Clear Sky Science · en
Generalisation of automatic tumour segmentation in histopathological whole-slide images across multiple cancer types
Why this matters for cancer care
Cancer diagnosis still relies on experts carefully examining glass slides of stained tissue under a microscope—a time‑consuming task made harder by growing case numbers and a shortage of pathologists. This study asks a simple but powerful question: can one artificial‑intelligence system reliably find cancerous areas in digital microscope images for many different tumour types, instead of building a separate tool for each cancer? If so, it could ease workloads, speed up diagnosis, and extend advanced analysis even to rarer cancers where data are scarce.
From glass slides to digital helpers
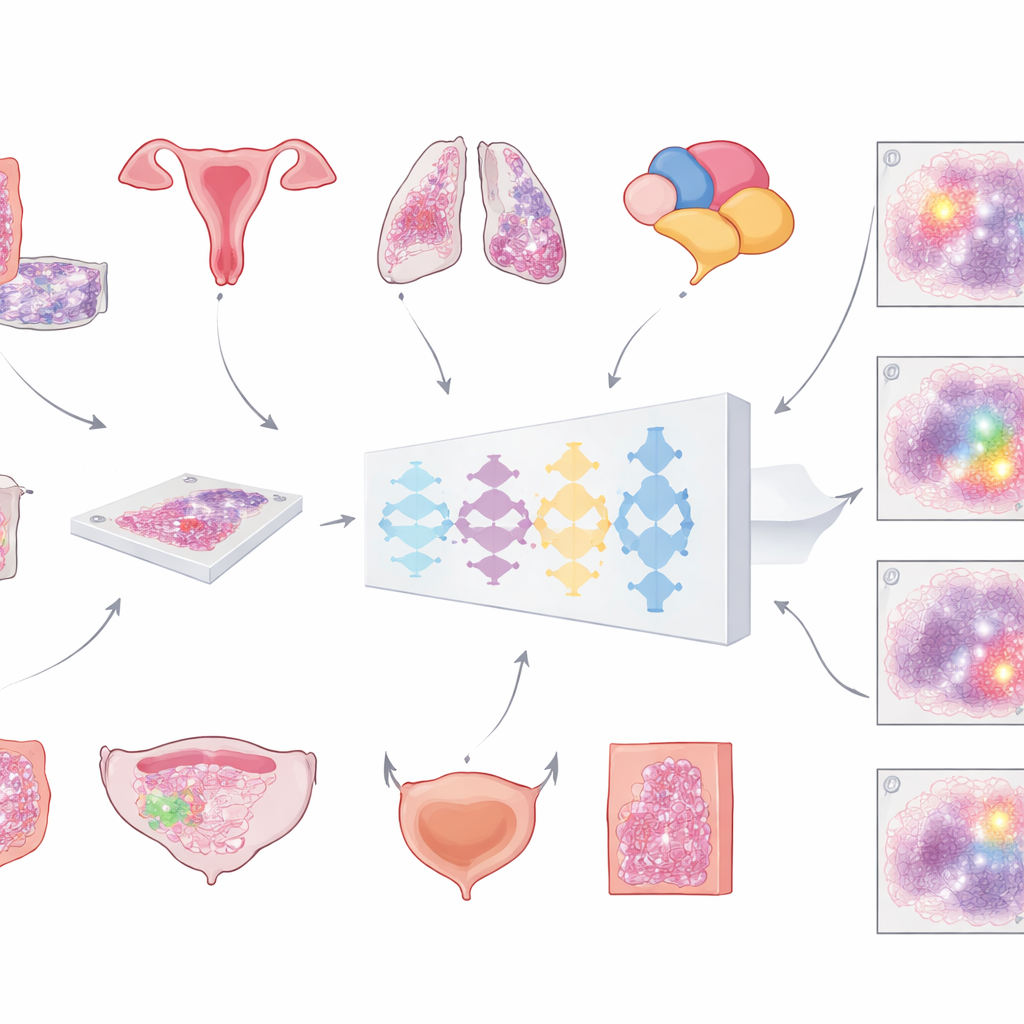
Modern hospitals are increasingly scanning microscope slides to create huge, detailed “whole‑slide images” of tumours. The first crucial step for any computer‑based analysis is to separate cancerous tissue from everything else—normal cells, fat, empty glass, and artefacts. Until now, most automated tools have been trained on just one cancer type, limiting where they can be used. The team behind this work set out to build a single, universal model that could spot tumour regions across multiple common cancers in slides stained with the routine haematoxylin and eosin dyes. Their vision was a general tool that could plug into many diagnostic workflows without redesigning it each time.
