Clear Sky Science · en
Large-scale single-cell analysis and in silico perturbation reveal dynamic evolution of HCC: from initiation to therapeutic targeting
Why liver cancer’s inner life matters
Hepatocellular carcinoma, the most common form of liver cancer, is notoriously difficult to treat. Many patients do not respond well to today’s drugs, and tumors often come back or spread. One big reason is that each tumor is a chaotic mix of different cells that evolve over time and talk constantly to their surroundings. This study uses powerful single-cell and spatial gene-reading technologies to watch that evolution unfold, cell by cell, from early tumor growth to advanced, treatment‑resistant disease, and to pinpoint weak spots that new therapies might exploit.
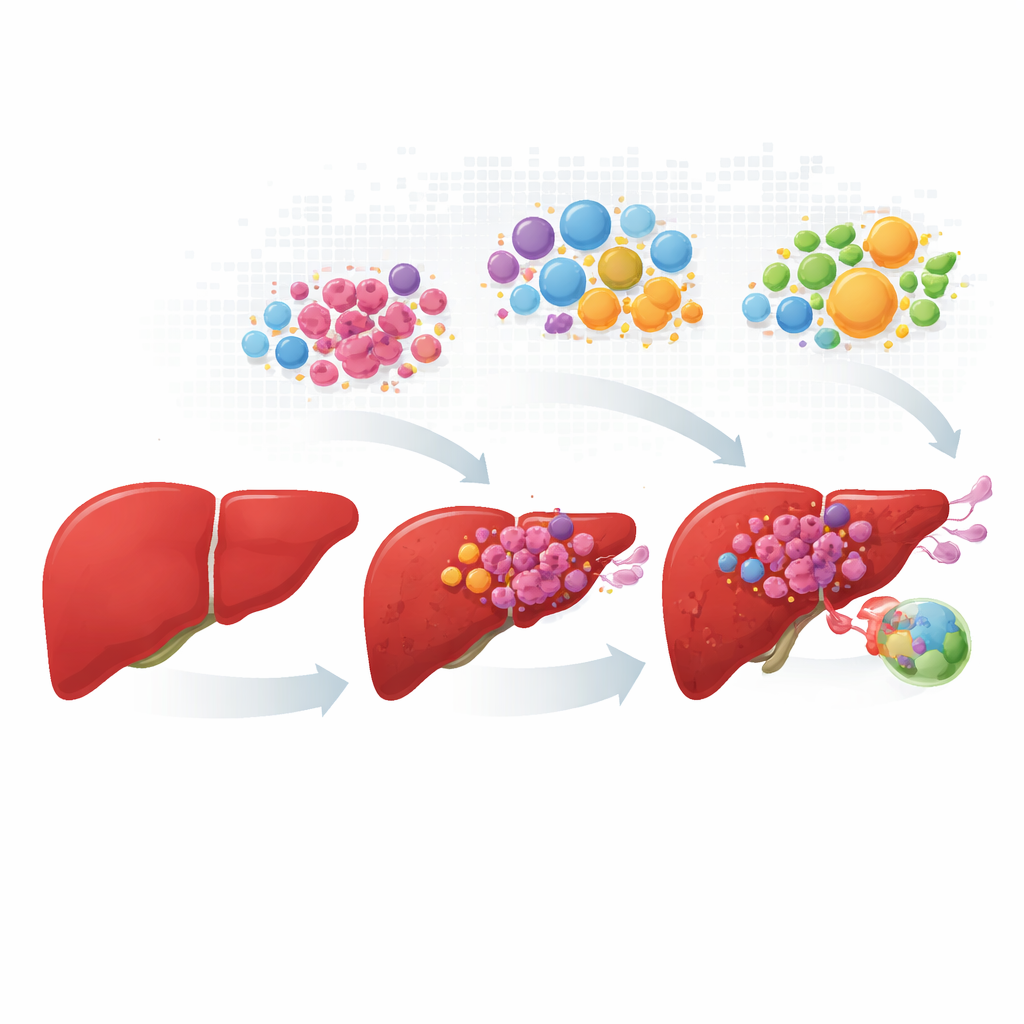
Following cancer cells from early change to spread
The researchers assembled and harmonized data from more than a hundred patients, covering normal liver, primary tumors, growths in blood vessels, and lymph node metastases. Instead of treating each tumor as a uniform mass, they profiled hundreds of thousands of individual cells, then mapped where those cells sit in actual tissue slices. Cancerous liver cells fell into four recurring “personalities.” One group kept many normal liver functions and metabolic tasks; the other three showed stress‑driven growth, high activity of the MYC growth pathway, or invasive, inflammation‑linked behavior. These four states appeared in different proportions as the disease advanced, with fast‑growing and invasive cells dominating in vessel and lymph node lesions.
