to What Is Available
On the held-out set of 171 fully characterized patients, all multimodal models beat their single-input counterparts, and the mixture-of-experts design performed best, reaching very high scores in distinguishing the three subtypes. Notably, when only MRI or only pathology was provided at test time, the multimodal model did not collapse; it performed about as well as dedicated single-modality models. This means a clinic could deploy one unified system that uses whatever information is available—pre-surgery scans alone, post-surgery tissue alone, or both together—rather than maintaining separate tools for each scenario.
Seeing What the Model Sees
To build trust in the system’s decisions, the researchers probed where the model “looks” and which image features matter most. Attention maps showed that the joint model spreads its focus more broadly across the tumor and its surroundings, in both MRI and tissue slides, and that more diffuse attention tended to coincide with correct predictions. A deeper analysis of learned features revealed patterns that match known medical clues: for example, MRI features highlighting contrast-enhancing tumor cores and distorted fluid spaces helped separate more aggressive tumors, while tissue features capturing classic cell shapes and textures helped recognize specific glioma subtypes. Interesting gaps also emerged: the model did not strongly encode some textbook signs of the most aggressive tumors, supporting the idea that it often treats that group as a “default” unless there is strong evidence for a more favorable subtype.
What This Could Mean for Patients
In plain terms, this work shows that an AI system combining brain scans and microscope images can classify brain tumors more accurately than systems that look at only one type of image, and that it can be trained even when the two image types are not available from the same patients. If further validated in larger and more diverse groups, such tools could help doctors estimate tumor subtype earlier and more widely, especially in settings where genetic testing is limited. While they are not a replacement for molecular tests, they may act as fast, low-cost guides that point surgeons and oncologists toward the most likely diagnosis and the most appropriate treatment path.
Citation: Saueressig, C., Scholz, D., Raffler, P. et al. Multimodal fusion of pathology and radiology foundation models for WHO 2021 glioma subtyping. npj Precis. Onc. 10, 118 (2026). https://doi.org/10.1038/s41698-026-01366-5
Keywords: glioma subtyping, multimodal imaging, artificial intelligence, MRI and pathology, brain cancer diagnosis