Training one model to see many cancers
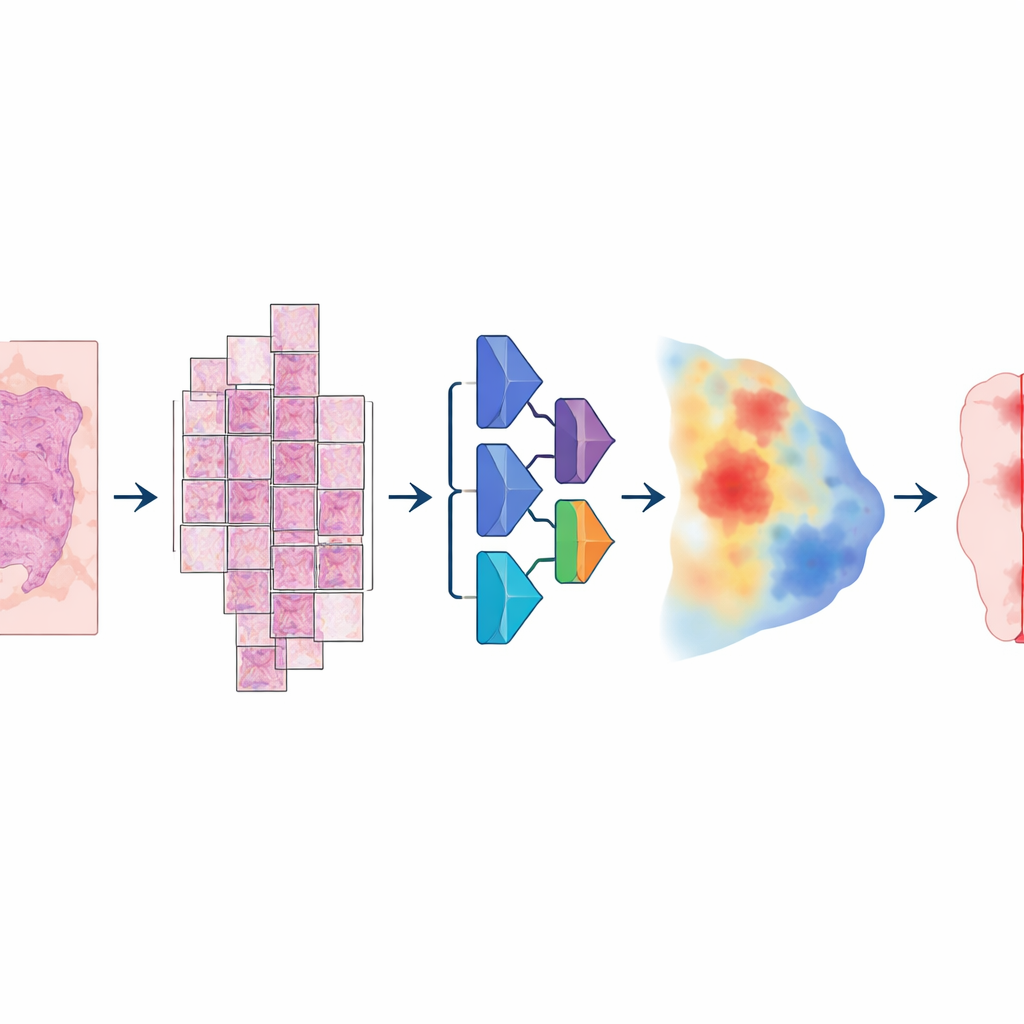
To build this model, the researchers assembled an unusually large and varied collection of digital slides: more than 20,000 whole‑slide images from over 4,000 patients with colorectal, endometrial, lung, and prostate cancer. All samples came from standard formalin‑fixed, paraffin‑embedded tissue and were scanned on two different high‑resolution slide scanners. A pathologist carefully outlined tumour areas on every slide, providing the “ground truth” the computer would learn from. The model followed a multi‑step pipeline: each enormous image was broken into large overlapping tiles, passed through a deep neural network that estimated, for every pixel, how likely it was to be tumour, and then reassembled into a smooth map that was finally converted into a clean tumour‑versus‑non‑tumour mask.
Putting the system to the test
Crucially, the team did not stop at training performance. They tested the same model on more than 3,000 additional patients across six cancer types—including breast and bladder cancers that were never used during training—and on slides from multiple hospitals and scanners. Accuracy was measured mainly with a standard overlap score (the Dice coefficient), which reaches 100% when the computer’s tumour outline perfectly matches the pathologist’s. For large, intact tumour samples in colorectal, endometrial, lung, prostate, and breast cancer, the average overlap exceeded 80% and often 90%. In large external collections from The Cancer Genome Atlas, drawn from many labs and scanners worldwide, performance again stayed above 80%, suggesting that the model generalises well beyond its home institution.
Where it struggles and how it compares
The main weakness emerged in early‑stage bladder cancers sampled by a procedure that produces tiny, fragmented tissue pieces. In these cases, the model often failed to mark any tumour at all, especially when the cancer area was very small. Yet when it did detect tumour, overlap with the pathologist’s outlines was high, and simple adjustments to the final thresholds improved results—hinting that the underlying network recognised the pattern but the post‑processing was too strict. The researchers also built four “specialist” models, each trained on a single cancer type, and found that none meaningfully outperformed the general model in its own domain. In contrast, these specialist systems largely failed when applied to other cancer types, while the general model remained robust. Compared with a popular, more generic medical segmentation tool that requires user hints, the new model usually performed as well or better while remaining fully automatic.
What this means for patients and doctors
For non‑experts, the key takeaway is that one well‑designed AI system can reliably highlight cancerous tissue on digital slides across several major tumour types, without needing custom versions for each disease or scanner. It does not replace the pathologist, but it can pre‑mark likely tumour regions, support consistent measurements, and free specialists to focus on the most challenging cases. The current version still misses some very small or early‑stage tumours—particularly fragmented bladder samples and likely other biopsy‑like tissues—so it is not yet suited to detecting the faintest traces of cancer. Nonetheless, the study shows that broad, “pan‑cancer” tumour segmentation is feasible in real‑world conditions and can form a robust first step for future automated tools that assess cancer grade, predict treatment response, or guide precision therapies.
Citation: Skrede, OJ., Pradhan, M., Isaksen, M.X. et al. Generalisation of automatic tumour segmentation in histopathological whole-slide images across multiple cancer types. npj Precis. Onc. 10, 107 (2026). https://doi.org/10.1038/s41698-026-01311-6
Keywords: digital pathology, deep learning, tumour segmentation, whole-slide imaging, pan-cancer model