Four tumor personalities with different futures
The four cancer cell states were not just academic categories; they tracked with how patients fared. Tumors rich in the liver‑like, metabolically active state were linked to better survival, whereas tumors dominated by the other three aggressive states tended to have worse outcomes. Yet even the “good” state came with a twist: it was strongly enriched in patients who did not benefit from the liver‑cancer drug sorafenib, likely because these cells preserve detox machinery that can inactivate medicines. By replaying how cells move along developmental paths, the team showed that invasive, inflammatory cells can branch toward more differentiated or more highly proliferative states, suggesting that tumor cells can switch identities, potentially escaping therapies aimed at a single state.
The supporting cast: immune and stromal accomplices
Cancer cells do not act alone. The study uncovered specialized immune cells and support cells that either fight tumors or quietly help them. Among macrophages, the tissue’s garbage‑collectors and sentinels, some types produced inflammatory signals that can aid anti‑tumor immunity, while others were metabolically rewired, suppressed helpful signals, and promoted blood vessel growth and invasion. These “bad actor” macrophages expanded in advanced samples. Likewise, certain endothelial cells lining blood vessels and fibroblasts that build connective tissue formed distinct subsets tied to poor prognosis. In advanced disease, a particular endothelial subtype and two fibroblast types created tightly connected hubs that remodeled the extracellular scaffold, fueled new blood vessel formation, and correlated with the most aggressive cancer cell states.
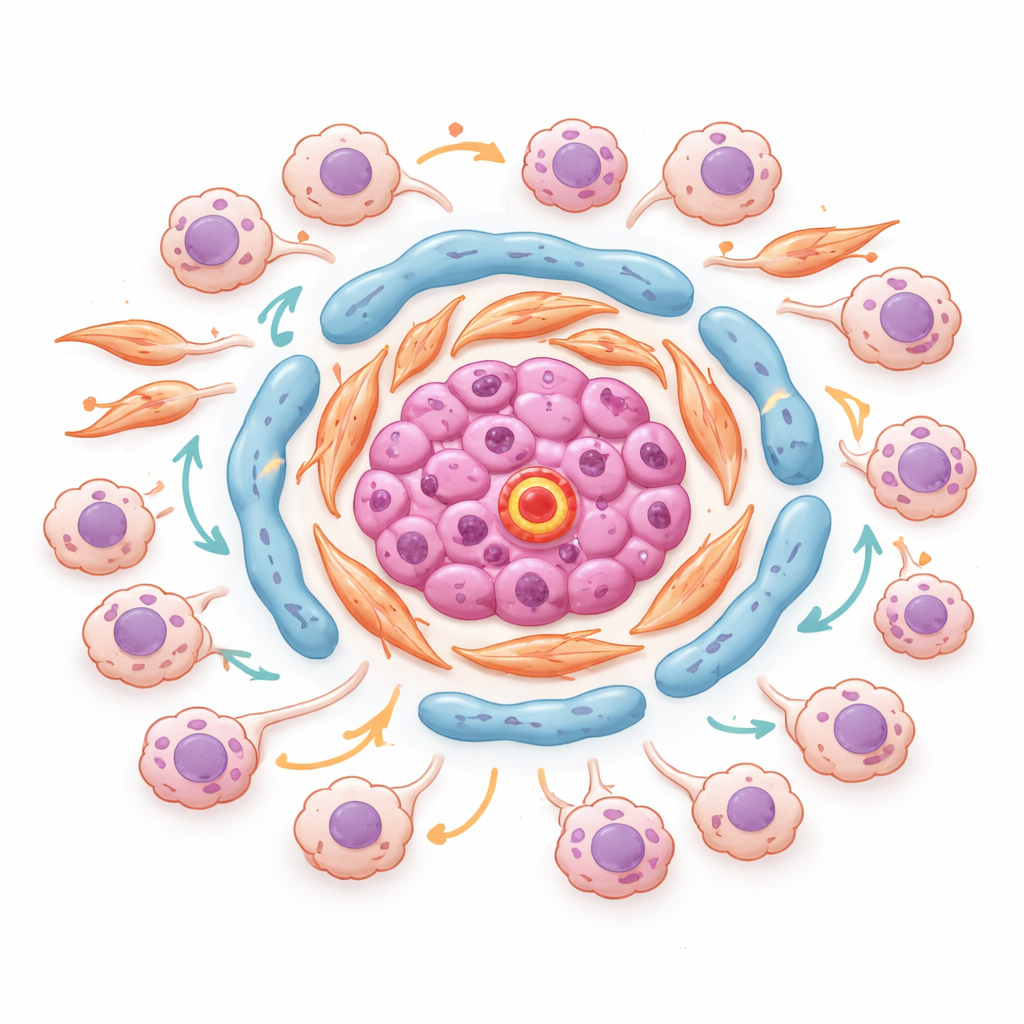
Seeing dangerous neighborhoods inside tumors
Using spatial transcriptomics, which measures gene activity without losing the map of where each spot sits in the tissue, the authors identified organized “neighborhoods” inside tumors. In one representative tumor, endothelial cells rich in a molecule called ESM1 clustered with lipid‑hungry and matrix‑producing fibroblasts at the core of stromal islands. Around these cores accumulated stress‑driven and invasive cancer cells, in regions with strong TGF‑beta and low‑oxygen signals. Key transcription factors that control how cells respond to stress and inflammation were shared between these cancer states and were most active exactly where tumor cells touched these stromal hubs. This suggests that specific micro‑niches at tumor–stroma borders actively encourage cancer cells to change state, become more aggressive, and resist therapy.
