Clear Sky Science · en
The next layer: augmenting foundation models with structure-preserving and attention-guided learning for local patches to global context awareness in computational pathology
Teaching Computers to Read Cancer Slides
When a pathologist examines a cancer biopsy under the microscope, they don’t just look at individual cells—they look at patterns, neighborhoods, and how tumor, immune cells, and normal tissue are arranged together. Today’s artificial intelligence systems for digital pathology are very good at spotting details in tiny image patches, but they often miss this bigger picture. This study introduces EAGLE-Net, a new AI approach that helps computers see cancer slides more like human experts do, by paying attention to both local details and the overall layout of tissue on the slide.
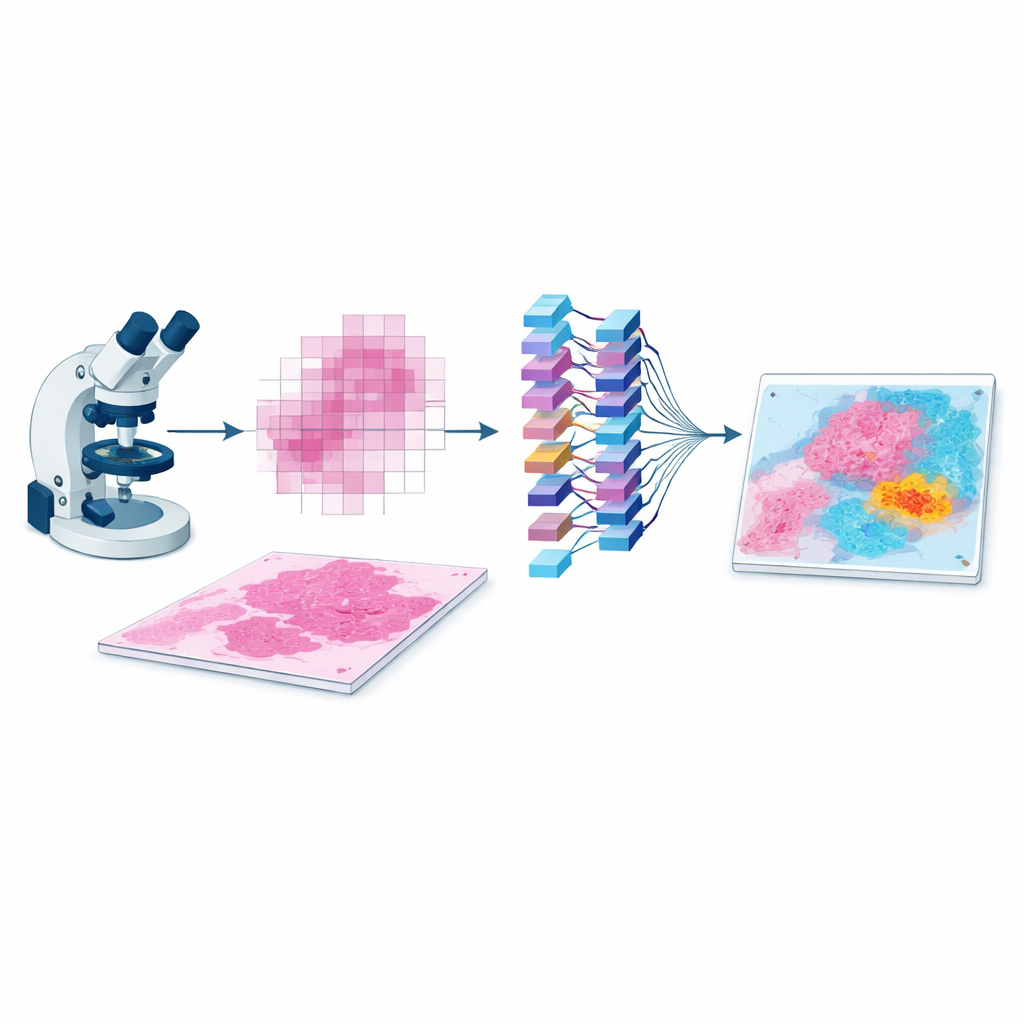

Why the Layout of Tumor Tissue Matters
A tumor is more than a clump of cancer cells. It lives in a busy neighborhood filled with blood vessels, immune cells, connective tissue, and areas of scarring or cell death. How these elements are arranged—their distances, boundaries, and mixtures—can indicate how aggressive the cancer is and how a patient might respond to treatment. Conventional AI systems in pathology usually chop a whole-slide image into thousands of small tiles and analyze them almost in isolation. They then try to guess the patient’s diagnosis or outcome by pooling information from all tiles. This strategy often ignores how tiles relate to each other in space, which can weaken predictions and make AI heatmaps look scattered or hard to interpret.
A New Way to Capture the Bigger Picture
EAGLE-Net is designed to bridge that gap between local detail and global structure. It starts from powerful “foundation models” that already know how to extract rich visual features from small slide patches. On top of that, it adds a new module that encodes where each patch came from on the slide, preserving the true geometry of the tissue rather than squeezing it into a distorted grid. Using multi-scale filters, EAGLE-Net learns patterns that span from tiny cell-level changes to broader tissue structures like tumor borders and surrounding stroma. It then uses an attention mechanism—essentially a way of assigning importance scores—to focus on patches and neighborhoods that are most relevant for predicting diagnosis or survival.
Letting the Model Learn From Neighborhoods, Not Just Dots
A key innovation in EAGLE-Net is how it teaches the network to value not only the most important tiles but also their nearby neighbors. During training, the method repeatedly identifies the tiles that the model finds most informative and then encourages it to consider surrounding tiles within a small radius as part of the same meaningful region. This “neighborhood-aware” learning nudges the model to form smooth, contiguous regions of attention that align with how pathologists view tumor fronts, immune clusters, and other microenvironments. At the same time, an additional term in the training process actively pushes the model to ignore background or empty areas, reducing the risk of false highlights on stray artifacts or whitespace.
Proving Its Value Across Many Cancer Types
The researchers tested EAGLE-Net on nearly 15,000 whole-slide images spanning 10 different cancers, including lung, kidney, stomach, uterine, thyroid, colorectal, and prostate tumors. They evaluated two main tasks: predicting how long patients would survive and classifying tumor types or grades. Across most cancer cohorts, EAGLE-Net matched or beat several leading attention-based methods, often improving survival prediction scores and classification accuracy by a few percentage points, which is meaningful at population scale. It also performed strongly when paired with three very different underlying foundation models, showing that its design is flexible and not tied to a single feature extractor.
Seeing Inside the Model’s Reasoning
Beyond raw accuracy, the team carefully examined where EAGLE-Net “looked” on the slides. Compared with other methods, its attention maps formed smoother, more coherent regions that hugged tumor boundaries and captured invasive edges, necrotic pockets, and immune cell clusters. Quantitative comparisons with expert-drawn tumor masks showed that EAGLE-Net’s highlighted regions overlapped better with the true tumor, made fewer false hits on normal tissue, and more faithfully reproduced complex tumor shapes. The model also devoted a larger share of its attention to tumor, necrosis, and immune compartments, and less to normal lung or blood vessels, mirroring what a pathologist would prioritize when judging prognosis.
What This Means for Future Cancer Care
In practical terms, EAGLE-Net demonstrates that adding spatial awareness and neighborhood reasoning on top of existing pathology AI can improve both performance and interpretability. Rather than treating a slide as a bag of disconnected tiles, the method learns to recognize biologically meaningful niches—tumor borders, immune-rich regions, and patterns of invasion—that matter for patient outcomes. Because it works with many different foundation models and does not require labor-intensive pixel-level labeling, EAGLE-Net could be broadly applied to large archives of digital slides. With further validation and integration into clinical workflows, such systems may help pathologists stratify patients more precisely, discover new tissue-based biomarkers, and ultimately guide more tailored cancer treatments.
Citation: Waqas, M., Bandyopadhyay, R., Showkatian, E. et al. The next layer: augmenting foundation models with structure-preserving and attention-guided learning for local patches to global context awareness in computational pathology. npj Precis. Onc. 10, 109 (2026). https://doi.org/10.1038/s41698-026-01312-5
Keywords: computational pathology, cancer prognosis, digital pathology AI, tumor microenvironment, EAGLE-Net