Finding a shared weak point for new treatments
To move from description to intervention, the team used a machine‑learning model trained on millions of cells to simulate “virtual knockouts” of genes and predict which changes could push harmful cells back toward healthier states. Comparing macrophages, endothelial cells, and cancer cells, they found a small set of genes whose removal helped normalize all three. Among these, HSP90B1 stood out: cancer cell lines depend heavily on it, it is overproduced in liver tumors, especially in those that resist targeted drugs and immunotherapy, and high levels predict poorer survival. HSP90B1 encodes a stress‑handling chaperone in the cell’s protein‑folding factory that supports many surface receptors and signaling pathways. Because selective drugs against this chaperone already exist and can also reshape the immune environment, the study proposes HSP90B1 as a promising target to overcome resistance, especially when combined with current liver‑cancer treatments.
What this means for patients and therapies
Taken together, this work shows that liver cancer is not one disease but a dynamic ecosystem of cell states and niches that shift as the tumor grows and spreads. By tying specific cancer cell personalities, support cells, and spatial neighborhoods to patient outcome and drug response, the study offers a roadmap for more precise treatment: matching therapies to dominant cell states, disrupting stromal hubs that cultivate aggressive behavior, and targeting shared dependencies like HSP90B1 that many tumor and microenvironment cells rely on. While further laboratory and clinical testing is needed, the atlas this team built provides a detailed blueprint for turning the tumor’s internal complexity from an obstacle into an opportunity for smarter, more durable liver cancer therapies.
Citation: Xia, P., Shuang, S., Fu, D. et al. Large-scale single-cell analysis and in silico perturbation reveal dynamic evolution of HCC: from initiation to therapeutic targeting. npj Precis. Onc. 10, 100 (2026). https://doi.org/10.1038/s41698-026-01307-2
Keywords: hepatocellular carcinoma, single-cell analysis, tumor microenvironment, spatial transcriptomics, HSP90B1